Sustainable Production of Glycolipid Biosurfactants by Halophilic Bacteria Utilizing Waste Vegetable Oil: Insights into Antimicrobial and Antiadhesive Properties
Department of Biotechnology, Institute of Science and Technology for Advanced Studies and Research – ISTAR, The Charutar Vidya Mandal (CVM) University, Vallabh Vidyanagar, Gujarat, India.
Corresponding author Email: nisha.daxini@cvmu.edu.in
DOI : http://dx.doi.org/10.13005/bbra/3460
ABSTRACT:Biosurfactants from halophilic bacteria represent a promising sustainable alternative to synthetic surfactants, particularly when produced from waste substrates. This study investigated biosurfactant production by halophilic bacteria isolated from coastal marine waters using waste vegetable oil as a carbon source. Six halophilic bacterial isolates were obtained from seawater samples collected from Dhuvaran, Dandi, Dwarka, and Daman coastal regions of India. Among these, isolates Dh4 and Dm2 exhibited superior biosurfactant production, achieving yields of 2.3 g/100 mL and 2.1 g/100 mL, respectively, in a mineral salt medium supplemented with 2% waste vegetable oil. The produced biosurfactants demonstrated significant surface-active properties, reducing surface tension and showing oil displacement zones of 7.5 cm and 7.8 cm for Dh4 and Dm2, respectively. Emulsification indices (E24) reached 42% and 47%, indicating strong emulsifying capacity. Preliminary biochemical characterisation revealed the presence of glycolipid biosurfactants, as confirmed by saponification and Fehling's tests. FTIR spectroscopy identified characteristic functional groups including C-H stretching (2923.57, 2853.24 cm⁻¹), C=O stretching (1714.39 cm⁻¹), and C=C stretching (1636.88 cm⁻¹), consistent with glycolipid structures. The biosurfactants exhibited potent antiadhesive activity against pathogenic bacteria, inhibiting biofilm formation by Bacillus subtilis (75.33%), Escherichia coli (39.3%), Pseudomonas aeruginosa (28.26%), and Salmonella paratyphi (13.5%) at a concentration of 5 mg/mL. Antimicrobial activity against these pathogens was also demonstrated. These findings highlight the potential of halophilic bacteria for sustainable biosurfactant production from waste oily substrates, offering promising applications in biomedical and environmental fields, particularly for combating biofilm-associated infections.
KEYWORDS:Antiadhesive Activity; Biofilm Inhibition; Biosurfactant; Glycolipid; Halophilic Bacteria; Sustainable Production; Waste Vegetable Oil
Introduction
Biosurfactants are amphiphilic molecules produced by microorganisms that possess both hydrophilic and hydrophobic moieties, enabling them to reduce surface and interfacial tension between liquids, gases, and solids.¹These naturally occurring compounds have garnered significant attention as eco-friendly alternatives to synthetic surfactants due to their biodegradability, low toxicity, high selectivity, and effectiveness under extreme environmental conditions.² Unlike petroleum-derived chemical surfactants, biosurfactants can be synthesised from renewable feedstocks, offering substantial environmental and economic benefits.³
Bacterial biofilms pose a critical challenge in both clinical and industrial settings, with approximately 40–80% of bacteria on Earth forming biofilms.⁴ These complex microbial communities are implicated in numerous chronic infections, including cystic fibrosis caused by Pseudomonas aeruginosa, periodontitis involving Porphyromonas gingivalis, and nosocomial infections associated with Enterococcus species.⁵˒⁶ Within bacterial biofilms, approximately 1% of cells exhibit antibiotic resistance, and biofilm-associated infections account for 95% of urinary tract infections and 87% of bloodstream infections related to medical devices.⁷ This alarming prevalence underscores the urgent need for novel antibiofilm strategies.
Biosurfactants have emerged as promising antibiofilm agents due to their ability to prevent microbial adhesion to surfaces, interfere with quorum-sensing signalling, and disrupt established biofilm structures.⁸˒⁹ Glycolipids such as rhamnolipids and sophorolipids, as well as lipopeptides like surfactin and iturin, have demonstrated significant antiadhesive and antimicrobial activities against various pathogens.¹⁰˒¹¹ The mechanisms underlying these activities include alterations in cell surface hydrophobicity, interference with bacterial communication systems, and disruption of biofilm matrix components.¹²
Halophilic bacteria, which thrive in high-salt environments, represent an underexplored resource for biosurfactant production.¹³ These extremophiles possess unique metabolic capabilities and can produce biosurfactants that maintain stability under harsh conditions of salinity, pH, and temperature.¹⁴ Furthermore, the utilization of waste substrates for biosurfactant production addresses both environmental pollution and production cost challenges. Waste cooking oils and used vegetable oils, which pose significant disposal problems, have been successfully employed as carbon sources for biosurfactant production, achieving concentrations of up to 67 g/L with various microbial species.¹⁵˒¹⁶
Despite growing interest in biosurfactant research, the exploration of halophilic bacteria from coastal marine environments for biosurfactant production using waste-oily substrates remains limited. The present study aimed to isolate and screen biosurfactant-producing halophilic bacteria from the coastal waters of Gujarat, India; optimize biosurfactant production using waste vegetable oil as a sustainable carbon source; characterize the biosurfactant through biochemical and spectroscopic analyses; and evaluate its antiadhesive, antibiofilm, and antimicrobial activities against clinically relevant pathogens. The findings contribute to the development of sustainable bioprocessing strategies and offer potential applications in combating antibiotic-resistant biofilm infections.
Materials and Methods
Chemicals and Instruments Used
Chemicals and Reagents
All chemicals and reagents used in this study were of analytical grade. Culture media components, including Zobell Marine Broth (ZMB) and Mineral Salt Medium (MSM) were procured from HiMedia Laboratories Pvt. Ltd., India. Organic solvents such as chloroform and methanol (HPLC grade) were obtained from Loba Chemie Pvt.Ltd Standard rhamnolipid used as a reference biosurfactant was purchased from Sigma-Aldrich (USA). Diesel oil was obtained from local automobile service garages in Anand, Gujarat, and used vegetable oil (palm oil) was collected from local vendors.
Instrumentation
Key laboratory instruments used in this study included a UV–Vis spectrophotometer (Shimadzu UV-1800) for OD measurements, a refrigerated centrifuge (REMI C-24BL) for cell separation, a rotary evaporator (Buchi R-215) for solvent evaporation, an FTIR spectrophotometer (PerkinElmer Spectrum Two), a GC–MS system (PerkinElmer Autosystem XL), and NMR and ESI–MS instruments available at SAIF Chandigarh. An orbital shaker incubator (REMI) and a laminar airflow cabinet (Air pac) were used for routine microbiological and culture handling procedures.
