Leaf Morphometric Studies in some species of the Cucurbitaceae Juss.
1Department of Botany, Maratha Vidya Prasarak Samaj’s, S.S.S.M. Arts, Science and Commerce College, Saikheda, Nashik, Maharashtra, India,
2Department of Botany, Kalwan Education Society’s, Art, Commerce and Science College, Kalwan, Nashik, Maharashtra, India.
Corresponding Author E-mail: kaleunipune@gmail.com
Download this article as:
ABSTRACT:Ten species of Cucurbitaceae Juss. were morphometrically analyzed based on their leaf characteristics using taxonomic analysis to provide information about the links between these types of plants. Numerical characteristics, including leaf length, petiole length, leaf breadth, and lamina length, are positively correlated with the resolved taxonomic relationships of different species within the same genus, according to taxonomic component analysis. A key factor in uniting the species within a genus is the principal component analysis results of five quantitative characters based on a similarity matrix, which demonstrate a significant correlation between leaf length and leaf breadth, leaf base nerve number, and the ratio of leaf lamina length to petiole length. However, they also significantly separate the species from one another. Morphometric features supported the Cucurbitaceae's current categorization. Following the extraction of the characters that significantly influenced the similarity between the chosen Cucurbitaceae species, it further displays the component matrix of Coccinia grandis (L.) Voigt, Ctenolepis garcinii (Burm.f.) C. B. Clarke, Cucumis melo L., Cucumis setosus Cogn., Diplocyclos palmatus (L.) C. Jeffrey, Momordica balsamina L., Momordica dioica roxb ex willd, Mukia maderaspatana (L.) Roem., Solena amplexicaulis (Lam.) Gandhi and Trichosanthes cucumerina L.
KEYWORDS:Cucurbitaceae; Leaf; Morphometric studies; Nashik district; Taxonomy
Introduction
The Cucurbitaceae Juss. is a significant family of flowering plants commonly known as the gourd or cucumber family. There are approximately 965 species and 100 genera from around the world.
The studied number of species is incorporated in the genera Coccinia, Ctenolepis, Cucumis, Diplocyclos, Momordica, Mukia, Solena, and Trichosanthes. India has a great diversity of Cucurbitaceae (gourd family) species, with over 94 identified species distributed across 31 genera.1
The numerical analysis and comparison of closely related species are revealed by morphometric research. 2,3 Morphometric analysis and principal component analysis can be used to separate and categorize closely related plant species in the examined Cucurbitaceae species using a similarity matrix. Because numerical taxonomy uses more and better characters, it has increased the quality of conventional taxonomy.4,5
Certain genera in the Cucurbitaceae family, including Coccinia, Ctenolepis, Cucumis, Diplocyclos, Momordica, Mukia, Solena, and Trichosanthes, have been the subject of similar studies. Leaf morphometric analysis is a statistical technique. Due to their therapeutic value, Cucurbitaceae species must be correctly identified. Therefore, morphometric investigations allow for the clear separation and grouping of plant species by numerical analysis and comparison of closely related taxa.6,7 Principal Component Analysis (PCA) and Cluster Analysis (CA) are two techniques that facilitate the Similarity Matrix. This kind of research has been done in the genus under review. A line ruler was used to measure the quantitative characteristics of the leaf, including its length, breadth, base nerve number, petiole length, and lamina length. were morphometrically analyzed. Five distinct quantitative features were used to support the Cucurbitaceae’s current categorization. All of these features made a substantial contribution to the delineation of the Cucurbitaceae. species under study. In subsequent research, we advise using this approach to do a thorough taxonomic analysis of the genus Cucurbitaceae. Principal component analysis in the Cucurbitaceae ten species with a similarity matrix (Table 2, 3). All things considered, the observed differences in leaf size, shape, venation, and lamina–petiole ratio offer helpful quantitative characteristics for differentiating Cucurbitaceae species and may have taxonomic and adaptive relevance.8,9 Overall, the PCA similarity structure suggests that leaf size–related traits cluster together, while lamina–petiole proportion contributes independently to variation among Cucurbitaceae species (Table 2, 3).
