Isolation, Identification and Media Optimization of Gut Bacteria Species from Danio rerio (zebra fish) Against the Spodoptera litura Larvae
Department of Microbiology, AVS College of Arts and Science (Autonomous), Tamil Nadu, India.
Corresponding Author Email: ssmmicro@gmail.com
DOI : http://dx.doi.org/10.13005/bbra/3474
ABSTRACT:Spodoptera litura control is genuinely expected to replace the deadly compound pesticides; the disadvantages of the invention pesticides are issues, an effect on human and animals, environmental pollution and killing the non-target insect and highly resistance response to an insect pest. This situation used as ecologically agreeable, moved nearer to the bio pesticide. In this study, four strains were isolated from the zebra fish gut, among the two strains the highest bacterial growth was observed in ZBF-2 Bacillus subtilis, ZBF-4 Bacillus cereus. In view of biochemical, physiological and molecular characterization the isolated zebra fish gut was distinguished as having a place with Bacillaceae family. Phylogenetic investigation of the 16S rDNA sequence of zebra fish gut bacteria are ZBF-2 Bacillus subtilis, ZBF-4 Bacillus cereus. In Danio rerio gut bacterial optimal growth conditions were tested with different media optimal growth level with ZBF-2 and ZBF-4 showed the highest growth rates under optimized conditions, including glucose as the key carbon source, peptone as the nitrogen source, pH levels of 4 to 7 and a temperature of 35ºC at 72hrs .The two strains, ZBF-2Bacillus subtilis and ZBF-4Bacillus cereus, were selected for 16S rRNA sequencing, and the resulting sequences identified as and get the accession number of Bacillus subtilis (PV297978) and Bacillus cereus (PV300533). These optimized conditions fostered the growth of gut microbes that are important for the secondary metabolites enhance for the bioactive compounds may be responsible for the insecticidal properties. And we further tested, with the isolated bacterial cell free supernatant of ZBF-2 and ZBF-4 produced 76-80% mortality at 48hrs in Spodopteralitura larvae. This study shows that, the symbiotic bacteria ZBF-2 Bacillus subtilis, ZBF-4 Bacillus cereus are virulent, and their secondary metabolites are also toxic and can be utilized in pest control programs.
KEYWORDS:Bacillus cereus; Bacillus subtilis;Gut bacteria; Larval morality; Optimization; Secondary metabolites
Introduction
Trillions of bacteria make up the vertebrate gut microbiome, which is essential for digestion, energy acquisition, nutrient absorption, and host health.1 Fish development, reproduction, and ecological interactions are all greatly impacted by gut microbes.2 Since fish larvae are initially sterile and pick up microorganisms from their surroundings, the microbiome is especially crucial during the early stages of development.3 Environmental factors such as salinity and temperature largely shape gut microbial communities.4 Additionally, variations in gut microbiota among individuals of the same species often reflect differences in environmental conditions.5 Exploring microbiome changes in wild hosts can provide insights into ecological adaptability. Fish gut microflora includes aerobic, facultative anaerobic and obligate anaerobic bacteria. Similar to humans, microbial composition changes with age, nutrition, and environmental conditions.6 Bacterial colonization occurs on the skin, gills, and intestine with zebrafish (Danio rerio) actively ingesting water as early as four days post-fertilization (dpf), allowing microbes to enter their digestive tract.7By adulthood, zebrafish gut microbiomes are well-established, with high bacterial abundance. Zebrafish, a freshwater omnivorous teleost fish native to South Asia, has become a widely used model organism in biomedical research.8 Bacillus species are known to produce a wide range of bioactive compounds with potential applications in medicine, agriculture, and industry. Bacillus megaterium produces megacin, a bacteriocin with activity against Gram-positive bacteria 9. Bacillus cereus produces cereulide, a toxin that can cause food poisoning,10bacilysin, an antibiotic with activity against Gram-positive bacteria.11
Bacillus subtilis produce’s bacitracin, an antibiotic with activity against Gram-positive bacteria 12and surfactin, a lipopeptide with antimicrobial and antiviral activities13. Bacillus spp. Produced by bacillomycin, a lipopeptide with antimicrobial and antifungal activities of bacillus sps14and fengycin, a lipopeptide with antimicrobial and antifungal activities 15. These bioactive compounds have potential applications in various fields, including medicine, agriculture, and industry. Only a minor fraction of bacterial populations can grow on standardlaboratory media,15 In fish, skin microbiota is estimated to have less than 0.01% culturable bacteria.16 Similar discrepancies have been observed in human gut microbiota, where cultural methods overestimated to bacteria populations.17 Despite extensive studies on fish gut microbiota, research has predominantly relied on culturable bacteria, limiting the understanding of total microbial diversity.18 Traditional microbial identification methods based on phenotypic and biochemical characteristics are time-consuming and may not accurately reflect the actual composition of microbial communities.19
Spodoptera litura, commonly known as the tobacco caterpillar or armyworm, is a significant agricultural pest that causes substantial damage to various crops, including tobacco, cotton, and vegetables.20 This polyphagous pest is widely distributed across Asia, Africa, and Australia, and its management is a major challenge due to its rapid adaptation to insecticides and resistant crop varieties.21 The larvae of S. litura feed on plant leaves, causing defoliation and reducing crop yields.22 Effective management of S. litura requires an integrated approach, including cultural, biological, and chemical controls. Chemical insecticides, such as pyrethroids, organophosphates, and carbamates, have been widely used to control S. litura populations 21 However, the excessive and indiscriminate use of these insecticides has led to the development of resistance, environmental pollution, and harm to human health.22 Some of the chemical insecticides commonly used against S. liturainclude Cypermethrin, Chlorpyrifos, Carbofuran. S litura, is a devastating agricultural pest that causes significant damage to various crops, including tobacco, cotton and vegetables. To manage this pest, chemical insecticides have been widely used, providing quick and effective control. However, the excessive and indiscriminate use of chemical insecticides has raised concerns about their environmental impact, human health risks, and the development of pesticide-resistant to S. liturapopulations. In recent years, researchers have explored alternative approaches, including the use of beneficial microorganisms, such as gut bacteria from zebrafish (Danio rerio), to control insect pests. Studies have shown that zebrafish gut bacteria, including Bacillus megaterium, Bacillus cereus, Bacillus subtilis, and other Bacillus species, possess insecticidal properties against various insect pests, including S. litura.23These bacteria produce bioactive compounds, such as lipopeptides, bacteriocins, and enzymes, which can be used to control insect pests.24,25 Injecting S. litura with these beneficial bacteria or their bioactive compounds offers a promising alternative to chemical insecticides. This approach has been shown to be effective in controlling S. litura populations while minimizing harm to the environment and human health.26Therefore, the full investigation and exploitation of gut bacteria can be significant for biological control of insect pests. The present study focuses on the isolation, identification and culturing of two strains of gut bacteria from Danio rerio againstinsecticidal efficacy on S. litura larvae.
