Oxadiazole Skeleton with In Silico Modelling and Docking Study with Neuroprotective Effects of Cholinesterase Inhibitors of (PDB ID: 7E3H) on Alzheimer’s Disease
1Department of Pharmaceutical Chemistry, PRES's college of Pharmacy (D and B Pharm), Chincholi, Sinner, Nashik Maharashtra, India..
2Department of Pharmaceutical Chemistry, Pravara Rural College of Pharmacy Pravaranagar, Rahata, Ahmednagar, Maharashtra, India.
3Department of Pharmaceutical Chemistry, Trimurti Gramin pharmacy College, Trimurti Nagar; Newasa Phata, Khadke, Newasa, Ahilayanagar, Maharashtra, India
Correspondence Author Email: rohit.bhor69@gmail.com
DOI : http://dx.doi.org/10.13005/bbra/3463
ABSTRACT:Memory loss and cognitive impairment are the main symptoms of Alzheimer's disease (AD), a neurodegenerative brain illness that affects millions of people globally. Alzheimer's disease is a neurodegenerative illness that is characterized by pathological features such as β-amyloid deposits and neurofibrillary tangles in the cerebral cortex. Therefore, identifying strong (PDB ID: 7E3H) inhibitors is an important and urgent objective for their potential use in Alzheimer's disease treatment. We have optimized the geometry of the analyzed structures using the AM3 semi-empirical approach. We have utilized offline resources for molecular docking research, including Marvin Sketch for chemical structure representation and the Protein Data Bank for PDB protein file downloads. These tools are a supplement to internet resources like PyRx 0.9, Swiss ADMET, and PubChem. We used the Swiss Protein Data Bank Viewer for protein synthesis. This work has shown that the structures under consideration generally form stronger complexes with the enzyme than the known inhibitor. In light of the molecular docking studies, the compounds leaders (S)-2-((4-((3-methyl-5-phenyl-1,2,4-oxadiazol-2(3H)-yl)methyl) phenyl)amino)-N-phenylacetamide AM1 and (R)-2-((4-((3-methyl-5-phenyl-1,2,4-oxadiazol-2(3H)-yl)methyl)phenyl)amino)-N-(2-nitrophenyl) acetamide AM2, and 1-(2-chlorophenyl)-3-((4-(((3S,5R)-3-methyl-5-phenyl-1,2,4-oxadiazolidin-2-yl)methyl) phenyl) amino) propane-1,2-dione AM 8 have been chosen. The Lipinski criterion compliance of the compound leaders' structures has been examined. Proponents of the medications on the list could suggest more research on treating Alzheimer's disease. In vitro and in vivo biological tests are still required despite the positive outcomes of in silico research.
KEYWORDS:Alzheimer’s disease; Docking; In silico Study; 1,3,4-oxadiazole; PDB:7E3H
Introduction
Alzheimer’s disease (AD) causes memory loss and cognitive deterioration.1 Perhaps because its cause is unknown, AD has no proven cure. Gliosis, neurofibrillary tangles, inflammation, synapse loss, and extracellular amyloid beta (A) plaque are the hallmarks of AD. Different ideas exist about Alzheimer’s disease.2 Numerous theories have been put forth, including the lysosome hypothesis, presenilin hypothesis, Ca2+ dysregulation theory, tau hypothesis, oligomer hypothesis, and β-amyloid hypothesis. Low acetylcholine is necessary for AD. Acetylcholine levels can be optimized using esterase inhibitors to stop degradation. Amyloid plaques and other forms of neurodegeneration, neurotoxicity, and snap to toxicity can result from an excess of A-amyloid peptides.3 Calcium-Sensing Receptor (CaSR) imbalance is associated with inflammation and neurological disorders such as Alzheimer’s. Lastly, lysosome hypothesis contends that pH gene abnormalities cause issues with the autophagy-lysosomal system. Acetylcholinesterase (AChE) are implicated in the sickness. By hastening the production of Alzheimer’s-like A peptide aggregation, they promote neurotoxicity.4 Low acetylcholine, aberrant amyloid, T protein aggregation, inflammation, and oxidative stress are all present in AD patients. AChE inhibitors are currently the focus of most Alzheimer’s disease research.5 Type-B carboxylesterase is acetylcholinesterase. common ancestor of carboxylesterase type B. Acetylcholine and choline esters are broken down by this serine hydrolase. Neurofibrillary tangles (tau pathology) and β-amyloid plaques in the cerebral cortex are pathological characteristics of Alzheimer’s disease (AD), a neurodegenerative illness.6 In this instance, neurofibrillary tangles are intimately associated with the disease’s symptoms and course, and the pathology of β-amyloid plaques varies greatly. Alzheimer’s disease (AD), the most common kind of dementia and a deadly neurological illness characterized by memory and cognitive problems, mostly affects elderly people.7 It is a neurological disorder that damages brain cells and progressively impairs cognitive abilities including speech, memory, attention, and judgment.8 There is currently no proven therapeutic therapy for the progressive and cognitive decline brought on