Materials and Methods
Study area
Some selected regions of Nashik district (Fig. 1, Table 1). Ten species of Cucurbitaceae were collected from the Nashik district (Fig. 2). The collected species were identified as per different Floras and other literature. 10–13
A straight ruler was used to measure the quantitative characteristics of the leaf, including its length, breadth, base nerve number, petiole length, and lamina length (Table 2). Measures of central tendency and values were used to process the twenty samples from each sample. The obtained mean values of each quantitative character were subjected to principal component analysis and cluster analysis. Principal Component Analysis (PCA) and Cluster Analysis (CA) are two techniques that facilitate the Similarity Matrix.6,7
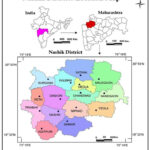 |
Figure 1: Study area- Nashik district
|
Table 1: Cucurbitaceae species studied from Nashik district, Maharashtra (India)
| Sr. No. | Species Name | Geographic coordinates | Location |
| 1. | Coccinia grandis ( L .) Voigt | 19.998333N, 73.938211E
19.953953N, 73.588881E 19.946355N, 73.533885E |
Chehadi Kh., Nashik
Anjaneri, Nashik Trimbak, Nashik |
| 2. | Ctenolepis garcinii (Burm.f.) C.B. Clarke | 20.037171N, 73.617537E | Shivangaon, Nashik |
| 3. | Cucumis melo L. | 20.038518N, 74.036701E
20.115918N, 73.761373E |
Shingave, Nashik
Ramshej, Nashik |
| 4. | Cucumis setosus Cogn. | 19.954333N, 73.663609E
19.911559N, 73.552948E 19.894479N, 73.516922E |
Maharavni, Nashik
Bhilmal, Nashik Dhadoshi, Nashik |
| 5. | Diplocyclos palmatus ( L .) C. Jeffrey | 19.954179N, 73.666506E
19.945142N, 73.551550E 19.887548N, 73.937324E 20.018823N, 74.015637E |
Maharavni, Nashik
Trimbak, Nashik Sinnar, Nashik Saikheda, Nashik |
| 6. | Momordica balsamina L. | 20.465704N, 74.066162E | Kalwan, Nashik |
| 7. | Momordica dioica roxb ex willd | 19.955480N, 73.663603E
19.897661N, 73.559149E 19.947809N, 73.570105E |
Maharavni, Nashik
Pahine, Nashik Pegalwadi, Nashik |
| 8. | Mukia maderaspatana ( L.) Roem. | 19.953574N, 73.666004E
19.930402N, 73.560467E |
Maharavni, Nashik
Trimbak, Nashik |
| 9. | Solena amplexicaulis ( Lam.) Gandhi | 19.912333N, 73.559200E
19.898341N, 73.532301E |
Bhilmal, Pahine, Nashik
Kojoli, Nashik |
| 10. | Trichosanthes cucumerina L. | 20.070255N, 73.675141E
20.000658N, 73.926040E 20.037567N, 73.717885E |
Girnare, Nashik
Lakhalgaon, Nashik Gangapur gaon, Nashik |
Results
Ten Cucurbitaceae species, including Coccinia grandis (L.) Voigt, Ctenolepis garcinii (Burm.f.) C. B. Clarke, Cucumis melo L., Cucumis setosus Cogn., Diplocyclos palmatus (L.) C. Jeffrey, Momordica balsamina L., Momordica dioica roxb ex willd, Mukia maderaspatana (L.) Roem., Solena amplexicaulis (Lam.) Gandhi, and Trichosanthes cucumerina L. The current classification of the ten Cucurbitaceae was supported by five different leaf morphometric characteristics (Fig. 2). Five-leaf quantitative characteristics of ten Cucurbitaceae species, including leaf length, leaf breadth, and the ratio of leaf length to leaf breadth, significantly contributed in the delimitation of the Cucurbitaceae species under study (Table 2).
Ten Cucurbitaceae species from the Nashik district, including the talukas of the Nashik district, are the subject of this study (Table 1). Significant differences were found across the ten Cucurbitaceae species (I1–I10) based on quantitative examination of leaf features (Table 2). There were significant interspecific variations in leaf size, with leaf length (LL) ranging from 3.9 cm (I10) to 12.4 cm (I8). Leaf breadth (LB) also varied, ranging from 3.52 cm (I5) to 9.41 cm (I8). The biggest leaf dimensions were consistently displayed by species I8, and the smallest values for both length and breadth were shown by species I10.
