Bioprospecting Agricultural Soils for Isolation and Characterization of Biosurfactant Producing Bacteria
Department of Microbiology, Bhavan’s Vivekananda College of Science, Humanities and Commerce, Secunderabad, India.
Corresponding Author E-mail: aparnachandra7673@gmail.com
Download this article as:
ABSTRACT:Biosurfactants are environmentally friendly surface-active compounds produced by various microorganisms, including bacteria such as Bacillus spp., recognized for their versatile capabilities, have garnered attention as potential biosurfactant producers. This study aims to elucidate the biosurfactant production process from these bacterial strains, highlighting their growth conditions, strain selection, extraction, purification, characterization and potential applications. The biosurfactant exhibit superior biodegradability,low toxicity and high efficacy in reducing surface and interfacial tension compared to synthetic surfactants. Biosurfactant plays a crucial role in inhibiting biofilm formation and extracellular polymeric substances, making them promising candidates for antimicrobial and antifouling applications. Additionally, they have been explored for bioremediation, enhanced oil recovery and pharmaceutical formulations due to their emulsification and antimicrobial properties. Understanding the significance of biosurfactant producing bacteria the present study was undertaken to isolate and characterize biosurfactant producing bacteria from agricultural fields. Five isolates identified were labelled asBS1, BS2, BS3, BS4 and BS5 respectively. The isolates were evaluated for multiple biosurfactant producing properties like emulsification index, Drop collapse test, hemolytic test, C-TAB test, Oil displacement test and Anti biofilm test. The five isolates were characterized using Morphological and Biochemical characterization and found to be belonging toBacillus sps as all are endospore forming and Gram-Positive rods. Among all the tested isolates BS1 and BS4 showed good biosurfactant production compared to BS2, BS3 and BS5.
KEYWORDS:Agricultural Soil Microbiota; Bioprospecting; Biosurfactant; Microbial Characterization; Surface Active Biomolecules
Introduction
Agricultural ecosystems are dynamic environments where a wide array of microbial communities thrive and contribute to soil fertility, plant growth promotion, and pollutant degradation. Among these microorganisms, biosurfactant-producing bacteria have garnered significant attention due to their ability to produce surface-active compounds that reduce surface and interfacial tensions.1 These biosurfactants, which include glycolipids, lipopeptides, phospholipids, and polymeric surfactants, are biodegradable, eco-friendly, and have diverse applications in agriculture, bioremediation, pharmaceuticals, and food industries.2
The excessive use of chemical fertilizers and pesticides in agriculture has led to detrimental effects on soil health, water quality, and biodiversity. In recent years, there has been a pressing demand for sustainable practices that minimize environmental damage while maintaining productivity. One promising approach is the exploitation of biosurfactant-producing bacteria for eco-friendly applications such as biocontrol agents, biofertilizers, and bioremediation tools.3 These microbes not only assist in the degradation of xenobiotic compounds but also facilitate nutrient availability by increasing the bioavailability of hydrophobic substances like hydrocarbons and pesticides.
Agricultural fields, especially those subjected to extensive pesticide use, represent a rich reservoir for isolating potent biosurfactant-producing strains. The exposure of indigenous microbial communities to such environmental stresses often results in the selection of metabolically versatile bacteria capable of degrading recalcitrant pollutants and producing biosurfactants to aid in this process.4 The isolation and characterization of such bacteria are crucial not only for understanding their ecological roles but also for exploiting their biosurfactants for industrial and agricultural applications.
Biosurfactants have been found to enhance the bioavailability of hydrophobic compounds, increase soil wettability, and promote plant-microbe interactions.5 Furthermore, some biosurfactant-producing bacteria possess antagonistic properties against phytopathogens, making them valuable candidates for biocontrol agents.6 For instance, lipopeptide biosurfactants produced by Bacillus subtilis have shown strong antifungal activity against Fusarium oxysporum and Rhizoctonia solani in controlled experiments.7
Characterization of biosurfactants involves evaluating their surface activity, emulsification ability, critical micelle concentration (CMC), and stability under diverse physicochemical conditions. Moreover, molecular identification of the biosurfactant-producing isolates and genetic analysis of biosynthetic pathways (e.g., srfA, rhlAB, licA) provide deeper insights into their metabolic capabilities and potential for commercial exploitation.8
Recent studies have employed advanced techniques such as whole-genome sequencing, LC-MS/MS, and FTIR spectroscopy to decipher the structural complexity of biosurfactants and to identify novel strains with enhanced productivity.9 For example, metagenomic analyses of pesticide-contaminated soils have revealed the presence of diverse bacterial taxa belonging to genera Pseudomonas, Bacillus, and Serratia, many of which carry biosurfactant biosynthetic genes and exhibit high emulsification index.10 These findings underscore the potential of agricultural soils as valuable niches for isolating new biosurfactant producers.