by aging in AD, despite the fact that the mechanism behind these symptoms is still unclear9. According to estimates, there are 35 million AD sufferers worldwide.10 By 2030, this figure is expected to rise to 65 million, and by 2050, it will reach 115 million. These numbers highlight the need to develop an effective therapy. The loss of cholinergic neurons in the basal frontal cortex, intracellular neurofibrillary tangles caused by oxidative stresses, protein hyperphosphorylation, and extracellular amyloid (A) plaques are the main pathogenic characteristics of AD.11 Inadequate cholinergic transmission is one aspect that is thought to play a significant role in AD. This leads to the development of behavioural, functional, and cognitive symptoms.12 Overview Alzheimer’s disease (AD), the most common kind of dementia and a deadly neurological illness characterized by cognitive and memory problems, mostly affects elderly people.14 The loss of cholinergic neurons in the basal frontal cortex, intracellular neurofibrillary tangles caused by oxidative stresses, hyperphosphorylation of the τ protein, and extracellular β-amyloid (βA) plaques are the most significant pathogenic hallmarks of AD. Inadequate cholinergic transmission is one aspect that is thought to play a significant role in AD. This leads to the development of behavioural, functional, and cognitive symptoms.15 The cholinergic theory states that cholinesterase enzymes will be suppressed as a result of the brain’s inadequate acetylcholine levels. Therefore, improving cholinergic system dysfunction is often the aim of treatment using receptor agonists or cholinesterase (ChE) inhibitors.16 FDA-approved medications based on the “one drug one target” strategy donepezil, galantamine, and rivastigmine are being used to treat Alzheimer patients by inhibiting AChE, highlighting the inadequacies of this strategy given the complexity of AD. On the other hand, the most cutting-edge treatment approaches now in use are predicated on the “one drug–many targets” approach, which promotes the use of drugs with multiple effects at different target sites.17 The medication that is most commonly administered and effective in treating AD at all stages is donepezil. Because AChE binds to both the catalytic and peripheral anionic sites, it is effectively inhibited. When developing novel medications and treatment strategies, this “double binding” characteristic should be taken into account.18 We developed, produced, and investigated the biological effects of new compounds consisting of oxadiazole and various secondary amine derivatives in this work, taking into consideration the chemical composition of the donepezil molecule.19 We also examined pertinent molecular modelling research.
Materials and Methods
Devices and Materials
In contemporary drug development, docking is frequently used to investigate the relationship between the target ligand receptor, the intended lead chemical compound, and its protein receptors. Computational methods were used to carry out the research electronically. In addition to online resources, we use tools like Protein Data Bank (available at public domain websites like www.rcsb.org/pdb) and Marvin Sketch to illustrate chemical structures.20 Auto dock Vina was built on top of PyRx V0.9.2. Table 1 includes the oxadiazole AM1 to AM10 derivatives.
Preparation of Protein
Ligand Preparation
The molecules are made in both two and three dimensions using the sketch tool in Discovery Studio. Marvin Sketch was used to create the compound, optimize it in three dimensions, and save it in Structure Data File.
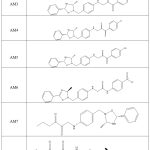 |
Table 1: Derivatives of Oxadiazole AM1 to AM10 |
Results
ADMET Prediction
In-silico ADMET are the key pharmacokinetic and safety properties used to evaluate how a drug behaves inside the body — from how it is absorbed into the bloodstream to how it is distributed, metabolized, eliminated, and whether it causes any toxic effects characteristics of possible drugs were estimated using a computer program called Swiss ADME prediction. The distribution constant of the molecule in 1-octanol in water, the number of N-ON bond and n-OH-NH hydrogen bond acceptor and donor sites, the CNS activity, the fraction of oral absorption by humans, the topological polar surface area (TPSA)and the count of H bond acceptors and donors were all determined. Any medication or synthetic compound’s ADME properties can be better understood with the help of the information presented here. ADMET Table 2 provided the predictions for Oxadiazole Derivatives AM1 through AM10. It was also found that there were pharmacological similarities, thumb rule (RO5) violations, and three-to-one rule violations. The five ideal characteristics of a given molecule are a MW of 500, a count of five H bond donors, and a count of ten H bond acceptors.