The leaf length to leaf width (LL/LB) ratio varied from 1.12 (I2) to 1.83 (I4), indicating a shift in leaf morphology from comparatively broad to more elongated forms. While the leaf outlines of species I1, I2, and I10 were more rounded, those of species I4 and I7 were somewhat extended. With values of 3 or 5, the leaf base nerve number (LBNN) displayed little change, suggesting a consistent venation pattern among the species. The remaining species had three basal nerves, whereas species I1, I3, I7, and I9 had five. The range of leaf lamina length (LLL) was 1.91 cm (I10) to 6.6 cm (I8). Petiole length (PL) varied greatly, with I7 having the smallest petiole (0.67 cm) and I8 having the longest (5.87 cm). As a result, there was significant variation in the ratio of leaf lamina length to petiole length (LLL/PL), which ranged from 1.09 (I10) to 9.45 (I7). Because of its extremely short petiole, species I7 had a noticeably high LLL/PL ratio, whereas species I5 also had a comparatively high ratio (4.25), suggesting a proportionately larger lamina.
The similarity matrix revealed strong positive correlations among most leaf quantitative characters of the studied Cucurbitaceae species. Leaf length (LL) showed very high correlation with leaf breadth (LB) (r = 0.99) and LL/LB ratio (r = 0.98), indicating that overall leaf size strongly influences leaf shape. Leaf breadth also correlated highly with petiole length (PL) (r = 0.92).
 |
Figure 2: Selected Cucurbitaceae species leaf morphology
|
Leaf base nerve number (LBNN) exhibited strong association with petiole length (r = 0.99) and moderate to high correlations with LL, LB, and LL/LB, suggesting coordinated variation between venation and leaf size traits. Leaf lamina length (LLL) showed moderate correlations with LL, LB, and PL, but a very high correlation with the LLL/PL ratio (r = 0.91), indicating that lamina length largely determines this ratio.
Table 2: Leaf quantitative characters of Cucurbitaceae species (Mean values in cm).
| I1 | I2 | I3 | I4 | I5 | I6 | I7 | I8 | I9 | I10 | |
| LL | 7.78 | 4.98 | 6.52 | 8.03 | 4.76 | 8.09 | 6.7 | 12.4 | 9.22 | 3.9 |
| LB | 6.46 | 4.49 | 5.03 | 4.45 | 3.52 | 5.27 | 4.3 | 9.41 | 6.68 | 3.8 |
| LL/LB | 1.18 | 1.12 | 1.28 | 1.83 | 1.34 | 1.55 | 1.69 | 1.31 | 1.37 | 1.15 |
| LBNN | 5 | 3 | 5 | 3 | 3 | 3 | 5 | 3 | 5 | 3 |
| LLL | 4.95 | 3.03 | 4.19 | 5.27 | 3.61 | 5.09 | 6.03 | 6.6 | 4.79 | 1.91 |
| PL | 2.82 | 1.95 | 2.33 | 2.76 | 1.01 | 3 | 0.67 | 5.87 | 4.43 | 1.99 |
| LLL/PL | 2.38 | 1.6 | 2.47 | 2.19 | 4.25 | 2.34 | 9.45 | 1.22 | 1.32 | 1.09 |
Abbreviations: LL: Leaf length; LB: leaf breadth; LL/LB: ratio of leaf length to leaf breadth; LBNN: leaf base nerve number; PL: petiole length; LLL: leaf lamina length; LLL/PL: ratio of leaf lamina length to petiole length.
I1: Coccinia grandis (L.) Voigt; I2: Ctenolepis garcinii (Burm.f.) C. B. Clarke; I3: Cucumis melo L.; I4: Cucumis setosus Cogn.; I5: Diplocyclos palmatus (L.) C. Jeffrey; I6: Momordica balsamina L., I7: Momordica dioica roxb ex willd; I8: Mukia maderaspatana (L.) Roem.; I9: Solena amplexicaulis (Lam.) Gandhi and I10: Trichosanthes cucumerina L.
Table 3: Principal component analysis in studied Cucurbitaceae species with Similarity Matrix.
| LL | LB | LL/LB | LBNN | LLL | PL | LLL/PL | |
| LL | 1 | 0.989535 | 0.981058 | 0.883728 | 0.667898 | 0.922456 | 0.315518 |
| LB | 0.989535 | 1 | 0.95025 | 0.87114 | 0.633455 | 0.922441 | 0.259168 |
| LL/LB | 0.981058 | 0.95025 | 1 | 0.842721 | 0.703153 | 0.867406 | 0.388662 |
| LBNN | 0.883728 | 0.87114 | 0.842721 | 1 | 0.645628 | 0.987279 | 0.313056 |
| LLL | 0.667898 | 0.633455 | 0.703153 | 0.645628 | 1 | 0.655343 | 0.906622 |
| PL | 0.922456 | 0.922441 | 0.867406 | 0.987279 | 0.655343 | 1 | 0.304805 |
| LLL/PL | 0.315518 | 0.259168 | 0.388662 | 0.313056 | 0.906622 | 0.304805 | 1 |
Abbreviations: LL: Leaf length; LB: leaf breadth; LL/LB: ratio of leaf length to leaf breadth; LBNN: leaf base nerve number; LLL: leaf lamina length; PL: petiole length; LLL/PL: ratio of leaf lamina length to petiole length.