In the context of green technology and sustainability, microbial biosurfactants offer a superior alternative to synthetic surfactants due to their low toxicity, high specificity, and environmental compatibility. Their application in bioremediation, especially in the degradation of petroleum hydrocarbons and organophosphates, is well documented.11 Additionally, their role in enhancing plant growth through improved nutrient uptake and pathogen suppression supports the development of bioformulations for integrated pest and nutrient management.
Despite significant progress in biosurfactant research, challenges remain in terms of large-scale production, cost-effectiveness, and regulatory approvals. Therefore, it is essential to explore native microbial strains from diverse ecosystems, particularly those with a history of agrochemical exposure, to identify robust biosurfactant producers with commercial potential.
This study aims to isolate and characterize biosurfactant-producing bacteria from agricultural fields with a history of pesticide usage. By focusing on morphological, biochemical, and molecular characterization, as well as assessing the biosurfactant activity using standard assays such as drop collapse, oil displacement, and emulsification index, this research endeavors to expand the repository of functional microbial strains for biotechnological applications. The study also discusses the possible implications of these findings in enhancing agricultural sustainability and environmental remediation.
Materials and Methods
Collection of soil sample
The soil sample used for the isolation of pesticide degrading bacteria was collected from agricultural field from Irwin village, Telangana where pesticide is used extensively. Samples were collected at a depth of 5cm to 10cm in the field. The soil sample was air dried at room temperature to remove the excess moisture and it was passed through a 2mm sieve to remove the unwanted debris and stored in a polyethylene bag in refrigerator for further experimental study.
Isolation and characterization of Bacteria
The soil samples werecollected and serial dilution were performed, 0.1 ml of sample, spread on nutrient agar plate and incubated at 37 °C for 24-48 hours.The bacterial isolates were streaked onto sterile nutrient agar plates using a sterile loop. These plates were subsequently incubated at 37°C for 24 hours. Morphological characteristics of the colonies, Gram staining andEndospore staining was performed for preliminary identification. Biochemical tests like Catalase, Oxidase, IMViC tests, Starch Hydrolysis, Nitrate reduction test and Gelatin Liquefaction were performed for tentative identification of bacteria as per the standard protocols mentioned in Laboratory manual in Microbiology.12
Identification using ABIS online software
The identification of bacterial isolates were performed using the Advanced Bacterial Identification Software (ABIS) Online, a bioinformatics-based platform for microbial classification.13
Biosurfactant production tests:
Emulsification Index (E24)
The emulsification index was assessed by adding 1.5 mL of various hydrocarbons (engine oil, crude oil, mustard oil, olive oil, petrol) to 1.5 mL of cell-free supernatant (CFS), followed by vortexing at high speed for 2 minutes. The mixture was then incubated at room temperature for 24 hours.14
Emulsification Index E24 (%) = (Height of the emulsified layer / Total height of liquid column) × 100
Hemolysis Test
Hemolysis activity was performed on 5% sheep blood agar plates. A loopful of 24 h old culture of each isolate was spot inoculated on blood agar medium, incubated at 37 °C for 48 hour. and observed for clear zone of hemolysis around the bacterial colonies indicating the presence of biosurfactant activity.15
C-TAB/Methylene-blue Test
The CTAB (Cetyl trimethyl ammonium bromide) agar plate method was performed to test the presence of anionic surfactants.100μl of test sample was added to wells in agar plates. The composition of agar medium is 0.5g of glucose, 0.25 g of peptone, 0.025g of beef extract 0.02 g of C-TAB, 0.001g of Methylene blue, 0.013g of yeast extract and 2g agar in 100ml water,sterilized and used for assay. The pH was adjusted to 7.2. After incubation presence of clear zone indicates the presence of anionic surfactant.16