Sample Collection and Isolation of Halophilic Bacteria
Seawater samples were collected from four coastal locations in India: Dhuvaran (22.2333° N, 72.7621° E), Dandi (20.8866° N, 72.7971° E), Dwarka (22.2378° N, 68.9627° E), and Daman (20.4090° N, 72.8300° E) during November 2023 to April 2024. Samples were collected in sterile glass bottles and immediately transported to the laboratory for processing. Serial dilutions (10⁻², 10⁻⁴, 10⁻⁶, 10⁻⁸) of water samples were prepared in sterile saline and spread-plated onto Zobell Marine Broth (ZMB) agar medium containing varying NaCl concentrations (5%, 10%, 15%, 20%, 25%). The ZMB medium composition included peptone (5.0 g/L), yeast extract (1.0 g/L), ferric citrate (0.1 g/L), magnesium chloride (8.8 g/L), sodium sulfate (3.24 g/L), calcium chloride (1.8 g/L), potassium chloride (0.55 g/L), sodium bicarbonate (0.16 g/L), and other trace minerals, with pH adjusted to 7.4–7.8. Plates were incubated at 37°C for 1–2 days, and morphologically distinct colonies were isolated and purified through repeated subculturing.
Screening for Biosurfactant Production
Oil Displacement Assay
Biosurfactant production was initially screened using the oil displacement method.¹⁷ Briefly, 20 mL of distilled water was added to a Petri dish, followed by 100 μL of diesel oil, which formed a thin oil layer on the water surface. Then, 100 μL of cell-free culture supernatant was gently placed on the oil surface. The diameter of the clear zone formed due to oil displacement was measured and recorded as an indicator of biosurfactant activity.
Emulsification Index (E24)
The emulsification index was determined according to the method described by Bodour et al.¹⁸. Equal volumes (4 mL) of cell-free culture broth and hydrocarbon substrate (diesel oil or vegetable oil) were mixed by vortexing for 2 minutes. The mixture was allowed to stand at room temperature for 24 hours. The E₂₄ value was calculated using the formula:
E24 (%) = (Height of emulsified layer / Total height of mixture) × 100
Biosurfactant Production
Well-isolated bacterial colonies were inoculated into 250 mL Erlenmeyer flasks containing 100 mL of Mineral Salt Medium (MSM) supplemented with different carbon sources, following previously established protocols (Datta et al., 2018; Radzuan et al., 2017). The MSM contained NaCl (0.5 g/L), KH₂PO₄ (3.0 g/L), MgSO₄·7H₂O (0.12 g/L), CaCl₂·2H₂O (0.013 g/L), NaNO₃ (15 g/L), K₂HPO₄ (4.4 g/L), and yeast extract (3.0 g/L). The medium was supplemented with various carbon sources to evaluate their effect on biosurfactant synthesis, including 2% waste vegetable oil, 2% diesel oil, 1% coconut oil, and engine oil. Cultures were incubated at 37 °C on a rotary shaker at 120 rpm for 5–7 days to study the growth profile and biosurfactant production kinetics.
Daily monitoring was performed using oil displacement assays to determine the onset and progression of biosurfactant production.
Biosurfactant Extraction and Quantification
Although biosurfactant production was monitored for a total duration of 5–7 days, the oil-displacement assay—which was the primary method used to quantify biosurfactant activity—showed that the highest activity occurred at 72 h. The displacement diameter increased progressively from 24 h to 72 h, after which no significant improvement was observed up to day 7. Thus, while the incubation period was extended to study the overall production kinetics, biosurfactant extraction was performed specifically at 72 h, corresponding to the peak oil-displacement activity and therefore the maximal biosurfactant yield.
At 72 h, the culture broth was centrifuged at 8,000 rpm for 15 minutes to obtain the cell-free supernatant. The supernatant was acidified to pH 2.0 using 6 N HCl and kept at 4 °C for 12 hours to allow biosurfactant precipitation. The precipitate was then extracted using a chloroform:methanol (2:1, v/v) solvent system, and the organic phase was separated using a separating funnel. Following solvent evaporation in a rotary evaporator, the biosurfactant was obtained as a honey-coloured extract and quantified gravimetrically.
Biochemical Characterization
Saponification Test
For the saponification test, 5 mL of dissolved biosurfactant was mixed with 2 mL of 2% NaOH and heated in a water bath at 60–70°C for 30 minutes. Formation of a gelatinous or opaque soap layer upon cooling indicated positive saponification.²¹
Fehling’s Test
Fehling’s reagents A and B were mixed in equal volumes and added to 2–3 mL of the biosurfactant solution. The mixture was heated in a boiling water bath for 5 minutes. Formation of a brick-red precipitate of copper(I) oxide indicated the presence of reducing sugars.²²
Ninhydrin Test
Biosurfactant samples were mixed with 0.2% ninhydrin reagent solution and heated in a boiling water bath for 5 minutes. The appearance of purple or blue color indicated the presence of amino acids or proteins.²³
Fourier Transform Infrared Spectroscopy (FTIR)
Functional groups in the crude biosurfactant were identified using FTIR spectroscopy (SICART, Vallabh Vidyanagar). Approximately 2 mg of dried biosurfactant was analyzed in the spectral range of 400–4000 cm⁻¹ using potassium bromide (KBr) pellets.
Antiadhesive and Antibiofilm Assays
Slide Test
Glass slides were coated with 1.5 mL of crude biosurfactant (0.5 to 50 μg/mL) in 6-well plates and incubated at 4°C for 18 hours, followed by washing twice with phosphate-buffered saline (PBS). Control slides contained PBS only. Aliquots of 1.5 mL of bacterial suspension, Pseudomonas aeruginosa MTCC 3457, Escherichia coli ATCC 23716, Bacillus subtilis MTCC 441, Salmonella paratyphi clinical strain were added to wells and incubated at 37°C for 24 hours. Glass slides were removed, rinsed with PBS, stained with 2% crystal violet for 5 minutes, washed, air-dried, and photographed under an optical microscope.
Microtiter Plate Method
Wells of sterile 96-well microtiter plates were filled with 100 μL of biosurfactant solution (0.5-5 mg/mL) and incubated at 37°C for 24 hours. After removal and washing with sterile distilled water, 200 μL of pathogen culture was added to each well and incubated for 24 hours. Wells were washed, stained with 0.1% crystal violet for 20 minutes, and destained with methanol. Glacial acetic acid in water (33%) was added, and absorbance was measured at 595 nm. Percentage inhibition of microbial adhesion was calculated as:
Microbial adhesion inhibition (%) = [(OD control – OD sample) / OD control] × 100
Antimicrobial Activity
Antimicrobial activity was assessed using the agar well diffusion assay²⁴. Pathogenic strains Pseudomonas aeruginosa MTCC 3457, Escherichia coli ATCC 23716, Bacillus subtilis MTCC 441, S. paratyphi clinical strain were incorporated into nutrient broth agar (1%, v/v) plates. Crude biosurfactants were pipetted into wells drilled in the agar, with rhamnose as a control. Plates were incubated aerobically at 37°C for 24 hours, and zones of inhibition were measured.