Table 2: ADMET Prediction of Oxadiazole Derivatives AM1 to AM10
| Comp. Code | Bioavailability Score | CYP31A2 inhibitor | CYP2C19 inhibitor |
CYP2C9 inhibitor | CYP2D6 inhibitor | CYP3A4 inhibitor |
Pgp substrate | BBB permeant | GI absorption | Dock score |
| AM1 | 0.55 | No | Yes | Yes | Yes | Yes | No | Yes | High | -11.5 |
| AM2 | 0.55 | No | Yes | Yes | No | Yes | No | No | High | -10.9 |
| AM3 | 0.55 | No | Yes | Yes | Yes | Yes | No | Yes | High | -10.5 |
| AM4 | 0.55 | No | Yes | Yes | Yes | Yes | No | Yes | High | -11.7 |
| AM5 | 0.55 | No | Yes | Yes | Yes | Yes | Yes | No | High | -11.5 |
| AM6 | 0.55 | No | Yes | Yes | No | Yes | No | No | High | -11.0 |
| AM7 | 0.55 | No | Yes | Yes | Yes | Yes | No | Yes | High | -10.1 |
| AM8 | 0.55 | No | Yes | Yes | Yes | Yes | No | No | High | -7.6 |
| AM9 | 0.55 | No | Yes | Yes | Yes | Yes | Yes | Yes | High | -11.4 |
| AM10 | 0.55 | No | No | Yes | Yes | Yes | Yes | No | High | -9.2 |
In-silico Molecular Docking Studies
After a review of the literature on oxadiazole derivative research and docking investigations, ten oxadiazole scaffolds in total were developed for our study. Molecular docking using Pyrex V0.9 has been used to forecast possible interactions between the protein and its inhibitors. Using molecular docking, the binding mechanism competence of acetylcholinesterase with ten oxadiazole drugs was examined. The protein molecules were in proximity to the synthetic ligand. An acceptable affinity for interacting with a target receptor was revealed by docking values ranging from 8 to 12 kcal/mol for our developed medicines. The reference medication donepezil, the natural ligand, and the synthetic compounds were all included in the docking experiments. Muegge#violations, Egan#violations, Veber#violations, and Ghose# #violations; #donors; #acceptors of H-bonds; # Lipinski the rotatable bonds for Oxadiazole Derivatives AM1 through AM10 are predicted in Table 3. AM 1, AM 8, and AM 2 all have binding energies of 12 kcal/mol. Compared to donepezil (12.76 K/cal), the docking findings for compounds AM 3, AM 7, and AM 9 (11.5 K/cal) and compound AM 5 are similar. The remaining molecule has good to moderate action, which is better than that of typical drugs. Critical amino acids in the ligand-binding domain of human AChE inhibitors have also been discovered. The non-covalent interactions between the examined ligands and the AChE inhibitor ligand-binding domain were identified. Small polar areas at each end of the big lipophilic pocket that is the active 7E3H | pdb_00007e3h site. These regions include crystalline water molecules that actively participate in ligand-protein interactions. According to X-ray diffraction analysis, the molecule of AM1 interacts with the active site of 7E3H | pdb_00007e3h during molecular docking. This is because a complex system of intermolecular hydrogen bonds is formed between the nitrogen atom of 1,3,4-oxadiazole rings and water molecules, H2O. With a calculation time of 10 seconds, the computed energy of inhibitor binding (1) with the enzyme’s active site is -11.5 kcal/mol. The root-mean-square deviation of atomic positions (RMSD) of the inhibitor’s predicted location in the enzyme’s active site is 1.4 Å, which is comparable to the findings of X-ray analysis. The molecule of AM2 interacts with the active site of 7E3H | pdb_00007e3h in addition, according to molecular docking data, because of the complex system of intermolecular hydrogen bonds that are formed between water molecules and the nitrogen atom of 1,3,4-oxadiazole rings. According to the molecular docking, AM3 and AM4 have formed the most stable complexes with 7E3H | pdb_00007e3h. (S)-2-((4-((3-methyl-5-phenyl-1,2,4-oxadiazol-2(3H)-yl)methyl)phenyl)amino)-N-phenyl acetamide AM1 and (R)-2-((4-((3-methyl-5-phenyl-1,2,4-oxadiazol-2(3H)-yl)methyl)phenyl)amino)-N-(2-nitrophenyl)acetamideAM2 and 1-(2-chlorophenyl)-3-((4-(((3S,5R)-3-methyl-5-phenyl-1,2,4-oxadiazolidin-2-yl)methyl) phenyl) amino) propane-1,2-dione AM effectively binds to 7E3H | pdb_00007e3h by the formation of two inter molecular hydrogen bonds. The bond length between the oxygen atom of valine and the hydrogen atom of the amine moiety is 2.853 Å. In light of the aforementioned findings, we set out to develop and manufacture new compounds based on the 1,3,4-oxadiazole scaffold. 2D and 3D Images of 1-(2-chlorophenyl)-3-((4-(((3S,5R)-3-methyl-5-phenyl-1,2,4-oxadiazolidin-2-yl)methyl) phenyl) amino) propane-1,2-dione AM 8 were shown in Figure 3. Additionally, the ortho position of the phenyl ring was exploited to introduce the monocyclic polar 1,3,4-oxadiazole motif with a five- or six-membered heterocyclic ring fused to benzene, respectively. In the third technique, linker characteristics were investigated by using 4- or 6-atom linkers to bridge the aryl or heteroaryl motif at the ortho position of the phenyl ring at position 5 of the oxadiazole core. These motifs and linkers may cooperate with amino acids in the AChE enzyme’s active site as donors or acceptors of H bonds.