Oil Spreading Assay
Oil spread assay was performed as per the standard protocol. 20 ml of distilled water was taken in a Petri dish, and 1ml of crude oil was overlaid uniformly on water surface. Gently add 100ul of cell free supernatant over the oil surface. Biosurfactant activity was measured by observation of clear zone on oil surface due to oil displacement.17
Thin Layer Chromatography (TLC)
Overnight grown cultures were centrifuged at 8000rpm for 15 minutes.Collect the supernatant and do acid precipitation with 6N Hcl. Incubate overnight at 40C.After incubation centrifuge again to collect precipitate.Add methanol to allow phase separation.The methanolic extract was then centrifuged to remove the undissolved fraction, while supernatant was collected and subjected to rotary vacuum evaporation at 30 °C. The yellowish- brown substance thus obtained was used for further analysis.. Preliminary characterization of the biosurfactant can be done by TLC, the components of the crude extract can be identified by TLC on silica gel 60 in chloroform: methanol: water (70:10:0.5).18
Antibacterial Assay
The antibacterial properties of partially purified biosurfactants were evaluated against lab isolates Bacillus subtilis, Pseudomonas putida, Escherichia coli and Staphylococcus aureus.Streptomycin disc is used as positive control. The antibacterial activity was done using the agar diffusion method. A volume of 100 µLof the test microorganism was evenly spread on a Muller Hinton agar plate. Wells were then created using a sterile well borer, and each well was filled with 100 μL of biosurfactant at a concentration of 1 mg/ml. The zones of growth inhibition were measured after a 24-hour incubation period, with results reported as the average of independent experiments.19
Antifungal Activity
The antifungal activity of the test samples (BS1 to BS5) was performed using the Agar Well Diffusion Method with Lab isolates Penicillium, Aspergillus, and Fusarium sps respectively. The inoculated plates are then incubated at 28–30°C for 48–72 hours.After incubation, the plates are examined for zones of inhibition, which indicate antifungal activity. The diameter of the inhibition zones (in cm or mm) is measured using a ruler or caliper, and the results are compared to the control to determine the effectiveness of the test samples.20
Antibiofilm Assay
The antibiofilm assay was performed to evaluate the ability of test samples to inhibit biofilm formation. Microtiter Plate Assay was performed to check biofilm inhibition. Overnight grown Pseudomonas aeruginosa (Biofilm producing culture) was used in assay. Each well is inoculated with 180 µL of the bacterial suspension along with 20ul of biosurfactant test samples at appropriate concentrations. Control wells contain only bacterial suspension without the test sample. The plates are incubated at 37°C for 24–48 hours to allow biofilm formation. After incubation, the wells were gently washed three times with sterile phosphate-buffered saline (PBS) to remove planktonic (free-floating) bacteria, leaving only the adherent biofilm. The biofilm was then fixed by air drying and stained with 0.1% crystal violet for 15 minutes. Excess stain was removed by washing with sterile distilled water, and the bound crystal violet is solubilized using 95% ethanol or acetic acid. The absorbance was measured at OD570 nm using a microplate reader to quantify biofilm biomass. A lower OD value in treated wells compared to the control indicates biofilm inhibition.21
Results
Morphological and Cultural Identification:
Table 1: Morphological and Cultural Identification
| S.NO | Sample | Gram Staining | Endospore Staining | Microscopic Observation |
| 1 | BS1 | Gram Positive | Spore former | Rod shaped chain |
| 2 | BS2 | Gram Positive | Spore former | Diplo Bacillus |
| 3 | BS3 | Gram Positive | Spore former | Clusters of Bacillus |
| 4 | BS4 | Gram Positive | Spore former | Chain shaped Bacillus |
| 5 | BS5 | Gram Positive | Spore former | Bacillus |
Table 1 depicts Morphological and cultural identification of bacterial isolates with gram staining. All the five isolates BS1, BS2, BS3,BS4 and BS5 are gram positive bacilli, spore forming bacteria but exhibited different morphologyunder microscopic observation.