Materials and Method
Sample Collection, Isolation of gut microbes
Zebrafish (Danio rerio) were collected from Edward Aquarium lies on the latitude of 11.6607636º, and longitude of 78.1504985º Salem, and acclimated for 24 hours before sample collection. They were anesthetized using tricaine methane sulfonate (MS-222) and humanely sacrificed. The gastrointestinal tract (GIT) was carefully dissected, and gut samples were stored at -80°C for further analysis. Gut microbes were isolated by serial dilution in sterile double-distilled water and cultured on Nutrient Agar at 28 °C for 24-48 hours. Pure cultures were obtained using the streak-plate method and stored at -80 °C. Further, these colonies were collected and sub-cultured in10ml of Nutrient broth. Description of samples used in this study as shown in (Table.1).
Morphological analysis of gut microbes
Bacterial isolates were sub-cultured on nutrient agar plates. After 24 hrs of incubation, the characteristic gut bacterial isolates were screened. Selected bacterial colonies were identified as Gram negative or Gram-positive microscopes organisms and biochemical characteristics of four gut bacteria were performed using KB001- HilMViC biochemical test kit.
Phylogenetic analysis of Bacterial isolates
Genomic DNA was isolated from the bacterial sample using PAR Life science and research bacterial genomic DNA isolation kit (PAR Life science and research PVT Ltd. Trichy India). 16S rRNA primers used in PCR reaction. PCR mixture containing 1µl DNA, 400 ng forward primer, 400ng reverse primer, 4µl dNTPs, 10µl 10X Taq DNA polymerase assay buffer, 1µl Taq DNA polymerase enzyme and distilled water. The total reaction volume was 100 µl. PCR amplification was carried out under following conditions step-I initial denaturing for 5mins at 94ᵒC, denaturing at 94ᵒC for the 30sec, step-ii annealing at 55˚C for 30s and step-iii. extension at 72˚C for 40 sec for 35 cycles. PCR product was obtained and gel was eluted using agarose gel extraction kit. After elution, the PCR products were sequenced using ABI sequencing machine (ABI 3500 XL Genetic Analyzer). 10µl of sequencing mixture contains 4µl Big Dye Terminator Ready Reaction Mix, 1µl Template (100 ng/µl), 2µl primer (10pmol/λ) and 3µl Milli Q water. PCR amplification of sequencing mixture was carried out with the following conditions. 25 cycles of amplification were started after initial denaturing at 96˚C for 1min, denaturing at 96˚C for 10sec, extension at 50˚C for 5sec and elongation at 60˚C for 4mins. DNA concentration can be determined by both UV spectrophotometer and quantitative analysis on Agarose gel. The gel documentation system with UV- light and images were captured. Data analysis was performed by sequencing Analysis_ version 5.4 software. Phylogenetic tree was constructed using MEGA 6 software, according to the Neighbour-joining tree method.
Growth of symbiotic bacteria at various culture media
The growth conditions of recovered isolates four bacteria strains were determined at different culture media: nutrient broth, LB broth, Tryptic soy broth, Muller Hinton broth and MacConkey broth. A single primary colony was innoculated in 150 ml conical flask containing 100ml of various culture media and incubated between 0 and 96 hrs. After incubation, the growth of the bacterial culture was determined at optical density (OD) of 600 nm.27
Growth of symbiotic bacteria at various pH
The growth conditions of recovered isolates four bacterial strains were determined at different pH. A single primary colony was inoculated in 100 ml conical flask containing 50ml of nutrient broth and incubated at the pH range of 5.0, 6.0, 7.0, 8.0 and 9.0 from 0 to 96 hrs. After incubation, the growth of the bacterial culture was determined at optical density of 600 nm.27
Growth of symbiotic bacteria at various Temperatures
The growth conditions of recovered isolates four bacterial strainswere determined at different temperatures. A single primary colony was innoculated in 100 ml conical flask containing 50ml of nutrient broth and incubated at the temperature 10 °C, 20 °C, 25°C, 30 °C, 35°C and 40°C for 48 hrs. After incubation, the growth of the bacterial culture was determined by optical density at 600 nm.27
Growth of symbiotic bacteria at various Carbon source
The growth conditions of recovered isolates four bacterial strains were determined at different carbon source: Glucose, Sucrose, Lactose, Starch and Fructose. As for the control, the commercialized media were used. A single primary colony was inoculated in 150 ml conical flask containing 100ml of nutrient broth. To the nutrient broth, 0.5g of above-mentioned carbon source was added separately and incubated from 0 to 96 hrs. After incubation, the growth of the bacterial culture was determined by optical density at 600 nm.
Growth of symbiotic bacteria at various Nitrogen source
The growth conditions of recovered isolates four bacterial strainswere determined at different nitrogen source: beef extract, peptone, yeast extract, soyabean and potassium nitrate. A single primary colony was inoculated in 150 ml conical flask containing 100ml of nutrient broth. To the nutrient broth, 0.5g of above-mentioned nitrogen source were added and incubated between 0 to 96 hrs. After incubation, the growth of the bacterial culture was determined by optical density at 600 nm.
Extraction of secondary metabolites from bacterial supernatant
The grown cultures were centrifuged at 10,000 rpm for 15 min after incubation, and the collected supernatants were filtered through 0.22 m membrane filter to remove the debris from pellets. The extract was concentrated, suspended in DMSO (Dimethyl sulfoxide), and stored at -20°C.28
Larval bioassay using crude bacterial metabolite of ZBF-2 and ZBF-4
From the broth culture, the bacterial culture of Bacillus subtilis (ZBF-2) and Bacillus cereus (ZBF-4)1 mg of secondary metabolite were extracted and dissolved in 1 ml of (DMSO) dimethyl sulfoxide (1mg/ml). The bacterial secondary metabolites were diluted to various concentrations like as 10-4, 10-5 and 10-6 ranging from 5, 10, and 20 µl/mL were injected into proleg of S. litura larvae. PBS (Phosphate Buffer Saline) were used as control. Mortality was recorded after 24 hrs of infection.29
Percentage of larval motility = Number of dead larvae / Number of total larvae × 100
Statistical Analysis
Statistical analysis was performed by using GraphPad Prism (version 5.1 San Diego, CA). The data are expressed as Mean ± SD. The significant difference between the Mean replicates was determined using Two-Way ANOVA.