Statistical Analysis
All experiments were carried out in triplicate, and the results are expressed as mean ± standard deviation. Statistical analysis was performed using ANOVA (Analysis of Variance) to determine the significance of differences among treatments. A p-value < 0.05 was considered statistically significant.
Results
Isolation and Characterization of Halophilic Bacteria
Six morphologically distinct bacterial isolates (Dh3, Dh4, Da2, Dm2, Dm3, Dw2) were successfully isolated from marine water samples collected from four coastal locations. The isolates exhibited diverse colony characteristics including varied pigmentation (yellow, orange, off-white, creamy white), colony morphology (round, pin-point), and texture (viscous, smooth). Gram staining revealed both Gram-positive and Gram-negative bacteria. All isolates demonstrated growth on ZMB medium with varying salt concentrations, confirming their halophilic nature.
Table 1: Colony Characteristics of Isolates
| Colony characteristics | Dh3 | Dh4 | Da2 | Dm2 | Dm3 | Dw2 |
| Size | Small | Small | Small | Small | Small | Small |
| Shape | Round | Pin point | Pin point | Pin point | Pin point | Pin point |
| pigmentation | Orange | Yellow | Off white | Off white | Creamy White | White |
| Opacity | Opaque | Opaque | Opaque | Transparent | Opaque | Transparent |
| Gram staining | Positive rods | Negative cocci | Negative rods | Positive cocci | Positive cocci | Positive rods |
| Elevation | Raised | Flat | Raised | Flat | Flat | Raised |
| Texture | Viscous | Viscous | Viscous | Viscous | Smooth | Viscous |
| Margin | Entire | Entire | Entire | Entire | Entire | Entire |
Screening for Biosurfactant Production
Initial screening using oil-displacement assays revealed that isolates Dm2 and Dh4 exhibited the highest biosurfactant production, with oil-displacement zones of 7.8 cm and 7.5 cm, respectively. The emulsification indices (E24) for these isolates were 47% and 42%, respectively, significantly higher than those of other isolates (Table 1). These values compared favourably with the rhamnolipid standard (93% E24, 7.9 cm oil displacement), indicating substantial biosurfactant production potential.
Table 2: Emulsification index and oil displacement of bacterial isolates
| Isolate | Emulsification Index (E24) | Oil Displacement (cm) |
| Dh4 | 42% | 7.5 |
| Dm2 | 47% | 7.8 |
| Rhamnolipid (standard) | 93% | 8.2 |
 |
Figure 2: a) Emulsification Index test. b) Oil displacement test for isolate Dm2 |
Biosurfactant Production Kinetics
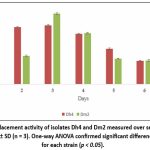
Time-course studies showed that biosurfactant production by both Dh4 and Dm2 isolates reached maximum levels on day 3 of incubation, with oil-displacement distances of 7.5 cm and 7.9 cm, respectively. Production subsequently declined after day 3, possibly due to nutrient depletion or Biosurfactant degradation. This growth-associated production pattern suggests that biosurfactant synthesis is coupled to bacterial metabolism during the exponential growth phase, consistent with findings in other Pseudomonas and Bacillus species.²⁵˒²⁶
 |
Figure 3: Biosurfactant production medium. |
Biosurfactant Production Using Different Oily Waste Substrates
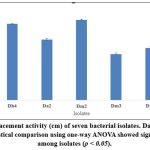
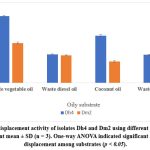
Both isolates were evaluated for biosurfactant production using various waste oily substrates. Dh4 demonstrated superior performance with waste vegetable oil (7.5 cm oil displacement), followed by diesel oil, engine oil, and coconut oil. Dm2 showed the highest activity with vegetable oil (4.2 cm) but lower activities with other substrates.
These results demonstrate the substrate specificity of biosurfactant production and highlight waste vegetable oil as the most suitable carbon source for both isolates, supporting sustainable waste valorization strategies.²⁷˒²⁸
Biosurfactant Extraction and Yield
Following acid precipitation and solvent extraction, biosurfactants were recovered as honey-colored viscous products. The maximum yields obtained were 2.3 g/L for Dh4 and 2.1 g/L for Dm2 when cultivated in MSM supplemented with 2% waste vegetable oil. These yields are comparable to those reported in other studies using halophilic bacteria and waste substrates, demonstrating the economic viability of this approach.²⁹˒³⁰
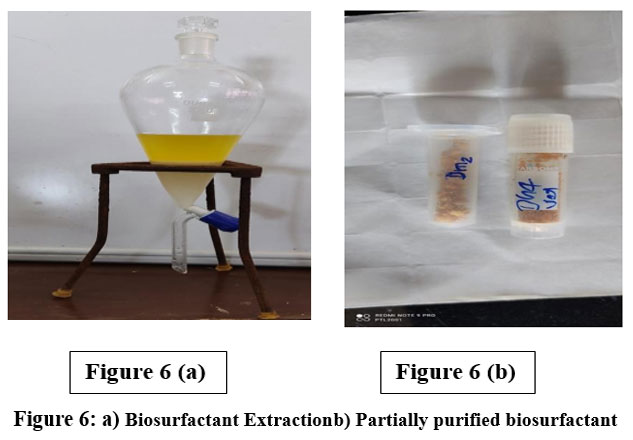 |
Figure 6: a) Biosurfactant Extraction, b) Partially purified biosurfactant. |
Biochemical Characterization
Preliminary biochemical tests revealed that both Dh4 and Dm2 biosurfactants tested positive for saponification and Fehling’s tests, indicating the presence of ester bonds and reducing sugars, respectively. The ninhydrin test was negative, suggesting minimal or no free amino acid content (Table 2). These results are consistent with glycolipid-type biosurfactants, such as rhamnolipids or sophorolipids, which are commonly produced by marine bacteria.³¹˒³²
 |
Figure 7: a) Fehling’s Test, b) Saponification Test. |
Table 3: Results of biochemical characterization tests
| Test | Dh4 | Dm2 | Interpretation |
| Saponification | Positive | Positive | Presence of ester bonds |
| Fehling’s | Positive | Positive | Presence of reducing sugars |