Discussion
Principal component analysis results of five quantitative characters based on a similarity matrix show significant correlations between leaf length, leaf width, ratio of leaf length to leaf breadth, leaf base nerve number, petiole length, leaf lamina length, and ratio of leaf lamina length to petiole length (Table 2, 3). About 31 Ficus species have been subjected to the two numerical taxonomy methods (principal component analysis with similarity matrix and cluster analysis) employed in this study.14 Ten Cucurbitaceae species were employed in the study for principal component analysis using a similarity matrix. Significant differences were found across the ten Cucurbitaceae species (I1–I10) based on quantitative examination of leaf features to be very useful for species identification within the family or genus (Table 2). Five Terminalia species and nine Ficus species underwent morphometric and phytochemical analyses using the hierarchical classification and visual interpretation of the taxonomic relationship in the current traditional classification of the genera. This study used the same approach as the Terminalia genus, here Cucurbitaceae studied member easily identify or segregate out different genera within the same family. 6,7
The outcomes of these taxonomical processes are often used as objective indicators of the similarities or differences between the taxa of ten species of Cucurbitaceae in order to arrange taxa in the hierarchical order for plant categorization.15 For plant reproductive organs, flowers or inflorescences are typically required for species identification or classification.16 The 10 species of the Cucurbitaceae family can be distinguished from one another using the leaf morphometry features utilized in this study. The leaf morphometry characteristics that significantly aided in distinguishing between members of the same genus and distinct species have been brought to light by this work. Nevertheless, “numerical taxonomy” and numerical techniques were used throughout the investigation.17
Conclusion
The findings validated the Cucurbitaceae’s present classification. Quantitative leaf features varied significantly across the ten species (Table 2). While leaf length (LL) varied from 3.9 cm (I10) to 12.4 cm (I8), leaf breadth (LB) ranged from 3.52 cm in species I5 to 9.41 cm in species I8. The largest leaves were found in species I8, while the smallest were found in species I10. The leaf length-to-breadth ratio (LL/LB) varied from 1.12 (I2) to 1.83 (I4), indicating a shift in leaf morphology from broader to more elongated forms. The leaf base nerve number (LBNN), which ranges from three to five, varied very little between species. While leaf lamina length (LLL) varied from 1.91 cm (I10) to 6.6 cm (I8), petiole length (PL) varied significantly, ranging from 0.67 cm in species I7 to 5.87 cm in species I8. Species I7 had the largest lamina length–to–petiole length ratio (LLL/PL) because to its small petiole. Overall, the LLL/PL ratio varied greatly (1.09–9.45) and exhibited minimal associations with the majority of size-related characteristics (LL, LB, and PL), suggesting that it is a largely independent character.
The lamina-petiole proportion contributed independently to variation among the Cucurbitaceae species, while leaf size-related variables clustered together, according to principal component analysis (PCA). These results show how quantitative leaf characteristics can be used taxonomically to differentiate Cucurbitaceae species. The ten Cucurbitaceae species were successfully identified, and their current taxonomy was supported by quantitative leaf morphometric features. The lamina–petiole ratio generated independent variance, underscoring its taxonomic relevance, while leaf size–related features demonstrated substantial grouping.
Acknowledgement
I am extremely grateful to Principal Prof. (Dr.) K. M. Khalkar and HOD Department of Botany, Dr. S. V. Gosavi, Maratha Vidya Prasarak Samaj’s, S.S.S.M. Arts, Science and Commerce College, Saikheda, Nashik, Maharashtra, India, 422210, for providing me with all the necessary facilities for the completion of this research.
Funding Sources
The authors received no financial support for the research, authorship, and/or publication of this article.
Conflict of Interest
The authors do not have any conflicts of interest.
Data Availability Statement
This statement does not apply to this article.
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Clinical Trial Registration
This research does not involve any clinical trials.
Permission to reproduce material from other sources
Not Applicable
Author Contributions
Suresh Ganpat Sabale- Data collection
Roshani Manik Shinde -Analyzed the data and served as the major advisor.
Suresh Ganpat Sabale- Wrote the research article (Introduction, Materials & Methods, Tables, Results, and Discussion) and aided as co-advisor.