Results
Sample Collection, Isolation of gut microbes
In order to isolate the bacillus species, four different cultures were isolated from the gut of danio rerio (Zebra fish) as shown in (Figure.1). These sample of gut region of Danio rerio are crushed using sterile motor and pestle and then serial dilution should be performed and then streaked on nutrient agar medium (NA). The different types of bacterial colonies on nutrient agar medium should be streaked on another plate to isolate pure colonies of bacillus species (Table.1).
 |
Figure 1: Sample collection and isolation of gut microbes form danio rerio(zebra fish). |
Table 1: Description of samples used in this study
| Sample name | Common or host name | Environment | Sample
Site |
Water habitat | Collection date |
| ZBF2 | Zebrafish(Danio rerio, superorder Ostariophysi) | Conventional lab aquaculture | Whole intestinal contents | F | 20-Jan-25 |
| ZBF4 | Zebrafish(Danio rerio, superorder Ostariophysi) | Conventional lab aquaculture | Whole intestinal contents | F | 20-Jan-25 |
*F– Fresh WaterAbbreviations: F, freshwater
Identification of gut microbes
The bacterial isolates on nutrient agar medium (NA)are suspected as bacillus species and these were identified according to their morphological and cultural characteristics and bio-chemical test
Morphological and cultural characteristics
Gut Bacterial isolates from Danio rerio (zebra fish) such as ZBF-1Bacillus megatorium, ZBF-2 Bacillus subtilis, ZBF-3Bacillus sps, ZBF-4Bacillus cereus strains were isolated from nutrient agar plate and their morphology and characteristics should be identified by their color, odor, Growth condition etc.
Identification of gut bacterial microbes
Bacteria were isolated from the gut of Zebra Fish (Danio rerio) using NA plates and bromothymol blue stain as an indicator. Four strains, ZBF-1 Bacillus megatorium, ZBF-2 Bacillus subtilis, ZBF-3 Bacillus sps, ZBF-4 Bacillus cereus, were identified as Gram-positive rod-shaped bacteria based on morphological as shown in (Fig. 2, 3, 4,5). The gut bacterial isolates of Danio rerio results that the isolates of bacterial culture are gram positive bacillus. Colonies of these isolates are highly convex, yellow color colony, Opaque, frosted glass appearance with either rough or smooth surface. The phenotypic characteristics of gut bacterial isolates were shown in (Table.2).
Table 2: Phenotypic characteristics of isolated bacteria
| Characteristics | ZBF1(bacillus megaterium) | ZBF2(Bacillus subtilis) | ZBF3(bacillus
sps) |
ZBF4(Bacillus
Cereus) |
| Morphology | G (+) rods | G (+) rods | G (+) rods | G (+) rods |
| Bioluminescence | – | – | – | – |
| Bacterial pathogenicity | + | + | + | + |
| Growth on 28℃ | + | + | + | + |
| Growth on 37 | + | + | + | + |
| Absorption of bromothymol blue | + | + | + | + |
| Catalase | + | + | + | + |
| Spore Formation | + | + | + | + |
| Colony Morphology | Yellow-colour | Yellow-colour | Yellow-colour | White-colony |
Biochemical identification
The bacterial isolates of gut bacteria of four different strains, ZBF-1 Bacillus megatorium, ZBF-2 Bacillus subtilis, ZBF-3 Bacillus spsand ZBF-4 Bacillus cereus were examined for bio-chemical characterization as shown in (Figure.6). The results of gut bacteria of four different strains were showed as gram positive rod-shaped bacteria. The results are shown in (Table.3).
Table 3: Biochemical tests for the bacterial strains of ZBF1, ZBF2, ZBF3 and ZBF4
| S.No. | Biochemical test | ZBF1 | ZBF2 | ZBF3 | ZBF4 |
| 1. | Indole test | – | – | + | + |
| 2. | Methyel red | _ | _ | _ | _ |
| 3. | Vogousproskarus | _ | _ | _ | _ |
| 4. | L-arabitol | _ | _ | _ | _ |
| 5. | D-Cellobiose | _ | _ | _ | _ |
| 6. | Beta-Galactosidase | _ | _ | _ | _ |
| 7. | H2S Production | _ | _ | _ | _ |
| 8. | D- methyl –D- Glycosides | – | – | – | – |
| 9. | Glutamyl ArylamidasepNa | + | + | _ | + |
| 10. | D-Glucose | + | _ | + | _ |
| 11. | Dextrose | + | + | – | – |
| 12. | Glucose | + | _ | + | _ |
| 13. | Beta-Glucosidase | + | _ | + | _ |
| 14. | D-Maltose | _ | _ | _ | _ |
| 15. | D-Mannitol | _ | _ | _ | _ |
| 16. | D-Mannose | + | _ | + | _ |
| 17. | Beta-Xylosidase | _ | _ | _ | _ |
| 18. | Alpha- Methyl-D- Glycosides | _ | _ | _ | _ |
| 19. | L-Proline Aryl amidase | + | + | + | + |
| 23. | Urease | + | _ | + | _ |
| 24. | D-Sorbitol | + | _ | + | _ |
| 25. | Sucrose | _ | + | + | _ |
| 27. | D-Trehalose | _ | _ | _ | _ |
| 28. | Citrate-sodium | – | _ | – | _ |
| 29. | Malonate | _ | _ | + | + |
| 30. | Alpha-Glucosidase | + | + | – | – |
| 31. | Succinate Alkalisation | + | _ | + | _ |
| 32. | L- Arabinose | – | – | – | – |
| 33. | Galactosaminidase | + | + | _ | _ |
| 34. | Alpha-Galactosidase | _ | + | _ | + |
| 35. | Phosphatase | + | _ | + | _ |
| 36. | Glycine Aryl amidase | + | + | _ | _ |
| 37. | Ornithine Decarboxylase | + | + | _ | _ |
| 38. | Lysine Decarboxylase | _ | _ | _ | _ |
| 39. | L-Histidine assimilation | + | _ | + | + |
| 40. | Esculin | _ | + | _ | + |
| 41. | Beta-Glucuronidase | + | _ | + | _ |
| 42. | Rhamnose | – | _ | – | _ |
| 43. | Adonitol | + | + | – | – |
| 44. | Salicin | _ | + | + | _ |
| 46. | L-Lactate assimilation | + | + | + | – |
‘+’ positive and ‘– ‘negative
Optimization of isolation of bacterial species
Carbon source
The growth of isolates ZBF-1, ZBF-2, ZBF-3 and ZBF-4 was tested with various carbon sources like as (glucose, sucrose, lactose, starch, fructose) in nutrient broth. After incubation for 0 to 96 hours, glucose was found to be the most effective carbon source, promoting the highest bacterial growth levels at 72 hours compared to the control. Glucose was identified as the key component for enhancing secondary metabolite production in these gut bacteria. Effect of various carbon sources on the growth condition of gut bacterial culture shown in (Figure.7).