Table 3: Prediction of Muegge#violations; Egan#violations; Veber#violations; Ghose#violations Lipinski#violations; #H-bonddonors; #H-bondacceptors; #Rotatable bonds for Oxadiazole Derivatives AM1 to AM10
| Comp code | Muegge #violations |
Egan #violations |
Veber #violations |
Ghose #violations |
Lipinski #violations |
#H-bond donors |
#H-bond acceptors |
#Rotatable bonds |
| AM1 | 0 | 0 | 0 | 0 | 0 | 2 | 4 | 8 |
| AM2 | 0 | 0 | 0 | 1 | 0 | 2 | 6 | 9 |
| AM3 | 1 | 0 | 0 | 1 | 0 | 2 | 5 | 8 |
| AM4 | 1 | 0 | 0 | 1 | 0 | 2 | 5 | 8 |
| AM5 | 0 | 0 | 0 | 0 | 0 | 3 | 6 | 8 |
| AM6 | 0 | 0 | 0 | 1 | 0 | 2 | 6 | 9 |
| AM7 | 0 | 0 | 0 | 0 | 0 | 2 | 5 | 9 |
| AM8 | 0 | 0 | 0 | 0 | 0 | 2 | 6 | 9 |
| AM9 | 0 | 0 | 0 | 0 | 0 | 2 | 5 | 8 |
| AM10 | 0 | 0 | 1 | 0 | 0 | 2 | 6 | 11 |
Discussion
The novel oxadiazole-based derivatives’ structure-activity relationship investigations showed that the disubstituted-1,3,4-oxadiazole pharmacophoric entity is well tolerated for AChE inhibition. AChE inhibitory activity was clearly stronger when a benzyl moiety was grafted at position 3 of the oxadiazole ring (AM1 and AM2) than when phenyl and 2-nitroaniline; 2-chlorobenzaldehyde moieties were used.23 It was interesting to note that only compound AM1, which had the benzyl moiety, exhibited strong AChE inhibitory action among oxadiazole derivatives with the hydrazide scaffold in the ortho position of the phenyl ring of the fifth position of the oxadiazole ring.24 The AChE inhibitory potential was enhanced by moving the hydrazide moiety into the 4-hydroxybenzaldehyde scaffold. Subsequent examination of amide derivatives showed that the lipophilic and electron-rich 2-chlorobenzaldehyde; 4-chlorobenzaldehyde; 4-hydroxybenzaldehyde containing derivatives attached to phenyl 1,3,4-oxadiazole motif (AM3; AM4, and AM5) were less tolerant of AChE inhibition than the polar and electron-poor 4-nitroaniline motif (AM6). 2D and 3D Images of (R)-2-((4-((3-methyl-5-phenyl-1,2,4-oxadiazol-2(3H)-yl)methyl)phenyl)amino)-N-(2-nitrophenyl) acetamide AM2 were shown in Figure 2.The inhibition of AChE was positively impacted by the addition of the 1,3,4-oxadiazole scaffold to the amide moiety. The 1,3,4-oxadiazole derivatives containing the N-amide scaffold were shown to be the most promising neuroprotective Alzheimer’s disease (AD) candidates in terms of their neuroprotective potential.25 Grafting the lipophilic electron-deficient glutaraldehyde at position 3 of the oxadiazole ring in N-amide derivatives shows notable neuroprotective potential against Alzheimer’s disease (AD).26 A slight inhibition of human acetylcholinesterase (AChE) was observed in the 1,3,4-oxadiazole amide derivatives (AM1 to AM10). Notably, the human acetylcholinesterase (AChE) inhibitory potential was significantly enhanced by adding the lipophilic electron-deficient aniline, 2-nitroaniline, and 2-chlorobenzaldehyde phenyl at position 3 of the oxadiazole ring (AM1-AM3) (Table 4).27 Research We looked into the pharmacokinetic properties of the developed ligands using the SWISS ADME platform.28 It was discovered that the planned compounds included one to four hydrogen bond donors. With a few exceptions, the anticipated stomach absorption rate of the proposed compounds was likewise found to be good. Crossing the blood-brain barrier (BBB) requires consideration of the topological polar surface area (TPSA).29 Our compounds’ TPSA values fell within a reasonable range, suggesting that practically all the molecules may pass through the blood-brain barrier with ease. Acetylcholinesterase inhibitors, on the other hand, are used to treat and control Alzheimer’s disease because they easily pass across the blood-brain barrier.30 It has been discovered that our developed compounds show outstanding BBB penetrability. It was anticipated that the 1-octyl alcohol and water partition coefficient would fall between 2.5 and 3 and that there would be one to three potential metabolic pathways.31 The Lipinski five-rule was maintained. Following oral administration, they are all easily absorbed and cross the blood-brain barrier (BBB). This suggests that practically every property of the ligand falls inside the permitted range. Table 3 lists the distinct ADMET values for each chemical. The goal of the docking study is to justify the biological activities of the target molecules and anticipate the binding interactions between the synthesized medication candidates and the AChE enzyme’s active site. In this investigation, Pyrex and Discovery Studio applications were utilized. Prior to docking the synthesized molecules, the protein data library was used to download the protein crystal structure of AChE (PDB: 7E3H) and confirm the docking process. With an RMSD of 0.42 Å, which is under the cut off limit (< 1 Å), the co-crystallized molecule (donepezil) was docked in the AChE enzyme’s active site to get a posture that overlapped the experimental pose during the validation stage. Nevertheless, of the currently produced series, analogue 10 was shown to be the most effective inhibitor of the targeted AChE enzymes. The most potent analogue 10’s comprehensive protein–ligand interaction (PLI) profile against the AChE enzyme reveals a number of noteworthy interactions with the enzyme’s active region, including residues like GLU4,ASP5,ALA6,GLU7,LEU8,LEU9, VAL10,THR11, VAL12, ARG13,GLU4.ASP5, ALA6,GLU7, LEU8,LEU9,VAL10,THR11, VAL12,ARG13,GLY14, GLY15 (Table 4). 2D and 3D Images of (S)-2-((4-((3-methyl-5-phenyl-1,2,4-oxadiazol-2(3H)-yl)methyl)phenyl)amino)-N-phenylacetamide AM1 were shown in Figure 1 and Ramachandran plot for 1-(2-chlorophenyl)-3-((4-(((3S,5R)-3-methyl-5-phenyl-1,2,4-oxadiazolidin-2-yl)methyl) phenyl) amino) propane-1,2-dione AM 8 were shown in Figure 4.