Biochemical tests performed for isolates:
Table 2: Biochemical Test
| S.NO. | Biochemical Test | BS1 | BS2 | BS3 | BS4 | BS5 |
| 1 | Catalase Test | +ve | +ve | +ve | +ve | +ve |
| 2 | Oxidase Test | -ve | +ve | +ve | +ve | +ve |
| 3 | Indole Test | -ve | -ve | -ve | -ve | -ve |
| 4 | Methyl Red test | -ve | -ve | -ve | -ve | -ve |
| 5 | Voges – Prousker Test | -ve | -ve | -ve | -ve | -ve |
| 6 | Citrate Utilization Test | -ve | -ve | -ve | -ve | -ve |
| 7 | Starch Hydrolysis Test | -ve | +ve | -ve | +ve | +ve |
| 8 | Nitrate Reduction Test | +ve | +ve | +ve | +ve | +ve |
| 9 | Gelatin Liquefaction Test | +ve | +ve | +ve | +ve | +ve |
The biochemical profiles of all the five bacterial isolates(BS-1 to BS-5) are presented in Table 2. All isolates showed catalase positive activity demonstrating their ability to decompose hydrogen peroxide into water and oxygen. All isolates were positive for nitrate reduction and gelatin liquefaction suggesting their ability to reduce nitrate to nitrite and production of extracellular proteolytic enzymes. For Oxidase test only BS1 showed negative test indicating slight variation in respiratory enzyme systems among isolates.All isolates were negative for IMViC tests. For starch hydrolysis BS2,BS4 and BS5 exhibited positive results while others were negative.
Identification using ABIS online software
The Morphological, Biochemical and Enzymatic test results were subjected to identification upto genus level using ABIS online software
Table 3: Predicted genera and percentage similarity of Isolates using ABIS online
| Test Isolate | Genus identified | % Similarity |
| BS1 | Bacillus firmus. | 80.2 |
| BS2 | Bacillus megaterium. | 87.7 |
| BS3 | Bacillus cirroflagelosus | 90.8 |
| BS4 | Bacillus megaterium | 86.8 |
| BS5 | Brevibacillus brevis | 96 |
Bacterial identification of the five biosurfactant producing isolates was performed using ABIS online software. The predicted genera and percentage similarities are presented in Table 3. All five isolates were identified as members of genus Bacillus and related genera specifically BS1 showing 80.2% similarity with Bacillus firmus,BS2 showing 87.7% similarity with Bacillus megaterium,BS 3 showing 90.8% similarity with Bacillus cirroflagelosus,BS4 showing 86.8% similarity withBacillus megateriumand BS5 showing 96% similarity withBrevibacillus brevis.
Biosurfctant Production Tests
Emulsification Index
Table 4: Emulsification Index of Isolates with different Oils
| EMULSIFICATION INDEX [E24] | ||||||||
| S.No. | Oils | Positive
Control (%) |
Negative
Control (%) |
BS1(%) | BS2(%) | BS3(%) | BS4(%) | BS5(%) |
| 1. | Engine Oil | 39 | 14 | 25 | 46 | 36 | 53 | 28 |
| 2. | Petrol | 25 | 3.0 | 11 | 28 | 21 | 28 | 21 |
| 3. | Mustard Oil | 36 | 14 | 28 | 21 | 18 | 25 | 18 |
| 4. | Olive Oil | 39 | 5.0 | 11 | 28 | 33 | 28 | 28 |
| 5. | Crude Oil | 53 | 18 | 28 | 14 | 7 | 21 | 7.0 |
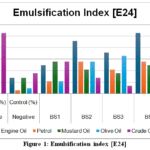 |
Figure 1: Emulsification index [E24]
|
The Emulsification Index (E24) analysis showed that among the isolates tested, BS4 exhibited the highest emulsification activity, particularly against engine oil (53%) and olive oil (28%), outperforming even the positive control(Table 4). BS2 also showed strong emulsification, while BS3 and BS5 demonstrated moderate to low activity (Figure 1).
Hemolysis Test
Table 5: Betahemolysis test of isolates
| S.No. | Sample | Type of Hemolysis | Zone Of Inhibition (Cm) |
| 1 | BS1 | Beta hemolysis | 2.4 |
| 2 | BS2 | Beta hemolysis | 2.5 |
| 3 | BS3 | Beta hemolysis | 1.8 |
| 4 | BS4 | Beta hemolysis | 1.7 |
| 5 | BS5 | Beta hemolysis | 2.5 |
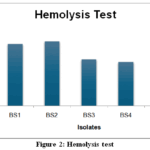 |
Figure 2: Hemolysis test
|
Hemolytic activity is a preliminary indicator of biosurfactant production and its potential cytotoxicity. In this study, all five bacterial isolates (BS1–BS5) exhibited beta hemolysis, characterized by clear zones around colonies on blood agar, confirming their ability to lyse red blood cells (Figure 2). The zone of hemolysis ranged from 1.7 cm (BS4) to 2.5 cm (BS2 and BS5), with BS2 and BS5 demonstrating the most pronounced activity(Table 5).