| Ninhydrin | Negative | Negative | Absence of free amino acids |
FTIR Spectroscopic Analysis
Fourier Transform Infrared (FTIR) spectroscopy of the crude biosurfactant produced by isolate Dh4 confirmed its glycolipid composition, exhibiting characteristic absorption bands consistent with rhamnolipid structures. The spectrum displayed major C–H stretching peaks at 2923.57 cm⁻¹ and 2853.24 cm⁻¹, corresponding to methylene (CH₂) and methyl (CH₃) groups of aliphatic fatty acid chains.³³ A strong C=O stretching peak at 1714.39 cm⁻¹ indicated ester linkages typical of glycerol-linked fatty acids, consistent with literature values for rhamnolipid biosurfactants.³4 The C=C band at 1636.88 cm⁻¹ reflected unsaturated fatty acids derived from waste vegetable oil, with these absorption signals characteristic of ester compounds in the hydrocarbon region.²¹ Critical identification bands included O–H vibration at 1343.38 cm⁻¹ representing hydroxyl groups from carbohydrate moieties, and C–O stretching at 1052.03 cm⁻¹ confirming glycosidic ether linkages between sugar and lipid moieties—the C–O–C absorption band is specifically assigned to the rhamnose molecule in biosurfactant structures.³⁵ The C–C bonding peak at 947.25 cm⁻¹ represented the aliphatic hydrocarbon backbone. A minor C–Cl band at 831.72 cm⁻¹ suggested trace chloroform residue from extraction procedures. Together, these peaks confirm a rhamnolipid-type glycolipid with rhamnose linked to hydroxydecanoic acids, matching literature values documented for Pseudomonas and Bacillus species-derived rhamnolipids.³¹˒³² FTIR spectroscopy is recognised as one of the best methods for determining functional groups and chemical bonds in unknown compound structures, enabling unambiguous identification of biosurfactant classes.³⁵
The Dm2 biosurfactant exhibited a similar glycolipid profile to Dh4 but with distinct spectral shifts indicative of subtle structural variations. A single C–H stretching band appeared at 2977.39 cm⁻¹, representing a +53.82 cm⁻¹ frequency shift relative to Dh4’s dual peaks at 2923.57 cm⁻¹, suggesting altered aliphatic chain composition with possible branching or shorter fatty acid chain lengths.³⁶ The carbonyl peak shifted to 1719.41 cm⁻¹ (+5.02 cm⁻¹ compared to Dh4), indicating a modified ester chemical environment—such carbonyl frequency variations reflect differences in functional group environments and sugar–lipid linkage geometries.³³ The C=C stretching at 1637.60 cm⁻¹ closely matched Dh4 (1636.88 cm⁻¹), indicating comparable unsaturated fatty acid incorporation from waste oil substrates.²¹ O–H hydroxyl stretching at 1341.94 cm⁻¹ was essentially superimposable with Dh4 (1343.38 cm⁻¹), confirming similar carbohydrate moiety composition. Notably, Dm2 lacked a distinct C–O glycosidic stretching peak in the 1052 cm⁻¹ region observed in Dh4, suggesting either reduced carbohydrate content, different sugar moieties, or altered glycosidic linkage configurations compared to Dh4.³³ The C–C aliphatic bonding peak appeared at 951.56 cm⁻¹ (+4.31 cm⁻¹ from Dh4), reflecting subtle variations in chain saturation or branching patterns. Critically, the absence of the C–Cl band at 831.72 cm⁻¹ indicated superior solvent removal and higher product purity compared to Dh4.³⁶ While both biosurfactants were confirmed as glycolipids that match rhamnolipid literature standards, with characteristic hydroxyl, carboxyl, and methoxyl groups, the spectroscopic differences suggest structural variations that may influence their differential antimicrobial and antibiofilm biological activities.³¹˒³³
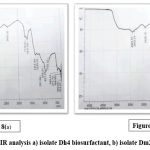 |
Figure 8: FTIR analysis a) isolate Dh4 biosurfactant, b) isolate Dm2 biosurfactant. |
Overall, FTIR spectra verified the glycolipid nature of both biosurfactants, consistent with rhamnolipid-type structures. The subtle spectral differences between Dh4 and Dm2 indicate variations in fatty acid and glycosidic components that may influence their physicochemical stability and bioactivity.³¹
Antiadhesive and Antibiofilm Activity
Slide Test
Microscopic examination of glass slides treated with biosurfactants from Dh4 and Dm2 demonstrated significant reduction in bacterial adhesion and biofilm formation compared to untreated controls. Visual assessment showed markedly reduced bacterial cell attachment for E. coli, B. subtilis, and S. paratyphi on biosurfactant-treated surfaces, indicating potent antiadhesive properties.
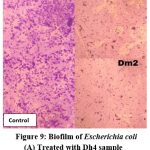 |
Figure 9: Biofilm of Escherichia coli. (A) Treated with Dh4 sample, (B) Treated with Dm2 sample. |
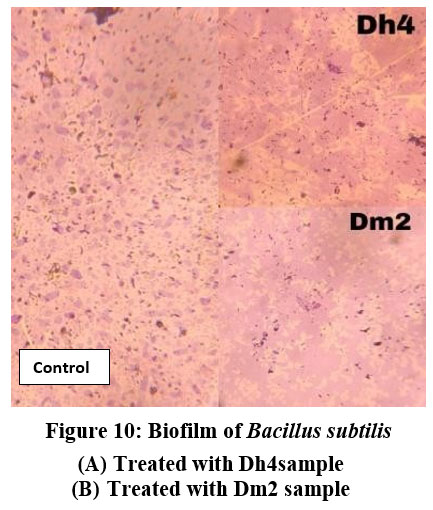 |
Figure 10: Biofilm of Bacillus subtilis. (A) Treated with Dh4 sample, (B) Treated with Dm2 sample. |
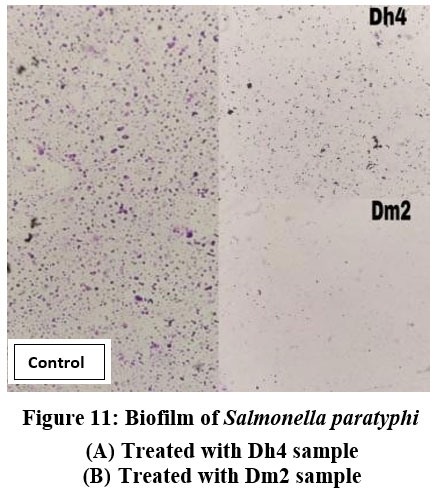 |
Figure 11: Biofilm of Salmonella paratyphi. (A) Treated with Dh4 sample, (B) Treated with Dm2 sample. |
This physical barrier effect suggests that biosurfactants modify surface properties, rendering them less favorable for initial bacterial attachment, a critical step in biofilm formation.9
Microtiter Plate Method
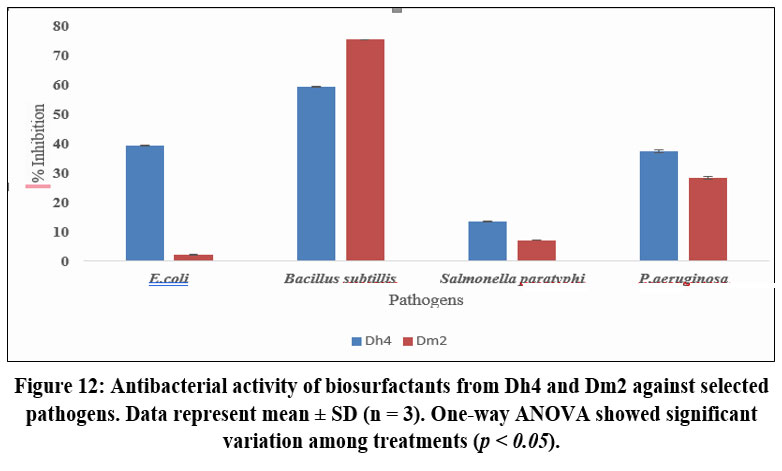