As for the third most active analogue AM1 and AM2 detailed protein–ligand interaction profile (PLI) demonstrated numerous key interactions with the catalytic activity of AChE enzymes such as GLU4, ASP5, ALA6,GLU7,LEU8,LEU9, VAL10,THR11, VAL12, ARG13,GLU4.ASP5,ALA6,GLU7, LEU8,LEU9,VAL10,THR11, VAL12,ARG13,GLY14, GLY15,ARG16LEU17,ARG1 (AM1) and GLU4,ASP5,ALA6,GLU7, LEU8,LEU9, VAL10,THR11, VAL12, ARG13,GLU4.ASP5, ALA6, GLU7,LEU8, LEU9,VAL10, THR11,VAL12,ARG13, GLY14,GLY15,ARG16, LEU17, ARG1 (AM2).
Table 4: Prediction of Name of Amino acid; Hydrophobicity; pKa; Avg. Isotropic Displacement and Molecular Docking score for Oxadiazole Derivatives AM1 to AM10 given by PDB: 7E3H
| Derivative | Name of Amino acid | Hydrophobicity | pKa | Avg. Isotropic Displacement | Molecular Docking |
| AM-1 | GLU4ASP5ALA6
GLU7 LEU8 LEU9 VAL10 THR11 VAL12 ARG13 |
-3.5-3.51.8
-3.5 3.8 3.8 4.2 -0.7 4.2 -4.5 |
4.312 | 68.78854.57848.32
55.85 43.52 43.291 37.848 38.868 40.941 62.77 |
-11.5 |
| AM-2 | GLU4ASP5ALA6
GLU7 LEU8 LEU9 VAL10 THR11 VAL12 ARG13 GLY14 GLY15 ARG16 LEU17 ARG18 |
-3.5-3.51.8
-3.5 3.8 3.8 4.2 -0.7 4.2 -4.5 -0.4 -0.4 -4.5 3.8 -4.5 |
4.33.94.3
12 12 12 |
68.78854.57848.32
55.85 43.52 43.291 37.848 38.868 40.941 62.77 43.211 40.129 51.628 37.528 42.052 |
-10.9 |
| AM-3 | GLU4ASP5ALA6
GLU7 LEU8 LEU9 VAL10 THR11 VAL12 ARG13 GLY14 |
-3.5-3.51.8
-3.5 3.8 3.8 4.2 -0.7 4.2 -4.5 -4.0 |
4.33.94.3
12 |
68.78854.57848.32
55.85 43.52 43.291 37.848 38.868 40.941 62.77 43.211 |
-10.5 |
| AM-4 | GLU4ASP5ALA6
GLU7 LEU8 LEU9 VAL10 THR11 VAL12 ARG13 |
-3.5-3.51.8
-3.5 3.8 3.8 4.2 -0.7 4.2 -4.5 |
4.33.94.3
12 |
68.78854.57848.32
55.85 43.52 43.291 37.848 38.868 40.941 62.77 |
-11.7 |
| AM-5 | GLU4ASP5ALA6
GLU7 LEU8 LEU9 VAL10 THR11 VAL12 ARG13 |
-3.5-3.51.8
-3.5 3.8 3.8 4.2 -0.7 4.2 -4.5 |
4.33.94.3
12 |
68.78854.57848.32
55.85 43.52 43.291 37.848 38.868 40.941 62.77 |
-11.5 |
| AM-6 | GLU4ASP5ALA6
GLU7 LEU8 LEU9 VAL10 THR11 VAL12 ARG13 |
-3.5-3.51.8
-3.5 3.8 3.8 4.2 -0.7 4.2 -4.5 |
4.33.94.3
12 |
68.78854.57848.32
55.85 43.52 43.291 37.848 38.868 40.941 62.77 |
-11.0 |
| AM-7 | GLU4ASP5ALA6
GLU7 LEU8 LEU9 VAL10 THR11 VAL12 ARG13 |
-3.5-3.51.8
-3.5 3.8 3.8 4.2 -0.7 4.2 -4.5 |
3.94.312 | 68.78854.57848.32
55.85 43.52 43.291 37.848 38.868 40.941 62.77 |
-10.1 |
| AM-8 | GLU4ASP5ALA6
GLU7 LEU8 LEU9 VAL10 THR11 VAL12 ARG13 |
-3.5-3.51.8
-3.5 3.8 3.8 4.2 -0.7 4.2 -4.5 |
4.33.94.3
12 |
54.57848.3255.85
43.52 43.291 37.848 38.868 40.941 62.77 62.78 |
-11.4 |
| AM-9 | GLU4ASP5ALA6
GLU7 LEU8 LEU9 VAL10 THR11 VAL12 ARG13 |
-3.5-3.51.8
-3.5 3.8 3.8 4.2 -0.7 4.2 -4.5 |
4.33.94.3
12 |
68.78854.57848.32
55.85 43.52 43.291 37.848 38.868 40.941 62.77 |
-7.6 |
| AM-10 | GLU4ASP5ALA6
GLU7 LEU8 LEU9 VAL10 THR11 VAL12 ARG13 GLY14 GLY15 ARG16 LEU17 ARG18 |
-3.5-3.51.8
-3.5 3.8 3.8 4.2 -0.7 4.2 -4.5 -0.4 -0.4 -4.5 3.8 -4.5 |
4.33.94.3
12 12 12 |
68.78854.57848.32
55.85 43.52 43.291 37.848 38.868 40.941 62.77 43.211 40.129 51.628 37.528 42.052 |
-9.2 |
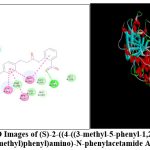 |
Figure 1: 2D and 3D Images of (S)-2-((4-((3-methyl-5-phenyl-1,2,4-oxadiazol-2(3H)-yl)methyl)phenyl)amino)-N-phenylacetamide AM1 |
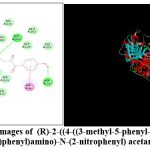 |
Figure 2: 2D and 3D Images of (R)-2-((4-((3-methyl-5-phenyl-1,2,4-oxadiazol-2(3H)-yl)methyl)phenyl)amino)-N-(2-nitrophenyl) acetamide AM2 |
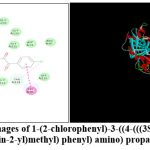 |
Figure 3: 2D and 3D Images of 1-(2-chlorophenyl)-3-((4-(((3S,5R)-3-methyl-5-phenyl-1,2,4-oxadiazolidin-2-yl)methyl) phenyl) amino) propane-1,2-dione AM 8 |
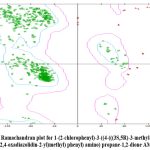 |
Figure 4: Ramachandran plot for 1-(2-chlorophenyl)-3-((4-(((3S,5R)-3-methyl-5-phenyl-1,2,4-oxadiazolidin-2-yl)methyl) phenyl) amino) propane-1,2-dione AM 8 |
Conclusion