Balasaheb Shantilal Kale- All possible corrections before and after sending the manuscripts.
References
- Renner, S. S.; Pandey, A. K. The Cucurbitaceae of India : Accepted Names , Synonyms , Geographic Distribution , and Information on Images and DNA Sequences. PhytoKeys 2013, 20 (1), 53–118. https://doi.org/10.3897/phytokeys.20.3948.
CrossRef - Bocxlaer, B. Van; Schultheiß, R. Comparison of Morphometric Techniques for Shapes with Few Homologous Landmarks Based on Machine-Learning Approaches to Biological Discrimination. Paleobiology 2010, 36 (3), 497–515.
CrossRef - Dinda, S.; Mondal, A. K. The Morphometric and Numerical Analysis of Five Species of Eragrostis Sp . Wolf . Based on Silica Bodies in Leaf Epidermal Cells. Plant Sci. 2018, 7 (5), 2213–2219.
CrossRef - Oso, O. A.; Jayeola, A. A. Digital Morphometrics : Application of Morpho Leaf in Shape Visualization and Species Delimitation , Using Cucurbitaceae Leaves as a Model. Plant Sci. 2021, 9 (e11448), 1–14. https://doi.org/10.1002/aps3.11448.
CrossRef - Hiremath, M. M.; Kotresha, K.; Makanur, N. S. Leaf Venation and Morphometric Studies in Some Members of Family Loranthaceae. J. Trend Sci. Res. Dev. 2025, 9 (2), 1011–1016.
- Deshmukh, S. A.; Waghmare, M. B.; Labhane, N. M.; Gaikwad, D. K. Leaf Morphometric Studies in The Genus Terminalia L . from Kolhapur District. Rev. J. Bot. 2013, 2 (2), 1–3.
- Jangam, A. P.; Jadhav, S. P.; Sutar, V. D.; Onkar, R. R. Leaf Morphometric Studies in Some Species of Ficus L . Rev. J. Bot. 2017, 6 (2), 29–31.
- Fatima, S.; Mahajan, M.; Protection, P. Morphometric Study of Genus Hibiscus from Family Malvaceae. IJRTI 2020, 5 (1), 84–89.
- Ángel, O.; Bonilla, D. L.; Valencia, S.; Ibarra, Á. G.; Morales, S.; Efraín, S.; Sánchez, T.; González, A. Leaf Morphometric Analysis and Potential Distribution Modelling Contribute to Taxonomic Differentiation in the Quercus Microphylla Complex. Plant Res. 2024, 137 (1), 3–19. https://doi.org/10.1007/s10265-023-01495-z.
CrossRef - Lakshminarasimhan, P.; Sharma, B. D. Flora of Nasik District (BSI); 1991.
- Almedia, M. R. Flora of Maharashtra. Thomas Paul Almeida Blatter Herb. St. Xavier’s Coll. 1996, 1.
- Singh, N. P.; Lakshminarasimhan, P.; Karthikeyan, S.; Prasanna, P. V. Flora of Maharashtra State, Dicotyledons. Flora India Ser. 2 2000, 1 (1), 1–871.
- Singh, N. P.; Lakshminarasimhan, P.; Karthikeyan, S.; Prasanna, P. V. Flora of Maharashtra State: Dicotyledones. Flora India Ser. 2 2001, 2 (1), 1–1096.
- Sonibare, M. A.; Jayeola, A. A.; Egunyomi, A. A Morphometric Analysis of the Genus Ficus (Moraceae). African J. Biotechnol. 2004, 3 (4), 229–235. https://doi.org/10.5897/ajb2004.000-2043.
CrossRef - Agbagwa, I. O.; Okoli, B. E. Fruit Epidermal Micromorphology in the Systematics of Abrus Adanson (Papilionaceae) in Parts of Tropical West Africa. Asian J. Plant Sci. 2005, 4 (6), 652–659.
CrossRef - Singh, G. Plant Systematics. New Hampsh. 2010, 3 (1), 1–717.
CrossRef - Soladoye, M. O.; Onakoya, M. A.; Chukwuma, E. C.; Sonibare, M. A. Morphometric Study of the Genus Senna in South-Western Nigeria. African J. Plant Sci. 2010, 4 (3), 44–52.
Abbreviations list
PCA: Principal Component Analysis
CA: Cluster Analysis
Accepted on: 30-01-2026
Second Review by: Dr. Jayath Kirthisinghe
Final Approval by: Dr. Eugene A. Silow