Quantitative analysis using the microtiter plate assay revealed dose-dependent antibiofilm activity for both biosurfactants. At a 5 mg/mL concentration, Dm2 biosurfactant exhibited the highest inhibition against B. subtilis (75.33%), followed by P. aeruginosa (28.26%), E. coli (2.26%), and S. paratyphi (7.2%). The Dh4 biosurfactant showed highest activity against B. subtilis (59.3%), E. coli (39.3%), P. aeruginosa (14.33%), and S. paratyphi (13.5%) (Figure 1). These results demonstrate that both biosurfactants possess significant antibiofilm potential, particularly against Gram-positive B. subtilis. The differential activity against various pathogens may be attributed to differences in bacterial cell wall composition and biofilm matrix characteristics.⁸˒³⁷
The superior antibiofilm activity observed in this study is comparable to or exceeds that reported for other microbial biosurfactants. For instance, biosurfactants from Lactobacillus rhamnosus showed similar concentration-dependent biofilm inhibition against pathogenic bacteria⁸ while lipopeptides from marine Bacillus species demonstrated effective biofilm disruption of P. aeruginosa and B. cereus.³⁸ The mechanisms underlying this antibiofilm activity likely include reduced bacterial adhesion, modifications in cell‐surface hydrophobicity, and the disruption of quorum-sensing systems, as reported in previous studies.¹²˒³⁹
Antimicrobial Activity
Agar well diffusion assays demonstrated that both biosurfactants possessed antimicrobial activity against the tested pathogenic bacteria. Dh4 biosurfactant produced inhibition zones of 12 mm against B. subtilis, 6 mm against S. paratyphi, 4 mm against E. coli, and 2 mm against P. aeruginosa. Dm2 biosurfactant showed zones of 3 mm, 7 mm, 3 mm, and 2 mm against the same pathogens, respectively (Table 4). These findings indicate direct antimicrobial effects in addition to antiadhesive properties.
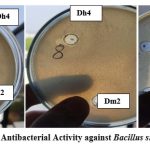 |
Figure 13: Dh4 and Dm2 Antibacterial Activity against Bacillus subtilis, S. paratyphi., E.coli |
Table 4: Antimicrobial activity (zone of inhibition) of biosurfactants
| Pathogen | Dh4 (mm) | Dm2 (mm) |
| B. subtilis | 12 | 3 |
| S. paratyphi | 6 | 7 |
| E. coli | 4 | 3 |
| P. aeruginosa | 2 | 2 |
The antimicrobial mechanisms of biosurfactants are multifaceted, involving membrane disruption, pore formation, and interference with essential bacterial processes.⁴⁰˒⁴¹ Glycolipid biosurfactants integrate into bacterial membranes, increasing permeability and causing cell lysis by disrupting lipid bilayer integrity. This membrane-targeting mechanism explains their broad-spectrum activity and potential to combat antibiotic-resistant pathogens.⁴²
Discussion
This study successfully demonstrated the isolation and characterisation of biosurfactant-producing halophilic bacteria from coastal marine environments, and their capacity to utilise waste vegetable oil as a sustainable carbon source. The findings contribute to the growing body of knowledge on marine microbial biosurfactants and their potential applications in biomedical and environmental sectors. Halophilic and halotolerant bacteria have emerged as promising biosurfactant producers due to their ability to thrive in extreme conditions and produce metabolites with unique properties.¹³˒¹⁴ The isolation of Dh4 and Dm2 from saline coastal waters aligns with previous studies reporting biosurfactant production by halophilic Bacillus, Halomonas, and Pseudomonas species from marine environments.²⁹˒⁴³ The morphological and biochemical characteristics of the isolated bacteria, including Gram-positive cocci (Dm2) and variable colony pigmentation, suggest potential taxonomic diversity warranting further molecular identification by 16S rRNA sequencing. The selection pressure exerted by high salinity, along with the presence of hydrocarbons in marine environments, likely drives the evolution of biosurfactant production in these bacteria. Biosurfactants facilitate hydrocarbon emulsification and uptake, thereby providing a competitive advantage for microbial survival and growth on hydrophobic substrates. ⁴⁴ This ecological adaptation translates into biotechnological potential for industrial biosurfactant production.
The successful utilization of waste vegetable oil for biosurfactant production represents a significant advancement toward sustainable bioprocessing. Waste cooking oils pose substantial environmental challenges due to improper disposal, contributing to water pollution and ecosystem degradation. ⁴⁵ By converting these waste streams into value-added biosurfactants, this approach addresses dual challenges of waste management and production economics. The biosurfactant yields obtained in this study (2.1–2.3 g/100 mL) are comparable to those reported for other waste-oil-based systems. For instance, Pseudomonas aeruginosa strains produced rhamnolipids at concentrations of 11.7–40 g/L from vegetable oil waste⁴⁶˒⁴⁷., while Bacillus species yielded lipopeptides at 0.011–0.05 g/L from frying oil waste. ⁴⁸˒⁴⁹ The relatively moderate yields observed in this study may be attributed to the halophilic nature of the isolates and non-optimized culture conditions. Optimising parameters such as C/N ratio, pH, temperature, and agitation could significantly enhance productivity.