C-TAB/Methylene-blue Test
No clear zones of inhibition were observed for any of the five samples in the C-TAB/Methylene-blue test. This indicates that none of the samples demonstrated anionic surfactant activity or the ability to inhibit microbial growth under the tested conditions.
Oil Spreading Assay
Oil spreading test was performed with crude oil.
Table 6: Oil spreading assay of biosurfactant
| S.No. | Sample | Oil Displacement (Cms) |
| 1 | Positive Control | 0.48 |
| 2 | Negative Control | 0 |
| 3 | BS1 | 0.46 |
| 4 | BS2 | 0.26 |
| 5 | BS3 | 0.43 |
| 6 | BS4 | 0.36 |
| 7 | BS5 | 0.15 |
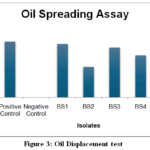 |
Figure 3: Oil Displacement test
|
Table 6 presents the results of Oil spreading assay done with crude oil. Among the isolates, BS1 and BS3 exhibited significant oil displacement (0.46 cm and 0.43 cm respectively), comparable to the positive control (0.48 cm), indicating strong biosurfactant activity. The presence of measurable oil displacement zones for all isolates except the negative control indicates biosurfactant production with varying efficiencies.
Extraction of crude sample and TLC
Table 7: Rf Values calculated from TLC sheet
| Sample | Rf Values |
| BS1 | 0.11 |
| BS2 | 0.22 |
| BS3 | 0 |
| BS4 | 0.22 |
| BS5 | 0.4 |
TLC analysis was performed to determine the Rf values of biosurfactant extracts obtained from five bacterial isolates. Among the samples BS5 showed highest Rf value(0.40) followed by BS2 and BS4 with identical Rf values of 0.22 while BS1 exhibited lower Rf value of 0.11. No detectable amount was seen in BS3. These results indicate variation in mobility of biosurfactant compounds among isolatessuggesting differences in their chemical composition and polarity.
Antibacterial Assay
Table 8: Antibacterial activity Biosurfactants
|
S.NO |
Bacteria |
Control |
Zone Of Inhibition (Cm) | ||||
| BS1 | BS2 | BS3 | BS4 | BS5 | |||
| 1 | Bacillus | 1.8 | 0 | 1.6 | 0 | 0 | 1.5 |
| 2 | Pseudomonas | 0 | 1.0 | 1.2 | 0 | 1.4 | 1.8 |
| 3 | Staphylococcus | 1.0 | 0 | 0 | 0 | 0 | 0 |
| 4 | Escherichia coli | 2.5 | 0 | 1.0 | 0 | 0 | 0 |
For Bacillus, the control showed an inhibition zone of 1.8 cm, while BS2 (1.6 cm) and BS5 (1.5 cm) exhibited some inhibitory effects, though slightly weaker than the control. BS1, BS3, and BS4 showed no inhibition against Bacillus. In the case of Pseudomonas, the control did not exhibit any inhibition, but BS1 (1.0 cm), BS2 (1.2 cm), BS4 (1.4 cm), and BS5 (1.8 cm) showed noticeable antibacterial activity, with BS5 demonstrating the strongest effect. For Staphylococcus, although the control had a 1.0 cm inhibition zone, none of the tested samples (BS1 to BS5) showed any antibacterial activity. Regarding Escherichia coli, the control inhibition was 2.5 cm, while BS2 exhibited only 1.0 cm of inhibition, indicating a much weaker effect. BS1, BS3, BS4, and BS5 showed no inhibition against E. Coli.
Antifungal Activity
The control inhibition zones for Penicillium notatum, Aspergillus niger, and Fusarium were 2.5 cm, 2.0 cm and 2.0 cm, respectively. None of the tested samples (BS1 to BS5) showed any inhibition against these fungi.The tested samples (BS1–BS5) were ineffective in inhibiting the growth of Penicillium, Aspergillus, and Fusarium, as they did not produce a measurable zone of inhibition.