The results were the same as those obtained by traditional methods when oxadiazole derivatives were docked into the binding site PDB:7E3H of the AChE protein molecules. Docking energy measurements showed a moderate yet advantageous interaction with acetylcholinesterase. The amino acid residues GLU4, ASP5, ALA6, GLU7, LEU8, LEU9, VAL10, THR11, VAL12, ARG13, GLU4.ASP5, and ALA6 are all strongly impacted by our tested substances. The reference standard donepezil and this binding mode are nearly identical. Therefore, it was believed that certain amino acid residues were essential for the interaction with inhibitors of cholinesterase. These drugs will have safer pharmacokinetic and toxicological characteristics, based on ADMET prediction data. The results of the study suggest that these compounds may have strong anti-Alzheimer properties and can be utilized to stop the progression of Alzheimer’s disease; however, further research is required before (S)-2-((4-((3-methyl-5-phenyl-1,2,4-oxadiazol-2(3H)-yl)methyl)phenyl)amino)-N-phenyl acetamide AM1 and (R)-2-((4-((3-methyl-5-phenyl-1,2,4-oxadiazol-2(3H)-yl)methyl)phenyl)amino)-N-(2-nitrophenyl) acetamide AM2 and 1-(2-chlorophenyl)-3-((4-(((3S,5R)-3-methyl-5-phenyl-1,2,4-oxadiazolidin-2-yl)methyl) phenyl) amino) propane-1,2-dione AM 8 can be considered as viable treatment for Alzheimer’s disease. ADMET predictions suggested a low risk of toxicity, supporting their potential as safe and effective Alzheimer’s disease candidates. Additionally, hydrogen bond analysis reinforced the stability of the protein–ligand interactions, highlighting the significance of these non-covalent forces in complex formation. The computational insights generated from this study provide a strong foundation for further synthetic, in vitro, and in vivo investigations aimed at validating their therapeutic efficacy. Ultimately, such advancements could contribute significantly to global Alzheimer’s disease control and eradication strategies.
Acknowledgement
The authors are thankful to Pravara Rural College of Pharmacy, Pravaranagar.
Funding Sources
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflict of Interest
The authors do not have any conflict of interest.
Data Availability Statement
This statement does not apply to this article.
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Clinical Trial Registration
This research does not involve any clinical trials.
Permission to Reproduce Material from Other Sources
Not Applicable
Author Contributions
Soniya Vaibhav Katore: In silico ADMET (Absorption, Distribution, Metabolism, Excretion)
Ajay Sanjay Mule: In silico ADMET (Absorption, Distribution, Metabolism, Excretion)
Neha Vilas Pagare: Drawing of Tafenoquine drug derivatives
Namrata Rajesh Adhav: SWISS ADMET Software Data
Harshali Narayan Anap: Docking images of 2D and 3D by Discovery Studio 2021
Rohit Jaysing Bhor8: The active amino residues, bond length, bond category, bond type, ligand energies, 1-Click Docking software and Research drafting
References
- Anquan Li, Jun Zhao, Chen Zhang, Qiuxia Jiang, Baofu Zhu, Hua Cao. Lewis Acid-Promoted Three-Component Cyclization for the Construction of Functionalized Oxazoles. The Journal of Organic Chemistry 2023, 88 (1), 27-38. https://doi.org/10.1021/acs.joc.2c01432
CrossRef - Hejun An, Shaoyu Mai, Qingqing Xuan, Yao Zhou, Qiuling Song. Gold-Catalyzed Radical-Involved Intramolecular Cyclization of Internal N-Propargylamides for the Construction of 5-Oxazole Ketones. The Journal of Organic Chemistry 2019, 84 (1) , 401-408. https://doi.org/10.1021/acs.joc.8b02334
CrossRef - Kate Lauder, Anita Toscani, Nicolò Scalacci, and Daniele Castagnolo . Synthesis and Reactivity of Propargylamines in Organic Chemistry. Chemical Reviews 2017, 117 (24) , 14091-14200. https://doi.org/10.1021/acs.chemrev.7b00343
CrossRef - Sho Suzuki and Akio Saito . Single-Step Synthesis of Iodinated Oxazoles from N-Propargyl Amides Mediated by I2/Iodosylbenzene/Trimethylsilyl Trifluoromethanesulfonate Systems. The Journal of Organic Chemistry 2017, 82 (22) , 11859-11864. https://doi.org/10.1021/acs.joc.7b01563