The combined biochemical and spectroscopic analyses strongly indicate that the biosurfactants produced by Dh4 and Dm2 are glycolipid-type compounds. The positive saponification test confirms the presence of ester linkages. In contrast, the Fehling’s test demonstrates reducing sugar components, both of which are characteristic of glycolipids such as rhamnolipids, sophorolipids, or trehalolipids.⁵⁰˒⁵¹ FTIR spectroscopy provided detailed molecular fingerprints consistent with glycolipid biosurfactants. The strong C–H stretching vibrations (2923, 2853 cm⁻¹) indicate aliphatic hydrocarbon chains typical of fatty acid moieties, while the carbonyl stretching peak (1714 cm⁻¹) suggests ester linkages between sugar and lipid components. These spectral features closely match those reported for rhamnolipids and other bacterial glycolipids.⁵²˒³²
The potent antiadhesive and antibiofilm activities observed in this study represent the most significant biomedical potential, with up to 75% inhibition of B. subtilis and 39% inhibition of E. coli at 5 mg/mL.⁸ These effects may be attributed to mechanisms such as surface modification⁹˒⁵³., quorum-sensing interference¹²˒³⁷., membrane disruption⁴⁰˒⁴¹., and EPS degradation 8. The direct antimicrobial activity further complements these properties, indicating dual functionality through membrane permeabilization, metabolic disruption, and synergistic antibiotic enhancement.⁴⁰˒⁵³˒⁵⁴
Clinically, such biosurfactants hold promise for use in medical device coatings, wound care formulations, combination therapies, and drug delivery systems. ⁵⁵–⁵⁹ Environmentally, this system demonstrates sustainability by utilising waste vegetable oil to produce biodegradable, non-toxic compounds⁶⁰˒⁶¹., and leveraging halophilic cultures to reduce contamination risks and production costs. Although biosurfactant production remains costlier than synthetic surfactants, ongoing optimization of substrate utilization and downstream processing continues to narrow this gap, making such eco-friendly systems increasingly viable for pharmaceutical, cosmetic, and environmental applications. ²⁷˒⁶²
Future Perspectives
Future research should focus on molecular identification to establish precise taxonomic classification of isolates Dh4 and Dm2. Production optimization via response surface methodology and fed-batch cultivation strategies will enhance yields to commercially viable levels. Mechanistic studies employing confocal microscopy, transcriptomic analysis, and metabolomic profiling will clarify the molecular mechanisms underlying biofilm disruption and antiadhesive activity. A comprehensive biocompatibility assessment, including cytotoxicity evaluations, hemolysis assays, and in vivo animal toxicity studies, will establish safe dosing ranges for clinical applications. In vivo efficacy validation using murine biofilm infection models, pharmacokinetic analyses, and combination therapy studies with conventional antibiotics will demonstrate therapeutic potential. Formulation development encompassing nanoparticle encapsulation, stabilization strategies, and medical device coatings will enhance clinical applicability. Scale-up validation in pilot bioreactors and downstream processing optimization will determine industrial feasibility. Regulatory pathway engagement with FDA/EMA, GMP implementation, and phased clinical trials will facilitate translation toward approved therapeutic products. At the same time, genetic engineering approaches will generate higher-producing strains for expanded applications in infection prevention, bioremediation, and the food industry sectors.
Conclusion
This study successfully demonstrated the production of antiadhesive glycolipid biosurfactants from halophilic bacteria isolated from coastal marine waters using waste vegetable oil as a sustainable carbon source. The isolates Dh4 and Dm2 produced biosurfactants with significant surface-active properties, yielding 2.1–2.3 g/100 mL. Biochemical and spectroscopic analyses confirmed the presence of glycolipid structures bearing characteristic functional groups, consistent with rhamnolipid-type biosurfactants. The produced biosurfactants exhibited potent antiadhesive and antibiofilm activities against clinically relevant pathogens, including B. subtilis, E. coli, P. aeruginosa, and S. paratyphi, with inhibition rates up to 75%. Direct antimicrobial activity was also demonstrated.
These findings highlight the dual benefits of waste valorization and production of bioactive compounds with significant biomedical application potential. The ability to prevent and disrupt bacterial biofilms positions these biosurfactants as promising alternatives or adjuvants to conventional antimicrobial therapies, particularly for combating device-related and chronic infections. Future research focused on molecular identification, structural elucidation, production optimization, and in vivo efficacy evaluation will advance the translational potential of these halophilic biosurfactants toward commercial and clinical applications.
The sustainable production paradigm demonstrated in this study—utilising extremophile bacteria and waste substrates—represents a model for circular-economy approaches in biotechnology, contributing to environmental sustainability while generating high-value products for healthcare and industry.
Acknowledgment
The authors gratefully acknowledge the support provided by the Institute of Science & Technology for Advanced Studies & Research (ISTAR), The Charutar Vidya Mandal (CVM) University. We thank ISTAR for providing the necessary laboratory facilities essential for conducting this research. We also acknowledge SICART, Vallabh Vidyanagar, for facilitating FTIR spectroscopy analysis.
Funding Sources
This work was financially supported as a Minor Research Project funded by The Charutar Vidya Mandal (CVM) University, Vallabh Vidyanagar, Gujarat, India, under Project Number CVMU/RDC/MRP/SPA/2023-24/30118/1449.
Conflict of Interest
The authors do not have any conflict of interest.
Data Availability Statement
This statement does not apply to this article.
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Clinical Trial Registration
This research does not involve any clinical trials.
Permission to reproduce material from other sources
Not Applicable
Author Contributions
Boski Thakkar: Conceptualization, Methodology, Investigation, Data Collection, Formal Analysis, Visualization, Writing – Original Draft Preparation.
Nisha Daxini: Supervision, Validation, Resources, Project Administration, Writing – Review & Editing.