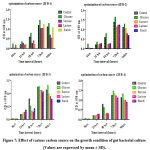 |
Figure 7: Effect of various carbon source on the growth condition of gut bacterial culture (Values are expressed by mean ± SD). |
Nitrogen source
The growth of isolates ZBF-1, ZBF-2, ZBF-3, and ZBF-4 was tested with different nitrogen sourceslike (beef extract, peptone, yeast extract, and soybean) in nutrient broth. After incubation for 0 to 96 hours, promoting the highest bacterial growth at 72 hours compared to the control. peptone showed the highest growth in the gut bacterial cultures ZBF-2, ZBF-4 as determined by optical density at 600 nm, making it the most effective nitrogen source for enhancing bacterial growth. Effect of various nitrogen source on the growth condition of gut bacteria culture as shown in (Figure.8).
 |
Figure 8: Effect of various nitrogen source on the growth condition of gut bacteria (Values are expressed by mean ± SD). |
Growth of gut bacteria at various Temperature
The growth of gut bacteria from isolates ZBF1, ZBF2, ZBF3 and ZBF4 was assessed at different temperatures (10°C, 20°C, 25°C, 30°C, 35°C and 40°C) using nutrient broth. After 24-48 hours of incubation, optical density at 600 nm was measured, with the highest growth observed at 35°C, indicating it as the optimal temperature for maximum growth ZBF and ZBF4 . Effect of various temperature on the growth condition of gut bacteria culture shown in (Figure.9).
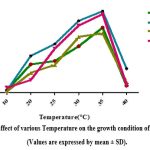 |
Figure 9: Effect of various Temperature on the growth condition of gut bacteria (Values are expressed by mean ± SD). |
Growth of gut bacteria at various pH
The growth of recovered isolates ZBF1, ZBF2, ZBF3, and ZBF4 was evaluated at different pH levels (5.0, 6.0, 7.0, 8.0, and 9.0) in nutrient broth. After incubating for 0 to 96 hours, the growth was measured by optical density at 600 nm. The highest growth was observed at pH 7, indicating it as the optimal pH for maximum bacterial growth. Effect of various pH on the growth condition of gut bacteria culture shown as (Figure.10).
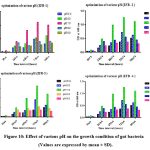 |
Figure 10: Effect of various pH on the growth condition of gut bacteria (Values are expressed by mean ± SD). |
Growth of gut bacteria at various culture media
The growth conditions of isolates ZBF1, ZBF2, ZBF3, and ZBF4 were tested in different culture media: nutrient broth, LB broth, Tryptic soy broth, Muller Hinton broth, and MacConkey broth. After incubating for 0 to 96 hours, the growth was measured by optical density at 600 nm. The highest growth was observed in nutrient broth, indicating it as the optimal medium for maximum bacterial growth. Gut bacterial growth on various media at different time intervals shown in (Figure.11).
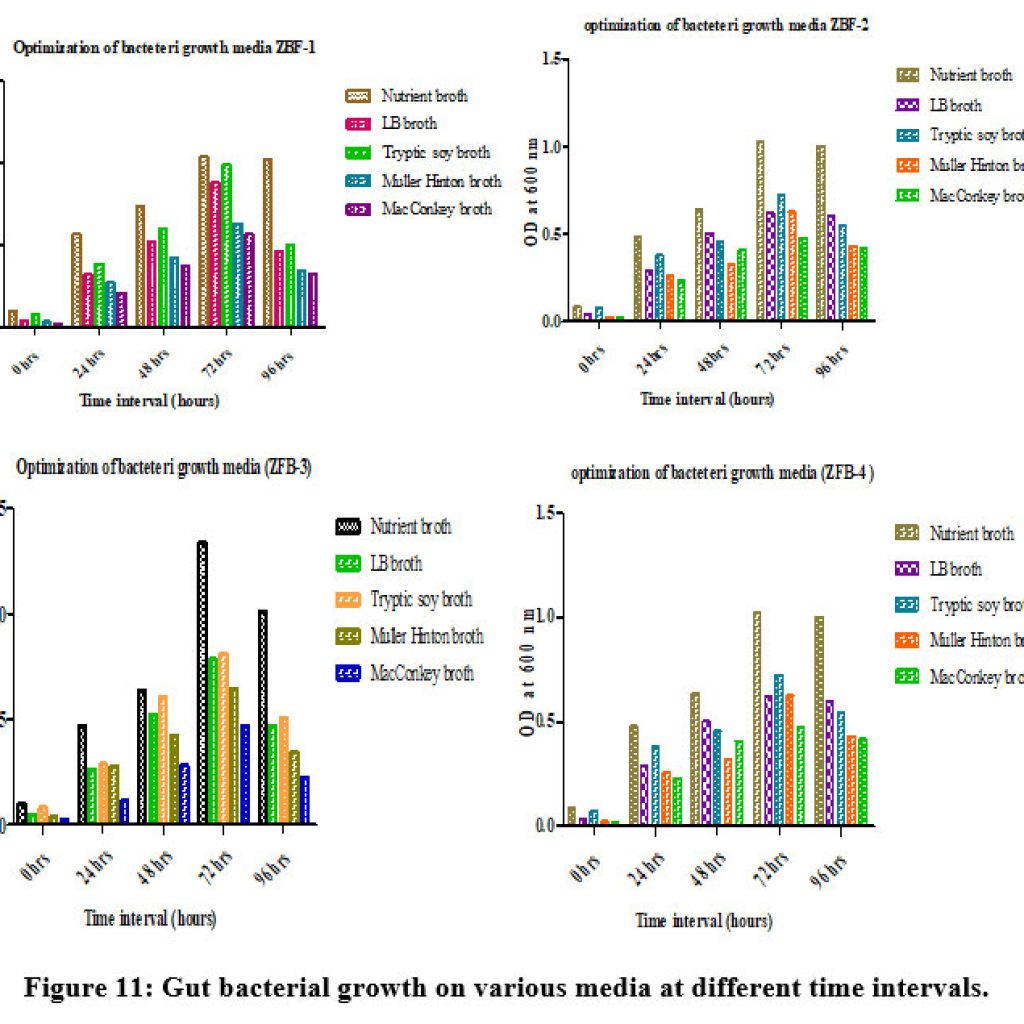 |
Figure 11: Gut bacterial growth on various media at different time intervals. |
Cell free supernatant extraction of ZBF-2 & ZBF-4 of gut bacteria
The cell-free supernatants from Bacillus subtilis ZBF-2 and Bacillus cereus ZBF-4, grown in nutrient broth, were extracted after centrifugation and sterile filtration. Bioactivity tests showed that Bacillus subtilis ZBF-2 exhibited strong antimicrobial activity against both Gram-positive and Gram-negative bacteria, while Bacillus cereus ZBF-4 showed moderate antibacterial effects, though less potent. Summarizing the cell-free supernatant extraction of Bacillus subtilis ZBF-2 and Bacillus cereus ZBF-4 (gut bacteria strains) and their biological activities as shown in (Table.4).