Antibiofilm Assay
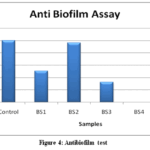 |
Figure 4: Antibiofilm test
|
Figure 4 demonstrates Anti biofilm assay with various isolates.Anti Biofilm assay of isolates showed 50.7% inhibition with BS1,96.3% with BS2, 32.6% with BS3 and 58.6% with BS5. However no Biofilm inhibitory activity was demonstrated with BS4 isolate. These results suggest that each isolate showed high variation in their Anti biofilm potential which is commonly noted in various soil bacteria in agricultural fields.
Discussion
Morphological, Cultural and Biochemical characterization of isolates showed that the isolates share several common metabolic traits with minor variations noted in oxidase and starch hydrolysis. These results indicate metabolic diversity among biosurfactant producing bacteria from agricultural soils. The ABIS based identification suggests majority of the isolates belong to Bacillusand closely related genera(Table 3). Species such as Bacillus megaterium and Bacillus firmus are known for their metabolic versatility,enzyme production and ability to synthesize surface active compounds supporting their potential role in biosurfactant production.
Emulsification indicates (E24) results demonstrate that all five isolates possess biosurfactant producing ability but the efficiency varied depending on hydrophobic substrate used.Such substrate dependent emulsification is commonly noted in soil bacteria. Among the tested isolates, BS4’s potential for bioremediation and hydrocarbon degradation is very high. Comparable emulsification efficiencies have been reported in all Bacillus species known for lipopeptide biosurfactant production.Moderate E24 values with petrol and vegetable oils suggest broad substrate specificity. The variation among isolates emphasizes the value of agricultural soils as sources of biosurfactant-producing bacteria with promising environmental applications such as oil spill cleanup and enhanced biodegradation.22,23.
Beta hemolysis is often associated with the production of potent lipopeptide biosurfactants, such as surfactin and iturin, which integrate into lipid membranes.24 The findings on betahemolysissuggest that the isolates, particularly BS2 and BS5, produce effective biosurfactants with potential antimicrobial or surface-active properties. However, the cytotoxic nature of such compounds must be further evaluated before industrial or environmental applications. The hemolysis test serves as a useful screening tool in identifying promising biosurfactant-producing strains from agricultural soils. The absence of clear inhibition zones in C-TAB/Methylene blue assay indicates that none of the isolates produced detectable anionic biosurfactants under experimental conditions. Similar observations have been reported in some Bacillusstrains where biosurfactant production depends on growth conditions and medium composition.
The results on Oil spreading activity align with previous studies demonstrating agricultural soils as rich sources of biosurfactant-producing microbes.25.The variation in oil displacement suggests diverse surfactant production capacities, warranting further molecular and structural characterization for potential industrial applications.Figure 3 demonstrates that Oil spreading assay results indicate that the isolates possess biosurfactant producing capability as reflected by displacement of crude oil on water surface. Higher oil displacement was noted in BS1 and BS3 suggesting greater biosurfactant activity than other isolates. Similar results are reported in earlier studies which produce surface active compounds capable of reducing surface tension and spreading oil. The variation among isolates may be due to differences in biosurfactant concentrationor molecular structure. These findings suggest the potential of isolates for application in hydrocarbon degradation and environment bioremediation.
Thin Layer Chromatography (TLC) analysis revealed the presence of different biosurfactant types among the bacterial isolates(Table 7). BS5 exhibited the highest Rf value (0.40), indicating the presence of relatively non-polar compounds, possibly glycolipids or lipopeptides. BS2 and BS4 had identical Rf values (0.22), suggesting similar biosurfactant profiles. The possible biosurfactant maybeSophorolipids produced by Cabdidabombicola, while BS1 (Rf = 0.11) likely produced more polar molecules which may be Rhamnolipids as per the previous studies. BS3 showed no detectable movement (Rf = 0), implying either absence or extremely polar biosurfactants. These findings correlate with earlier studies where Rf values were used to distinguish surfactant classes aiding further biochemical characterization.26,27
Biosurfactants produced by species of Bacillus are known to exhibit antimicrobial properties by disrupting microbial cell membranes. The absence of inhibition against Staphylococcus suggests selective antimicrobial activity of the produced compounds. Variations in inhibition zones among isolates may be attributed to differences in biosurfactant concentration and molecular composition. These findings highlight the potential of the isolates for antimicrobial and biotechnological applications.