References
- Banat IM, Franzetti A, Gandolfi I, et al. Microbial biosurfactants production, applications and future potential. Appl Microbiol Biotechnol. 2010;87(2):427-444. doi:10.1007/s00253-010-2589-0
CrossRef - Naughton PJ, Marchant R, Naughton V, Banat IM. Microbial biosurfactants: Current trends and applications in agricultural and biomedical industries. J Appl Microbiol. 2019;127(1):12-28. doi:10.1111/jam.14243
CrossRef - Campos JM, Stamford TL, Sarubbo LA, de Luna JM, Rufino RD, Banat IM. Microbial biosurfactants as additives for food industries. Biotechnol Prog. 2013;29(5):1097-1108. doi:10.1002/btpr.1796
CrossRef - Flemming HC, Wuertz S. Bacteria and archaea on Earth and their abundance in biofilms. Nat Rev Microbiol. 2019;17(4):247-260. doi:10.1038/s41579-019-0158-9
CrossRef - Datta D, Ghosh S, Kumar S, et al. Microbial biosurfactants: Multifarious applications in sustainable agriculture. Microbiol Res. 2024;279:127551. doi:10.1016/j.micres.2023.127551
CrossRef - Paganelli FL, Willems RJ, Leavis HL. Optimizing future treatment of enterococcal infections: attacking the biofilm. Trends Microbiol. 2012;20(1):40-49. doi:10.1016/j.tim.2011.11.001
CrossRef - Sharma D, Sharma BS, Shukla AK. Biotechnological approach of microbial lipase: a review. 2011;10(1):23-40. doi:10.3923/biotech.2011.23.40
CrossRef - Patel M, Gaba S, Maurya R, Mitra D, Chaudhary S. Inhibition of bacterial adhesion and antibiofilm activities of biosurfactant producing Lactobacillus rhamnosus with cell-bound biosurfactant. Microb Pathog. 2021;161:105272. doi:10.1016/j.micpath.2021.105272
CrossRef - Hamza F, Satpute S, Banpurkar A, Kumar AR, Zinjarde S. Biosurfactant from a marine bacterium disrupts biofilms of pathogenic bacteria in a tropical aquaculture system. FEMS Microbiol Ecol. 2017;93(11):fix140. doi:10.1093/femsec/fix140
CrossRef - Barale SS, Ghane SG, Sonawane KD. Purification and characterization of antibacterial surfactin isoforms produced by Bacillus velezensis AMB Express. 2022;12(1):7. doi:10.1186/s13568-022-01348-3
CrossRef - Yan X, Gu S, Cui X, et al. Antimicrobial, anti-adhesive and anti-biofilm potential of biosurfactants isolated from Pediococcus acidilactici and Lactobacillus plantarum against Staphylococcus aureus Microb Pathog. 2019;127:12-20. doi:10.1016/j.micpath.2018.11.039
CrossRef - Singh S, Bhatia S. Quorum sensing inhibitors: curbing pathogenic infections through inhibition of bacterial communication. Iran J Pharm Res. 2021;20(2):486-514. doi:10.22037/ijpr.2020.113470.14318
- Márquez-Villa JM, Mateos-Díaz JC, Rodríguez-González JA, Camacho-Ruíz RM. Optimization of lipopeptide biosurfactant production by Salibacterium 4CTb. Microorganisms. 2022;10(5):983. doi:10.3390/microorganisms10050983
CrossRef - Donio MBS, Ronica FA, Viji VT, et al. Isolation and characterization of halophilic Bacillus BS3 able to produce pharmacologically important biosurfactants. Asian Pac J Trop Med. 2013;6(11):876-883. doi:10.1016/S1995-7645(13)60156-X
CrossRef - Liu G, Zhong H, Yang X, et al. Advances in applications of rhamnolipid biosurfactant in environmental remediation: A review. Biotechnol Bioeng. 2017;115(4):796-814. doi:10.1002/bit.26517
CrossRef - Md Badrul Hisham NH, Ibrahim MF, Ramli N, Abd-Aziz S. Production of biosurfactant from used cooking oil by Bacillus HIP3 for heavy metals removal. Molecules. 2019;24(14):2617. doi:10.3390/molecules24142617
CrossRef - Morikawa M, Hirata Y, Imanaka T. Structure–function relationship of lipopeptide biosurfactants. Biochim Biophys Acta. 2000;1488(3):211-218. doi:10.1016/S1388-1981(00)00124-4
CrossRef - Bodour AA, Drees KP, Maier RM. Distribution of biosurfactant-producing bacteria in undisturbed and contaminated soils. Appl Environ Microbiol. 2003;69(6):3280-3287. doi:10.1128/AEM.69.6.3280-3287.2003
CrossRef - Edel W, Kampelmacher EH. Comparative studies on the isolation of “sublethally injured” salmonellae in nine European laboratories. Bull World Health Organ. 1973;48(2):167-174.
- Nayarisseri A, Singh P, Upadhyay SK. Screening and characterisation of biosurfactant-producing Bacillus subtilis strain ANSKLAB03. 2018;14(6):304-314. doi:10.6026/97320630014304
CrossRef - Patowary K, Patowary R, Kalita MC, Deka S. Characterization of biosurfactant produced during degradation of hydrocarbons using crude oil. Front Microbiol. 2017;8:279. doi:10.3389/fmicb.2017.00279
CrossRef - Rajan A, Sudha V, Abraham TE. Enzymatic modification of cassava starch by fungal lipase. Ind Crops Prod. 2010;27(1):50-59. doi:10.1016/j.indcrop.2007.07.003
CrossRef - Cioffi N, Rai M, Barakat MA. Antimicrobial Nanomaterials for Food Packaging. Academic Press; 2018.
- Gomaa EZ. Antimicrobial and anti-adhesive properties of biosurfactant produced by lactobacilli isolates. J Gen Appl Microbiol. 2013;59(6):425-436. doi:10.2323/jgam.59.425
CrossRef - Ciurko D, Krzemińska M, Głowacka K, Woźniak-Budych M, Jurga S. Production and characterization of lipopeptide biosurfactant by Pseudomonas antarctica RSC Adv. 2023;13(33):23192-23205. doi:10.1039/D3RA03408A
CrossRef - Wu B, Xiu J, Yu L, Huang L, Yi L, Ma Y. Biosurfactant production by Bacillus subtilis SL for enhanced oil recovery. Sci Rep. 2022;12:7785. doi:10.1038/s41598-022-12025-7
CrossRef - Gaur VK, Sharma P, Sirohi R, et al. Production of biosurfactants from agro-industrial waste and waste cooking oil. Bioresour Technol. 2022;343:126059. doi:10.1016/j.biortech.2021.126059
CrossRef - Mulligan CN. Sustainable production of biosurfactants using waste substrates. In: Advancements in Biosurfactants Research. Springer; 2023:57-77. doi:10.1007/978-3-031-21682-4_3
CrossRef - Suriya M, Mayavu P, Shanmugasundaram S, Sugesh S. Glycolipid biosurfactant from halophilic Bacillus Int J Adv Res Biol Sci. 2016;3(9):161-169. doi:10.22192/ijarbs.2016.03.09.023
CrossRef - Barakat KM, Hassan SW, Darwesh OM. Biosurfactant production by haloalkaliphilic Bacillus Egypt J Aquat Res. 2017;43(3):205-211. doi:10.1016/j.ejar.2017.09.001
CrossRef - Albasri HM, Almohammadi AA, Alhhazmi A, et al. Production and characterization of rhamnolipid biosurfactant from Geobacillus stearothermophilus. Front Microbiol. 2024;15:1358175. doi:10.3389/fmicb.2024.1358175
CrossRef - Cerqueira dos Santos S, Araújo Torquato C, de Alexandria Santos D, et al. Rhamnolipids from Pseudomonas aeruginosa with antiviral and antimicrobial activity. Sci Rep. 2024;14:4629. doi:10.1038/s41598-024-54828-w
CrossRef - Hosseini S, Sharifi R, Habibi A, Ali Q. Rhamnolipids from Pseudomonas oryzihabitans during crude oil biodegradation. Front Microbiol. 2024;15:1459112. doi:10.3389/fmicb.2024.1459112
CrossRef - Deepika KV, Raghuram M, Chari B. Rhamnolipid production by Pseudomonas aeruginosa KVD-HR42. Afr J Microbiol Res. 2017;11(6):218-231. doi:10.5897/AJMR2015.7881
CrossRef - FTIR spectroscopy as a method to determine chemical bonds in biosurfactants. PubMed Central; 2019.