Table 4: summarizing the cell-free supernatant extraction of Bacillus subtilis ZBF-2 and Bacillus cereus ZBF-4 (gut bacteria strains) and their biological activities:
| Bacterial Strain | Incubation Time | Supernatant Collection Method | Bioactivity Assay | Larvicidal Activity | Feeding Inhibition |
| ZBF-2 (Gut Bacteria) | 24-48 hours | Centrifugation followed by sterile filtration (0.22 µm filter) | Spodoptera litura larvicidal bioassay | 65-75% mortality at high concentration | Reduced feeding observed |
| ZBF-4 (Gut Bacteria) | 24-48 hours | Centrifugation followed by sterile filtration (0.22 µm filter) | Spodoptera litura larvicidal bioassay | 50-60% mortality at high concentration | Mild feeding inhibition |
16S rRNA Sequencing Identification
Genomic DNA of two isolated gut bacterial strains was used for PCR amplification with 16S rDNA primers. The amplified fragments (1260-1500 bp) were analyzed on an agarose gel as shown in (Figure.12). BLAST analysis revealed 98% sequence similarity with Bacillus subtilis for ZBF-2 and Bacillus cereus for ZBF-4. A phylogenetic tree was constructed using the Neighbour-Joining method in Mega 6 software as shown in (Figure.13A, B). The 16S rDNA sequences were submitted to GenBank, with accession numbers PV297978 (Bacillus subtilis) and PV300533 (Bacillus cereus).
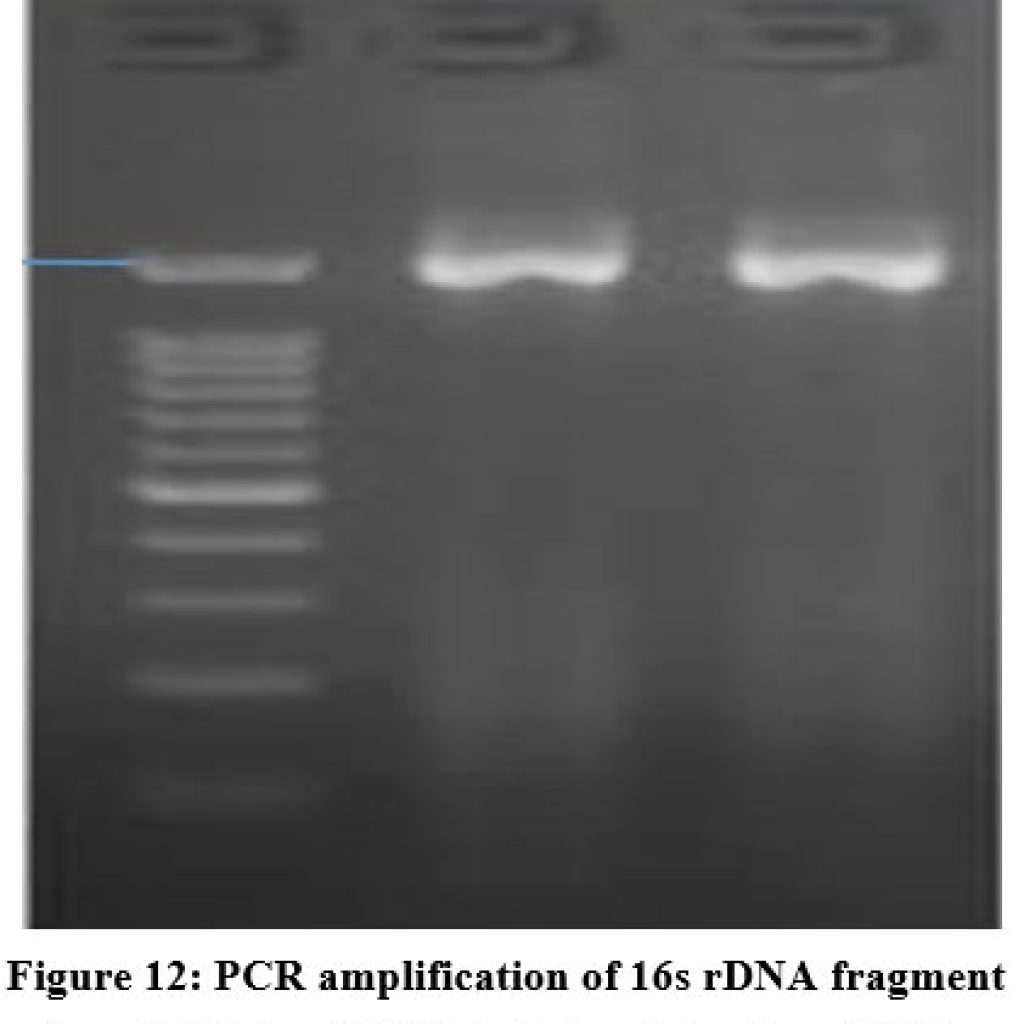 |
Figure 12: PCR amplification of 16s rDNA fragment from ZBF-1 and ZBF-4 strains. The size of PCR amplified product is ~1500bp. |
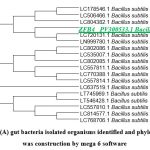 |
Figure 13: (A) gut bacteria isolated organisms identified and phylogentic tree was construction by mega 6 software |
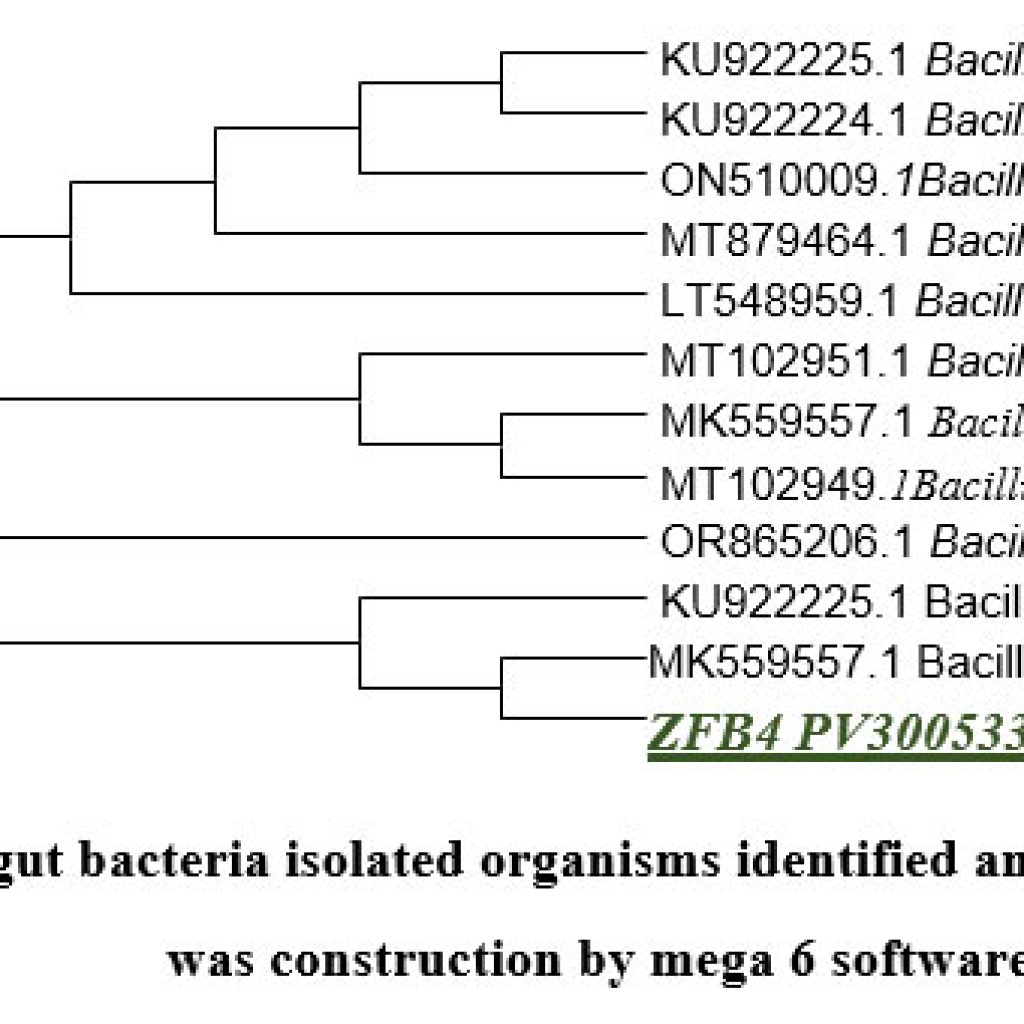 |
Figure 13: (B) gut bacteria isolated organisms identified and phylogentic tree was construction by mega 6 software. |
Secondary Metabolites from gut bacteria and larval toxicity
Virulence of gut bacteria Bacillus subtilis (ZBF-2) and Bacillus cereus (ZBF-4) and larval toxicity. From the broth culture, the extracted secondary metabolite of Bacillus subtilis (ZBF-2) and Bacillus cereus (ZBF-4) 1mg was dissolved in 1 ml of (DMSO)dimethyl sulfoxide(1 mg/ml). The larvicidal efficacy of secondary metabolite on S. litura was studied based on various concentration of secondary metabolite. After 24hrs of treatment the highest mortality was recorded in 10-4 concentration in all treatments. Mortality of S. litura depends on the concentration of secondary metabolites injected into larvae remaining concentration low levels mortality was observed (Figure.14,15).
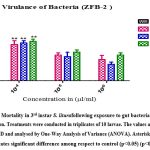 |
Figure 14: Mortality in 3rd instar S. liturafollowing exposure to gut bacteria in different concentration. Treatments were conducted in triplicates of 10 larvae. |
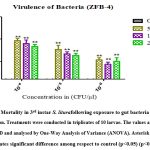 |
Figure 15: Mortality in 3rd instar S. liturafollowing exposure to gut bacteria in different concentration. Treatments were conducted in triplicates of 10 larvae. |
Discussion
Ray et al. 30 study reported that explored the influence of enzymatic activity and gut microbiota on fish digestion, emphasizing the role of factors like food type, digestive enzyme function, and exposure time. Their research highlighted that gut microbiota, including genera such as Bacillus, Enterobacteriaceae, and Acinetobacter, positively affects digestion by producing enzymes that aid in food breakdown. However, the varying digestive tract composition across fish species complicates determining the exact impact of this microbiota. In our study, we identified several Bacillus species, including Bacillus megaterium, Bacillus subtilis, Bacillus spp., and Bacillus cereus, which aligns with previous study of the 30 findings on the significance of Bacillus in fish digestion. Despite this, both studies face challenges in understanding the interactions of these species within the complex gut microbiota of different fish species. The gut microbiota is producing varieties of bioactive compounds such as proteins, enzymes, antibacterial and antifungal compounds were more responsible to the pharmaceutical and insecticidal agent of the microbiota organisms.