CrossRef - Savannah Jordan, Samuel A. Starks, Michael F. Whatley, and Mark Turlington . Highly Stereoselective Synthesis of Terminal Chloro-Substituted Propargylamines and Further Functionalization. Organic Letters 2015, 17 (19) , 4842-4845. https://doi.org/10.1021/acs.orglett.5b02408
CrossRef - Ádám Sinai, Dóra Vangel, Tamás Gáti, Petra Bombicz, and Zoltán Novák . Utilization of Copper-Catalyzed Carboarylation–Ring Closure for the Synthesis of New Oxazoline Derivatives. Organic Letters 2015, 17 (17) , 4136-4139. https://doi.org/10.1021/acs.orglett.5b01860
CrossRef - Helene Chachignon, Nicolò Scalacci, Elena Petricci, and Daniele Castagnolo . Synthesis of 1,2,3-Substituted Pyrroles from Propargylamines via a One-Pot Tandem Enyne Cross Metathesis–Cyclization Reaction. The Journal of Organic Chemistry 2015, 80 (10) , 5287-5295. https://doi.org/10.1021/acs.joc.5b00222
CrossRef - Marcos A. P. Martins, Clarissa P. Frizzo, Aniele Z. Tier, Dayse N. Moreira, Nilo Zanatta, and Helio G. Bonacorso . Update 1 of: Ionic Liquids in Heterocyclic Synthesis. Chemical Reviews 2014, 114 (20) , PR1-PR70. https://doi.org/10.1021/cr500106x
CrossRef - Yancheng Hu, Ruxia Yi, Chunxiang Wang, Xiaoyi Xin, Fan Wu, and Boshun Wan . From Propargylamides to Oxazole Derivatives: NIS-Mediated Cyclization and Further Oxidation by Dioxygen. The Journal of Organic Chemistry 2014, 79 (7) , 3052-3059. https://doi.org/10.1021/jo5001719
CrossRef - Chiara Pizzo and S. Graciela Mahler . Synthesis of Selenazoles by in Situ Cycloisomerization of Propargyl Selenoamides Using Oxygen–Selenium Exchange Reaction. The Journal of Organic Chemistry 2014, 79 (4) , 1856-1860. https://doi.org/10.1021/jo402661b
CrossRef - Yancheng Hu, Ruxia Yi, Fan Wu, and Boshun Wan . Synthesis of Functionalized Oxazoles via Silver-Catalyzed Cyclization of Propargylamides and Allenylamides. The Journal of Organic Chemistry 2013, 78 (15) , 7714-7726. https://doi.org/10.1021/jo401330t
CrossRef - Xinzhang Yu, Xiaoyi Xin, Boshun Wan, and Xingwei Li . Base-Catalyzed Cyclization of N-Sulfonyl Propargylamides to Sulfonylmethyl-Substituted Oxazoles via Sulfonyl Migration. The Journal of Organic Chemistry 2013, 78 (10) , 4895-4904. https://doi.org/10.1021/jo4004635
CrossRef - Anton V. Gulevich, Alexander S. Dudnik, Natalia Chernyak, and Vladimir Gevorgyan . Transition Metal-Mediated Synthesis of Monocyclic Aromatic Heterocycles. Chemical Reviews 2013, 113 (5) , 3084-3213. https://doi.org/10.1021/cr300333u
CrossRef - Wei Zhou, Chen Xie, Jianlin Han, and Yi Pan . Catalyst-Free Intramolecular Oxidative Cyclization of N-Allylbenzamides: A New Route to 2,5-Substituted Oxazoles. Organic Letters 2012, 14 (18) , 4766-4769. https://doi.org/10.1021/ol302031z
CrossRef - A. Stephen K. Hashmi, Maria Camila Blanco Jaimes, Andreas M. Schuster, and Frank Rominger . From Propargylic Amides to Functionalized Oxazoles: Domino Gold Catalysis/Oxidation by Dioxygen. The Journal of Organic Chemistry 2012, 77 (15) , 6394-6408. https://doi.org/10.1021/jo301288w
CrossRef - Dougal J. Ritson, Christian Spiteri, and John E. Moses . A Silver-Mediated One-Step Synthesis of Oxazoles. The Journal of Organic Chemistry 2011, 76 (9) , 3519-3522. https://doi.org/10.1021/ jo1025332
CrossRef - Barry M. Trost, Jean-Philip Lumb, and Joseph M. Azzarelli . An Atom-Economic Synthesis of Nitrogen Heterocycles from Alkynes. Journal of the American Chemical Society 2011, 133 (4) , 740-743. https://doi.org/10.1021/ja110117g
CrossRef - Simon Doherty, Julian G. Knight, A. Stephen K. Hashmi, Catherine H. Smyth, Nicholas A. B. Ward, Katharine J. Robson, Sophie Tweedley, Ross W. Harrington, and William Clegg . Efficient Cycloisomerization of Propargyl Amides by Electrophilic Gold(I) Complexes of KITPHOS Monophosphines: A Comparative Study. Organometallics 2010, 29 (18) , 4139-4147. https://doi.org/10.1021/om1006769