- Sandhu M, Paul AT, Proćków J, et al. PCB-77 biodegradation potential of biosurfactant-producing bacteria. Front Microbiol. 2022;13:952374. doi:10.3389/fmicb.2022.952374
CrossRef - Adnan M, Siddiqui AJ, Noumi E, et al. Biosurfactant from Lactobacillus acidophilus exhibits antibiofilm activity. Biomol Biomed. 2023;23(6):1051-1068. doi:10.17305/bb.2023.9324
CrossRef - Song B, Wang YZ, Wang GY, et al. Lipopeptide 6-2 from Bacillus amyloliquefaciens anti-CA. Mar Pollut Bull. 2016;108(1-2):62-69. doi:10.1016/j.marpolbul.2016.04.062
CrossRef - Deep A, Chaudhary U, Gupta V. Quorum sensing and bacterial pathogenicity. J Lab Physicians. 2011;3(1):4-11. doi:10.4103/0974-2727.78553
CrossRef - Wiman E, Zattarin E, Aili D, et al. Novel broad-spectrum antimicrobial lipopeptides. Sci Rep. 2023;13:4104. doi:10.1038/s41598-023-31185-8
CrossRef - Ali N, Pang Z, Wang F, Xu B, El-Seedi HR. Lipopeptide biosurfactants from Bacillus: Types, production, biological activities, and applications in food. J Food Qual. 2022;2022:3930112. doi:10.1155/2022/3930112
CrossRef - Kourmentza K, Gromada X, Michael N, et al. Antimicrobial activity of lipopeptide biosurfactants against foodborne pathogens and spoilage microorganisms and their cytotoxicity. Front Microbiol. 2021;11:561060. doi:10.3389/fmicb.2020.561060
CrossRef - Wu Y, Xu M, Xue J, Shi K, Gu M. Characterization and enhanced degradation potentials of biosurfactant-producing bacteria isolated from a marine environment. ACS Omega. 2019;4(1):1645-1651. doi:10.1021/acsomega.8b02653
CrossRef - Kubicki S, Bollinger A, Katzke N, Jaeger KE, Loeschcke A, Thies S. Marine biosurfactants: Biosynthesis, structural diversity and biotechnological applications. Mar Drugs. 2019;17(7):408. doi:10.3390/md17070408
CrossRef - Caporusso A, Radice M, Biundo A, et al. Waste cooking oils as a sustainable feedstock for bio-based applications: A systematic review. J Biotechnol. 2025;400:48-65. doi:10.1016/j.jbiotec.2025.02.0
CrossRef - Hazra C, Kundu D, Ghosh P, Joshi S, Dandi N, Chaudhari A. Screening and identification of Pseudomonas aeruginosa AB4 for improved biosurfactant production using agro-based materials. J Chem Technol Biotechnol. 2010;85(11):1450-1456. doi:10.1002/jctb.2480
CrossRef - Nitschke M, Costa SG, Contiero J. Rhamnolipid surfactants: An update on the general aspects of these biomolecules. Biotechnol Prog. 2005;21(6):1593-1600. doi:10.1021/bp050239p
CrossRef - Jajor P, Piłakowska-Pietras D, Krasowska A, Łukaszewicz M. Surfactin analogues produced by Bacillus subtilis grown on rapeseed cake. J Mol Struct. 2016;1126:141-146. doi:10.1016/j.molstruc.2016.02.014
CrossRef - Hentati D, Chebbi A, Mahjoubi M, et al. Investigation of halophilic bacterial biosurfactant for environmental applications: A review. Ecotoxicol Environ Saf. 2019;184:109623. doi:10.1016/j.ecoenv.2019.109623
CrossRef - Rahman KS, Rahman TJ, McClean S, Marchant R, Banat IM. Rhamnolipid production by Pseudomonas aeruginosa using low-cost materials. Biotechnol Prog. 2002;18(6):1277-1281. doi:10.1021/bp020071x
CrossRef - Gaur VK, Regar RK, Dhiman N, et al. Biosynthesis and characterization of sophorolipid biosurfactant by Candida: Application as food emulsifier and antibacterial agent. Bioresour Technol. 2019;285:121353. doi:10.1016/j.biortech.2019.121353
CrossRef - Sultana S, Sultana R, Al-Mansur MA, et al. Industrially potent rhamnolipid-like biosurfactant from oil-degrading Bacillus velezensis RSC Adv. 2024;14(34):24516-24533. doi:10.1039/d4ra03632f
CrossRef - Ceresa C, Rinaldi M, Chiono V, et al. Harnessing the potential of biosurfactants for biomedical and pharmaceutical applications. 2023;15(8):2156. doi:10.3390/pharmaceutics15082156
CrossRef - Ndlovu T, Rautenbach M, Vosloo JA, Khan S, Khan W. Characterisation and antimicrobial activity of biosurfactant extracts produced by Bacillus amyloliquefaciens and Pseudomonas aeruginosa. AMB Express. 2017;7(1):1-11. doi:10.1186/s13568-017-0375-8
CrossRef - Bjerk TR, Severino P, Jain S, et al. Biosurfactants: Properties and applications in drug delivery, biotechnology and ecotoxicology. Bioengineering (Basel). 2021;8(8):115. doi:10.3390/bioengineering8080115
CrossRef - Ohadi M, Forootanfar H, Dehghannoudeh N, Banat IM, Dehghannoudeh G. Role of surfactants and biosurfactants in the wound healing process: A review. J Wound Care. 2023;32(Suppl 4a):xxxix-xlvi. doi:10.12968/jowc.2023.32.Sup4a.xxxix
CrossRef - Subramaniam MD, Kim IH, Jeon JH, Kim HJ. Antimicrobial lipopeptides: Efficacy, applications, and challenges. Front Microbiol. 2020;11:577975. doi:10.3389/fmicb.2020.577975
- Kaczorek E, Pacholak A, Zdarta A, Smułek W. Impact of biosurfactants on microbial cell properties increasing hydrocarbon bioavailability. Colloids Interfaces. 2018;2(3):35. doi:10.3390/colloids2030035
CrossRef - Wang X, Niu C, Liu M, et al. Application of biosurfactants in medical sciences: Advancements in wound healing, drug delivery, and tissue engineering. 2024;29(11):2590. doi:10.3390/molecules29112606
CrossRef - Edwards KR. Surfactants based on renewable raw materials. J Ind Ecol. 2003;7(3-4):47-62. doi:10.1162/108819803323059398
CrossRef - Banat IM. Biosurfactants production and possible uses in microbial enhanced oil recovery and oil pollution remediation: A review. Bioresour Technol. 1995;51(1):1-12. doi:10.1016/0960-8524(94)00101-6
CrossRef - Soares da Silva RCF, de Almeida DG, Brasileiro PPF, et al. Production, formulation and cost estimation of a commercial biosurfactant. 2019;30:191-201. doi:10.1007/s10532-018-9830-4
CrossRef
Accepted on: 12-12-2025
Second Review by: Dr. Saeed Kewedar
Final Approval by: Dr. Eugene A. Silow