Biosurfactant-producing bacteria isolated from agricultural soils exhibited significant anti-biofilm activity, particularly strains BS2 and BS5. The strain BS4 showed no activity, indicating strain-specific variation. These findings suggest potential application in agricultural and clinical sectors for biofilm inhibition. The reduction in absorbance confirms disruption of pre-formed biofilms, consistent with earlier studies.28,29The biosurfactants amphipathic nature likely contributes to membrane destabilization and biofilm dispersal. Further structural characterization and genetic analysis are required to elucidate biosurfactant mechanisms. The results align with sustainable biocontrol strategies using eco-friendly microbial solutions.
Conclusion
The present study demonstrates that the biosurfactant-producing isolates (BS1–BS5) exhibited diverse functional properties across multiple assays, indicating their potential for varied biotechnological applications. Emulsification studies revealed substrate-specific efficiency, with certain isolates showing superior activity against hydrocarbons and vegetable oils. All isolates exhibited β-hemolytic activity, supporting their biosurfactant-producing capability. Antimicrobial activity was selective, with BS2 and BS5 showing comparatively higher inhibition against bacterial strains, particularly Bacillus and Pseudomonas, while no activity was observed against Staphylococcus or fungal strains.
In the oil spreading assay, BS1 demonstrated the highest displacement, closely comparable to the positive control, indicating effective surface activity. Notably, antibiofilm analysis identified BS4 as the most potent inhibitor, suggesting its relevance in controlling microbial biofilms. Thin Layer Chromatography (TLC) further confirmed the presence of diverse biosurfactant types, including glycolipids and lipopeptides, as indicated by varying Rf values.
Overall, BS1 and BS4 emerged as the most promising isolates, exhibiting strong emulsification and antibiofilm properties, respectively, while BS2 and BS5 showed significant antimicrobial potential. These findings highlight the functional diversity of biosurfactants and emphasize their suitability for targeted applications in environmental, industrial, and biomedical fields.
Acknowledgement
The authors sincerely thank the Principal and Management of Bhavan’s Vivekananda College of Science, Humanities, and Commerce for their continued support and encouragement to carry out this research project.
Funding Sources
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflict of Interest
The authors do not have any conflict of interest.
Data Availability Statement
This statement does not apply to this article.
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Clinical Trial Registration
This research does not involve any clinical trials.
Permission to reproduce material from other sources
Not Applicable.
Author Contributions
Farjeen Fatima: Work Execution, Data Analysis, and First draft of paper.
Aparna: Plan of work, Conceptualization, Supervision, Article drafting, Reviewing, and Editing.
References
- Gudiña EJ, Rangarajan V, Sen R, Rodrigues LR. Potential therapeutic applications of biosurfactants. Trends Pharmacol Sci. 2015;36(9):675-687.
- Santos DKF, Rufino RD, Luna JM, Santos VA, Sarubbo LA. Biosurfactants: multifunctional biomolecules of the 21st century. Int J Mol Sci. 2016;17(3):401.
CrossRef - Singh P, Cameotra SS. Biosurfactant production by microbes isolated from oil-contaminated soil. Environ Technol Innov. 2021;22:101490.
- Mulligan CN. Environmental applications for biosurfactants. Curr Opin Colloid Interface Sci. 2021;56:101521.
- Das P, Mukherjee S, Sen R. Biosurfactants: the green generation of specialty molecules for industrial applications. Trends Biotechnol. 2022;40(1):58-72.
- Sha Y, Wang Y, Lu Z. Antifungal lipopeptides produced by Bacillus spp.: a biocontrol perspective. Microbiol Res. 2023;267:127265.
- Zhang L, Sun C, Feng J. Isolation and antifungal activity of lipopeptides from Bacillus subtilis against common soil-borne phytopathogens. J Appl Microbiol. 2020;129(4):811-823.
- Chen CY, Baker SC, Darton RC. The application of FTIR spectroscopy in biosurfactant analysis. J Colloid Interface Sci. 2017;504:101-110.