CrossRef - Nimesh C. Misra and H. Ila . 4-Bis(methylthio)methylene-2-phenyloxazol-5-one: Versatile Template for Synthesis of 2-Phenyl-4,5-functionalized Oxazoles. The Journal of Organic Chemistry 2010, 75 (15) , 5195-5202. https://doi.org/10.1021/jo100941f
CrossRef - Jianmin Zhang, Elena A. Polishchuk, Jie Chen and Marco A. Ciufolini . Development of an Oxazole Conjunctive Reagent and Application to the Total Synthesis of Siphonazoles. The Journal of Organic Chemistry 2009, 74 (23) , 9140-9151. https://doi.org/10.1021/jo9018705
CrossRef - Marcos A. P. Martins, Clarissa P. Frizzo, Dayse N. Moreira, Lilian Buriol and Pablo Machado. Solvent-Free Heterocyclic Synthesis. Chemical Reviews 2009, 109 (9) , 4140-4182. https://doi.org/10.1021/ cr9001098
CrossRef - Jianmin Zhang and Marco A. Ciufolini. Total Synthesis of Siphonazoles by the Use of a Conjunctive Oxazole Building Block. Organic Letters 2009, 11 (11) , 2389-2392. https://doi.org/10.1021/ ol900455m
CrossRef - Ying-ming Pan, Fei-jian Zheng, Hai-xin Lin and Zhuang-ping Zhan. Brønsted Acid-Catalyzed Propargylation/Cycloisomerization Tandem Reaction: One-Pot Synthesis of Substituted Oxazoles from Propargylic Alcohols and Amides. The Journal of Organic Chemistry 2009, 74 (8) , 3148-3151. https://doi.org/10.1021/jo8027533
CrossRef - Marcos A. P. Martins, Clarissa P. Frizzo, Dayse N. Moreira, Nilo Zanatta and Helio G. Bonacorso. Ionic Liquids in Heterocyclic Synthesis. Chemical Reviews 2008, 108 (6) , 2015-2050. https://doi.org/10.1021/cr078399y
CrossRef - Quansheng Shen, Wen Huang, Jialiang Wang and Xigeng Zhou. Ln[N(SiMe3)2]3/RNH2 Catalyzed Monoaddition of Terminal Alkynes to Nitriles: A Novel and Concise Access to the Synthesis of Ynones. Organometallics 2008, 27 (2) , 301-303. https://doi.org/10.1021/om700891k
CrossRef - Filip Paulsen, Sebastian Clementson, Henrik von Wachenfeldt, Michał Antoszczak, Simon Fridolf, Daniel Strand. Synthesis of Siphonazole B Through Domino Cycloisomerization‐Oxazolonium Ion Rearrangements. Chemistry – A European Journal 2025, 31 (40) https://doi.org/10.1002/ chem.202501394
CrossRef - Nicholas Jankowski, Norbert Krause. Efficient and Stereoselective Synthesis of ( Z )‐2‐Oxazolines by Transition Metal‐Free Cycloisomerization of Internal Propargylic Amides in Hexafluoroisopropanol (HFIP). Asian Journal of Organic Chemistry 2025, 33 https://doi.org/10.1002/ajoc.202500262
CrossRef - Kensuke Kiyokawa, Kazuki Kawanaka, Satoshi Minakata. Amino‐λ 3 ‐iodane‐Enabled Electrophilic Amination of Arylboronic Acid Derivatives. Angewandte Chemie 2024, 136 (12) https://doi.org/10.1002/ange.202319048
CrossRef - Kensuke Kiyokawa, Kazuki Kawanaka, Satoshi Minakata. Amino‐λ 3 ‐iodane‐Enabled Electrophilic Amination of Arylboronic Acid Derivatives. Angewandte Chemie International Edition 2024, 63 (12) https://doi.org/10.1002/anie.202319048
CrossRef - Xu Chen, Xinrui Li, Wenbo He, Miao Wang, Ang Gao, Liping Tong, Shun Guo, Huaiyu Wang, Guoqing Pan. Rational multivalency construction enables bactericidal effect amplification and dynamic biomaterial design. The Innovation 2023, 4 (5) , 100483. https://doi.org/10.1016/j.xinn.2023.100483
CrossRef - Leila Kamrani Tamardash, Mohammad Bakherad, Ali Keivanloo, Behrouz Notash. An Efficient Synthesis of New Pyrazole‐Linked Oxazoles via Sonogashira Coupling Reaction. ChemistrySelect 2023, 8 (18) https://doi.org/10.1002/slct.202204985
CrossRef
Abbreviations List
mg/kg: Milligram/ kilograms;
Sec: seconds;
Kcal: kilocalorie;
Mol.Wt: Molecular Weight;
Gm: Gram;
LEU: Leucine;
THR: Threonine;
ALA: Alanine;
MET: Methionine;
PHE: Phenylalanine;
WHO: World health association;
Log P: Partition coefficient
SDF: format: Structure Data File
CaSR: Calcium-Sensing Receptor
Accepted on: 12-11-2025
Second Review by: Dr. Saeed Kewedar
Final Approval by: Dr. Jagdish Chandra Joshi