Dhayalan et al. 31 focused on isolating Bacillus thuringiensis from the gut of Systomussarana and optimizing culture parameters for maximal protease enzyme production using response surface methodology (RSM) and Box-Behnken design (BBD). Their study emphasized optimizing culture conditions to enhance enzyme yield, with a focus on protease production and its peptide profile. In contrast, our study evaluated factors such as carbon and nitrogen sources, temperature, pH, and different broth cultures to optimize bacterial growth rate of the parameters. The findings in our result suggest the main role of carbon, nitrogen, minerals, vitamin, temperature and pH source in promoting for the bacterial growth, with optimal growth at room temperature in especially in nutrient broth when compare to other broth. Our study, offers a broader examination of growth conditions. Despite the differences, both studies underscore the significance of tailored conditions for enhancing microbial activity, with implications for fish nutrition and enzymatic processes.
Mansooreh Jami et al. 32 studied the diversity of Actinobacteria in the gut microbiota of two freshwater fish species, Schizothoraxzarudnyi and Schizocyprisaltidorsalis. They cultured 277 isolates using three different agar media, identifying a range of Actinobacteria, primarily from the Streptomycetaceae family, along with other genera such as Saccharomonospora, Micromonospora, Nocardiopsis, and Arthrobacter. Their focus was on the diversity of secondary-metabolite biosynthetic genes (e.g., PKS I, NRPS, PhzE) and their antimicrobial activity against human and fish pathogens. In contrast, our study investigates the bacterial genus Bacillus, using PCR-based 16S rRNA gene sequencing and partial sequencing, which differs in both the bacterial genus and the methodological approach to the sequencing were identified and similarity to the BLAST tool alignment base pair of the nucleotide.
In present study, the identification of Bacillus subtilis and Bacillus cereus through 16S rRNA gene sequencing aligns with the method used by Jami et al.33 for taxonomic identification. However, previous study, focuses more specifically on Bacillus species, known for their antimicrobial properties and insecticidal activity. Whereas, Jami et al.32 explored the broader diversity of Actinobacteria. Both studies highlight the potential of freshwater fish gut microbiota as a source of bacteria with antimicrobial and bioactive compound-producing capabilities of these organisms. While Jami et al. 33examined a wide array of Actinobacteria with potential for secondary metabolite production, our study’s submission to the GenBank further supports the genetic characterization and global sharing of Bacillus isolates. In conclusion, while both studies focus on microbiota from freshwater fish, your research is more narrowly focused on Bacillus species, whereas Jami et al.33 emphasized a wider range of Actinobacteria and their secondary metabolite gene diversity.
Park et al.34 study reported that X. hominickii isolated from S. monticolum caused significant larval mortality when injected into different larvae of S. exigua, P. xylostella, H. assulta, P. brassicae, and D. melanogaster. Vicente-Diez et al. 35 found that the symbiotic bacterial supernatant from P. laumondii resulted in 64% mortality in P. spumarius nymphs. Ngugi et al.36 study observed that the bacterial supernatant of Bacillus subtilis at a concentration of 3.6×10-9 induced 68-88% larval mortality in Spodoptera litura within 24 and 48 hours. In our present study, the supernatant of isolated gut bacteria Bacillus subtilis and Bacillus cereus, injected at a concentration of 40 μl/ml, produced 76-80% mortality at 48 hours, confirming the pathogenicity of the gut bacteria secondary metabolites.