CrossRef - Patel R, Mehta H, Jadhav S. Recent advances in biosurfactant research: genomics to applications. Bioresour Technol Rep. 2022;19:101136.
- Sarkar D, Dutta A. Metagenomic insights into pesticide-contaminated soils for biosurfactant-producing bacterial communities. Front Environ Sci. 2023;11:1142032.
- Lima RV, de Souza AF, Nascimento R. Role of biosurfactants in soil remediation and plant growth. Appl Soil Ecol. 2021;157:103759.
- Reddy MG, Reddy MN, Saigopal DVR, Mallaiah KV. Laboratory Experiments in Microbiology. Himalaya Publishing House; Accessed March 7, 2026.
- ABIS Online – Advanced Bacterial Identification Software. Accessed March 7, 2026.
- Nayarisseri A, Singh P, Singh SK. Screening, isolation and characterization of biosurfactant producing Bacillus subtilis strain ANSKLAB03. 2018;14(6):304.
CrossRef - Verma R, Agrawal M, Rani ADL. Screening of biosurfactant production in bacteria isolated from oil and pesticide contaminated soil of Ranchi district. J Sci Res. 2022;66(4):1-8.
CrossRef - da Silva AF, Banat IM, Giachini AJ, Robl D. Fungal biosurfactants: from nature to biotechnological product—bioprospection, production and potential applications. Bioprocess Biosyst Eng. 2021;44(10):2003-2024.
CrossRef - Walter V, Rodrigues L, Teixeira J, Oliveira R. Oil spreading technique for biosurfactant screening: recent advances and applications. J Biotechnol. 2023;375:65-72.
- Ibrahim HMM. Characterization of biosurfactants produced by novel strains of Ochrobactrumanthropi HM-1 and Citrobacter freundii HM-2 from used engine oil-contaminated soil. Egypt J Pet. 2018;27(1):21-29.
CrossRef - Giri SS, Sen SS, Jun JW, Sukumaran V, Park SC. Antibacterial activity of biosurfactant produced by Bacillus spp. and its application in aquaculture. 2021;535:736336.
- Kavitha D, Anand T, Prabhakaran R. Screening and evaluation of antifungal activity of biosurfactant producing bacteria. J Mycol Plant Pathol. 2016;46(2):194-199.
- Reis RS, Pacheco GJ, Pereira AG, Freire DMG. Biosurfactants: production and applications. In: Chamy R, Rosenkranz F, eds. Biodegradation – Life of Science.IntechOpen; 2013.
CrossRef - Banat IM, Makkar RS, Cameotra SS. Potential commercial applications of microbial surfactants. Appl MicrobiolBiotechnol. 2000;53(5):495-508.
CrossRef - Mulligan CN. Environmental applications for biosurfactants. Environ Pollut. 2005;133(2):183-198.
CrossRef - Carrillo PG, Mardaraz C, Pitta-Alvarez SI, Giulietti AM. Isolation and selection of biosurfactant-producing bacteria. World J Microbiol Biotechnol. 1996;12(1):82-84.
CrossRef - Satpute SK, Banpurkar AG, Dhakephalkar PK, Banat IM, Chopade BA. Methods for investigating biosurfactants and bioemulsifiers: a review. Crit Rev Biotechnol. 2010;30(2):127-144.
CrossRef - Joshi S, Bharucha C, Jha S, Yadav S, Nerurkar A, Desai AJ. Biosurfactant production using molasses and whey under thermophilic conditions. Bioresour Technol. 2008;99(1):195-199.
CrossRef - Makkar RS, Cameotra SS. An update on the use of unconventional substrates for biosurfactant production and their new applications. Appl MicrobiolBiotechnol. 2002;58(4):428-434.
CrossRef - Gudiña EJ, Teixeira JA, Rodrigues LR. Biosurfactants produced by marine microorganisms with therapeutic applications. Mar Drugs. 2015;13(7):4030-4050.
- Sharma D, Ansari MJ, Al-Ghamdi A, Adgaba N. Biosurfactants: types, production, and environmental applications. Environ Chem Lett. 2021;19(2):1177-1195.
Accepted on: 27-03-2026
Second Review by: Dr. Chittran Roy
Final Approval by: Dr. Eugene A. Silow