This present study reported that, the gut bacteria Bacillus subtilis ZBF-2 and Bacillus cereus., ZBF-4 showed optimal growth levels with the help of the added carbon (glucose) and nitrogen (peptone) sources to commercialized media. In addition, optimal temperature for Bacillus subtilis and Bacillus cereus., was found to be 30 and 35ºC. Among the tested pH at 7.0 showed the best bacterial growth rate for the bacterial species Bacillus subtilis and Bacillus cereus., through enhancing for the secondary metabolites production. Isolated gut bacterial cell free supernatant shows the higher concentration in Bacillus subtilis and Bacillus cereus., produced 76- 80% larval mortality at 48hrs. When compared to the control groups.
Conclusion
In conclusion, this study reported that, the zebrafish gut microbes ZBF-2 Bacillus subtilis, ZBF-4 Bacillus cereus showed optimal growth level supplemented with carbon (glucose) and nitrogen (peptone) source to commercialized media. In addition, optimal temperature for ZBF-2 Bacillus subtilis, ZBF-4 Bacillus cereus was found to be 30 and 35ºC. Among the tested pH at 7.0 showed the best bacterial growth for ZBF-2 Bacillus subtilis, ZBF-4 Bacillus cereus through enhancing the secondary metabolites production of the gut bacteria. Cell free supernatant of isolated gut bacteria ZBF-2 Bacillus subtilis, ZBF-4 Bacillus cereus with 10-4 µl/ml concentration produced 76- 80% larval mortality at 48hrs. The bioassays revealed a dose- and time-dependent effect, with higher concentrations causing larval mortality at 48 hours. This confirms the pathogenicity of ZBF-2 Bacillus subtilis, ZBF-4 Bacillus cereus strains. Our studies suggest that the larvicidal activity from the cell free supernatant extract could be used for production of novel class of insecticides properties in future.
Acknowledgement
We would like to thank the Department of Microbiology, AVS College of Arts and Science (Autonomous), Salem, Tamil Nadu, India for providing the infrastructural facility for carrying out this research work.
Funding Sources
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflict of Interest
The authors do not have any conflict of interest.
Data Availability Statement
This statement does not apply to this article.
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Clinical Trial Registration
This research does not involve any clinical trials.
Permission to reproduce material from other sources
Not Applicable
Author contributions
Salem Sundarrajan Maithili: Visualization, Supervision, Writing – Review and Editing.
Kandhasamy Lalitha: Conceptualization, Methodology, Writing – Original Draft.
Muthu Ganesh boopathi: Data Collection, Analysis, Writing – Review and Editing and authors gave approval for publication.
Reference
- Budd, K.Effects of diet, habitat, and phylogeny on the fecal microbiome of wild African savanna (Loxodonta africana) and forest elephants (L. cyclotis). Ecol. Evol. 2020;10(12); 5637-5650. doi: 10.1002/ece3.6305
CrossRef - Cui, X. Research progress of the gut microbiome in hybrid ” Microorganisms.2022:10(5);891.
https://doi.org/10.3390/microorganisms10050891
CrossRef - Stephens, W. Z et al. The composition of the zebrafish intestinal microbial community varies across development.” The ISME journal. 2016;10(3): 644-654. https://doi.org/10.1038/ismej.2015.140
CrossRef - Fortunato, Elvira, Pedro B, and Rodrigo M, et al. Oxide semiconductor thin‐film transistors: a review of recent advances. Mater. 2012: 24(22): 2945-2986.
CrossRef - Su, L. The different clinical characteristics of corona virus disease cases between children and their families in China–the character of children with COVID-19.Emerg.Infect.2020;9(1): 707-713. doi: 10.1080/22221751.2020.1744483
CrossRef - Ringø, E, Electron microscopy of the intestinal microflora of fish. Aquaculture2003;227(1-4):395-415. https://doi.org/10.1016/j.aquaculture.2003.05.001
CrossRef - Bates A,Two zebras and a cardiac arrest: a case report of concomitant Brugada syndrome and an anomalous coronary artery. Heart J. Case Rep.(2020); 4(6) :1-6. doi: 10.1093/ehjcr/ytaa425
CrossRef - Engeszer R,Patterson Andrew A. Rao and David M. Parichy et al. Zebrafish in the wild: a review of natural history and new notes from the field.” Zebrafish.2007;41: 21-40.
CrossRef - Kumar D, Vinod Kumar AK, Verma Prashant Singh, and Ashutosh Dubey,et al., Characterization and immobilization of partially purified alkaline protease extracted from rhizospheric soil bacterium, Bacillus megaterium strain EN-2 and Bacillus subtilis strain EN-3Research J. Microbiol. Res 2014;8(1): 33-39. https://doi.org/10.5897/AJMR2013.6167
CrossRef - Ehling-S, Monika, E F, and Michel G, et al. Food–bacteria interplay: pathometabolismofemetic Front. Microbiol. 2015;(6):704doi: 10.3389/fmicb.2015.00704
CrossRef - Abriouel, H, Charles MAP Franz, Nabil Ben O, and Antonio G. “Diversity and applications of Bacillus bacteriocins. FEMS Microbiol. Rev. 2011: 35(1);201-232. doi: 10.1111/j.1574-6976.2010.00244.
CrossRef - Luise D, Micol B, Vincenzo M, Chiara S, Paolo Bosi, Andrea L, Flaminia F et al. “Bacillus probiotic supplementation diminish the Escherichia coli F4ac infection in susceptible weaned pigs by influencing the intestinal immune response, intestinal microbiota and blood metabolomics. J. Anim. Sci. Biotechnol.2019: 10(1): 74. doi: 10.1186/s40104-019-0380-3
CrossRef - Yakimov AP. Seregina TA. Kholodnyak AA. Kreneva RA. Mironov AS. PerumovDAand Timkovskii, AL, et al. Possible function of the ribT gene of Bacillus subtilis: theoretical prediction, cloning, and expression. Acta Naturae (англоязычнаяверсия), 2014:6(3):106-109: https://doi.org/10.32607/20758251-2014-6-3-106-109.
CrossRef - Kim -J, Jae-KH Jong-SP, Jin-S L, Soon-H L, Joon-Il C, and Keun-S K. et al. Various enterotoxin and other virulence factor genes widespread among Bacillus cereus and Bacillus thuringiensisJMicrobiol Biotechnol.2015:25(6):872-879. doi: 10.4014/jmb.1502.02003.
CrossRef - Wang XQDL, Zhao LL, ShenCL Jing and CS, Zhang et al.Application and mechanisms of Bacillus subtilis in biological control of plant disease. In Role of rhizospheric microbes in soil: Volume 1: Stress management and agricultural sustainability, Singapore: Springer Singapore, 2018:225-250. DOI:10.1007/978-981-10-8402-7_9
CrossRef - Bernadsky, G, and RosenbergE et al. Drag-reducing properties of bacteria from the skin mucus of the cornetfish (Fistulariacommersonii).Microbial Ecol.1992: 24(1): 63-76. https://doi.org/10.1007/BF00171971.
CrossRef - Langendijk, Petra S, Frits Schut Gijsbert J, Jansen, Gerwin C,Raangs, Ger R., Kamphuis, MH, Wilkinson and Gjalt W. Welling.et al. Quantitative fluorescence in situ hybridization of Bifidobacterium spp. with genus-specific 16S rRNA-targeted probes and its application in fecal samples. Environ. Microbiol.1995; 61(8);3069-3075.
CrossRef - MunroPOA, Barbour and TH Blrkbeck,Comparison of the gut bacterial flora of start‐feeding larval turbot reared under different conditions. Appl. Microbiol. 1994: 77, no. 5 560-566.
CrossRef - Cahill Marian M. Bacterial flora of fishes: a review. Microbiol ecol. 1990: 19(1) 21-41. doi: 10.1007/BF02015051.
CrossRef - Gandhi K, and RH. Patil. Field resistance of Spodoptera litura (Fab.) to conventional insecticides in India. Crop Prot. 2016: 88;103-108. https://doi.org/10.1016/j.cropro.2016.06.009
CrossRef - Armes Nigel J, John A. Wightman Deepak R. Jadhav and Gangavalli V. Ranga R et al. Status of insecticide resistance in Spodoptera litura in Andhra Pradesh, India. Pestic. Sci.1997: 50,(3) 240-248.
CrossRef - Sharma, S and Tejinder S B, et al. Effects of elevated temperature and carbon dioxide on food consumption and growh of Spodoptera litura Fabricius on cauliflower.Agrometeorol..2018:20(4)305-310. DOI: https://doi.org/10.54386/jam.v20i4.571
CrossRef - Singh A, Singh S, Singh, R, Kumar S, Singh, SK. and Singh I.K, et al. Dynamics of Zea mays transcriptome in response to a polyphagous herbivore, Spodoptera litura.FunctIntegr Genomics2021: 21(5): 571-592. doi: 10.1007/s10142-021-00796-7
CrossRef - Yakimov, APA. S. Afanasev M. A. Khodorkovskiy, and M.G Petukhov et al. Design of stable α-helical peptides and thermostable proteins in biotechnology and biomedicine. Acta Naturae. 2016: 8,4 (31):70-81.
CrossRef - Wang, Yutang, Lixia Li, Hua Liu, Tong Zhao, Chenshuo Meng, Zhande Liu, and Xuebo Liu et al. Bioactive compounds and in vitro antioxidant activities of peel, flesh and seed powder of kiwi fruit.” J. Food Sci. Technol.2018;53(9): 2239-2245. https://doi.org/10.1111/ijfs.13812
CrossRef - Kaur, M, Chadha P, KaurS, Kaur A, Kaur R, YadavAK, Kaur R et al.Schizophyllumcommune induced genotoxic and cytotoxic effects in Spodoptera litura.” Sci Rep.2018: 8(1): 4693. https://doi.org/10.1038/s41598-018-22919-0
CrossRef - Vijayakumari Nadaraja, A, Jayakumaran Nair, A, Prameela M, Hari N, BalakrishnanN et al. Evaluation of currently employed food preservation conditions to tackle biofilm forming food pathogens. Food Saf. 2018: 38(1);12407. https://doi.org/10.1111/jfs.12407
CrossRef - Park J, Park BS, Wang P, Patidar SK, Kim JH, Kim SH, Han MS, et al. Phycospheric native bacteria Pelagibacabermudensis and Stappia sp. ameliorate biomass productivity of Tetraselmis striata (KCTC1432BP) in co-cultivation system through mutualistic interaction.” Plant Sci. 2017: 8; 289. https://doi.org/10.3389/fpls.2017.00289
CrossRef - Lalitha, K., Kalaimurgan, D, Nithya K, Venkatesan S. and Shivakumar MS. Et al. Antibacterial, antifungal and mosquitocidal efficacy of copper nanoparticles synthesized from entomopathogenic nematode: Insect–host relationship of bacteria in secondary metabolites of Morganella morganiisp.PMA1Arab. J. Sci. Eng..2020: 45(6); 4489-4501. DOI:10.1007/s13369-020-04487-6
CrossRef - Ray AK, K. Ghosh and EJA N, Ringø et al. Enzyme‐producing bacteria isolated from fish gut: a review. Nutr. 2012: 18(5); 465-492. DOI:10.1111/j.1365-2095.2012.00943.x
CrossRef - Dhayalan A, Velramar B, Govindasamy B, Ramalingam KR, Dilipkumar A, and PachiappanP et al.Isolation of a bacterial strain from the gut of the fish, Systomussarana, identification of the isolated strain, optimized production of its protease, the enzyme purification, and partial structural characterization Genet. Eng. &Biotechnol.(2022): 20(1);24. https://doi.org/10.1186/s43141-022-00299-3
CrossRef - Jami SA, Ajamhassani M, and Naveh VH,et al. Biochemical characteristics of digestive enzymes of codling moth, Cydia pomonella (Lepidoptera: Tortricidae).Phytoparasitica. 2025: 53(2); 30.
CrossRef - Jami M, Lai Q, Ghanbari M., Moghadam MS, Kneifel W, and Domig, KJ, et al. Phylogenetic diversity and biological activity of culturable Actinobacteria isolated from freshwater fish gut microbiota. Res .2015: 175;6-15. https://doi.org/10.3389/fmicb.2015.00737
CrossRef - Park Y, Kang S, Sadekuzzaman M, Kim H, Jung J.K and Kim Y, et al. Identification and bacterial characteristics of Xenorhabdushominickii ANU101 from an entomopathogenic nematode, Steinernema monticolum. Invertebr. Pathol 2017;144: 74-87. doi: 10.1016/j.jip.2017.02.002.
CrossRef - Vicente-Díez I, Blanco-PérezR,ChelkhaM, PuellesM, Pou A, and Campos-Herrera R et al. Exploring the use of entomopathogenic nematodes and the natural products derived from their symbiotic bacteria to control the grapevine moth, Lobesiabotrana(Lepidoptera:Tortricidae). Insects.(2021):12,(11)1033. https://doi.org/10.3390/insects12111033
CrossRef - Ngugi Ceciliah N,etal. Characterization and evaluation of entomopathogenic nematodes for the management of tomato leafminer (Tuta Absoluta Meyrick).” PhD diss., University of Nairobi, 2021. http://erepository.uonbi.ac.ke/handle/11295/157047
Accepted on: 18-Dec-2025
Second Review by: Dr. Vivek Anand
Final Approval by: Dr. Eugene A. Silow