COI-Based DNA Barcoding and Phylogenetic Analysis of Catharsius molossus (Coleoptera: Scarabaeidae) from Tiruchirappalli, Southern India.
1Department of Zoology, Holy Cross College (Autonomous), Tiruchirappalli, Tamil Nadu. India.
2Department of Zoology, Nehru Memorial College, Affiliated to Bharathidasan University, Tiruchirappalli District, Tamil Nadu, India.
Corresponding Author Email: kaliyaperumalsaravanan72@gmail.com
DOI : http://dx.doi.org/10.13005/bbra/3465
ABSTRACT:Dung beetles (Coleoptera: Scarabaeidae) play a crucial role in ecosystem functioning, yet their molecular taxonomy and phylogenetic relationships remain underexplored in tropical regions. India has almost 400 species of dung beetle representing 30 genera, even though genetic data on much of the taxa are still scarce in the global repositories. This paper used the methodology of mitochondrial cytochrome c oxidase subunit I (COI) DNA barcoding to determine the identification of Catharsius molossus specimens that were taken in Tiruchirappalli in Tamil Nadu. Amplification of genomic DNA was done using universal COI primers and sequencing produced 19 good quality barcode sequences. The sequences were uploaded to the BOLD Systems v3 and NCBI GenBank (Accession No. PP396739). Phylogenetic analysis using the Neighbor-Joining (NJ) method in MEGA11 was done to verify morphological identifications, which supported clustering at a species level. These findings validate that COI is a good molecular identification marker of dung beetles and they can be useful in future biodiversity survey and phylogenetic studies in India.
KEYWORDS:Biodiversity; Dung beetles; Catharsius molossus; DNA barcoding; Phylogenetic analysis.
Introduction
Dung beetles are a group of ecologically important insects, which plays an important role in the provision of key ecosystem services including nutrient recycling, soil aeration, and secondary seed dispersal.1,2,3. It is well known that dung beetles are great bioindicators of biodiversity and ecological impact assessment because they are sensitive to changes in the environment in terms of the community structure.4-6 Among them, the genus Onthophagus is one of the most divergent lineages, with over 2100 described species belonging to the subfamily Scarabaeinae, including 16 known tribes at the moment.7,8
There exists a rich dung beetle fauna in India, containing almost 400 species spread in 30 genera and 9 tribes. The Western Ghats are biodiversity hot spots of the country and particularly known to hold the richness and endemism of the species.9 Taxonomic resolution of dung beetles is difficult because of phenotypic plasticity, convergent morphology and the existence of cryptic species complexes even though they have environmental significance.6,8 Thus, the interdisciplinary methods that include the application of molecular tools have become critical in solving the boundaries of species and enhancing phylogenetic hypotheses.
DNA barcoding, specifically the mitochondrial cytochrome c oxidase subunit I (COI) gene, has been found to be very useful in enhancing species delimitation, uncovering latent diversity, and expediting the process of identifying Scarabaeinae globally.10,11,12 However, Indian dung beetle, Catharsius molossus from the Tiruchirappalli region remain severely underrepresented in global reference databases such as BOLD and GenBank, limiting their use in ecological monitoring, evolutionary studies, and conservation planning.
Materials and Methods
Materials
Specimens: Adult Catharsius molossus collected from agricultural fields in Woraiyur, Tiruchirappalli, India.
Preservation: Specimens stored in sterile jars containing 100% molecular-grade ethanol and maintained at −20 °C.
Tissue: Hind leg muscle tissues (~20–25 mg) dissected under sterile conditions using autoclaved tools.
DNA Extraction Kit: HiPure Multi-Sample DNA Purification Kit (HiMedia, India).
DNA Quality Assessment: 0.8% agarose gel electrophoresis and NanoDrop 2000 spectrophotometer (Thermo Scientific).
PCR Primers: LCO1490: 5′-GGTCAACAAATCATAAAGATATTGG-3′
HCO2192: 5′-TAAACTTCAGGGTGACCAAAAAATCA-3′
PCR Product Purification: GeneJET PCR Purification Kit (Thermo Scientific).
Sequencing Platform: ABI 3500 Genetic Analyzer using BigDye Terminator v3.1 Kit (Applied Biosystems).
Sequence Editing Software: CodonCode Aligner v9.0.
Databases for Validation: NCBI GenBank (BLASTn) and BOLD v3.
Phylogenetic Tools: MEGA v11 with MUSCLE alignment; NJ method with 500 bootstrap replicates.
Methods
Study area and Specimen Collection
Adult dung beetles (C. molossus) were captured in agricultural fields at Woraiyur, Tiruchirappalli, southern India (10.813° N, 78.685° E). The sampling took place through the use of standard entomological sweep nets as well as hand collection. The specimens were shifted into the sterile insect jar and stored instantly in 100 percent molecular-grade ethanol to avoid degradation of DNA. Samples were kept at -20° C until further treatment.
Tissue preparation and DNA isolation
Hind leg muscle tissues were carefully excised from each specimen under sterile conditions. Approximately 20–25 mg of muscle tissue was placed in sterile 1.5 mL microcentrifuge tubes for DNA extraction. Genomic DNA was isolated using the HiPure Multi-Sample DNA Purification Kit with slight adjustments to the manufacturer’s instructions. Initially, the tissues were rinsed twice with wash buffer to remove residual ethanol. Samples were then digested in a mixture of 20 μL Proteinase K and 200 μL lysis buffer at 55°C for one hour. Following the addition of 200 μL ethanol, the lysates were transferred onto silica spin columns. Bound DNA was washed sequentially with pre-wash and wash buffers before being eluted in 100 μL of elution buffer. DNA integrity was examined on a 0.8% agarose gel, while yield and purity were quantified using a NanoDrop 2000 spectrophotometer.
PCR Amplification of Mitochondrial COI Gene
The 658 bp barcode fragment of the mitochondrial cytochrome oxidase I (COI) gene was amplified using the universal primers viz., LCO1490 and HCO2192
Isolated DNA samples were arranged in 96-well plates, with corresponding sample information recorded for reference. Polymerase chain reaction (PCR) was carried out for all specimens using the above primers, and the target fragment was successfully amplified in each case. The PCR products were purified with the GeneJET PCR Purification Kit (Thermo Scientific, EU-Lithuania) to eliminate primer dimers and other residual contaminants. Product quality was evaluated on a 1.8% agarose gel with a 100 bp DNA ladder as the size marker, confirming that the amplified fragments were suitable for sequencing.
Purification and sequencing
The GeneJET PCR Purification Kit was used to purify the PCR products to eliminate any remaining primers and other reagents. The purified amplicons were sequenced in both directions in an ABI 3500 Genetic Analyzer using ABI Prism BigDye, Terminator v3.1 Cycle Sequencing Kit. After sequencing, the products were purified by ethanol/EDTA precipitation method and resuspended in 12 mL of Hi-Di formamide. Two ml of the samples were then incubated at 95 °C in 5 minutes before being separated using capillary electrophoresis.
Sequence edit and alignment
Obtained sequences were then checked in codon code aligner v9.0 to check the ambiguities within the sequence. Ambiguous bases were trimmed, and consensus sequences were generated from forward and reverse reads. Final sequences were exported in FASTA format and validated by BLASTn searches against the NCBI GenBank database.
DNA Barcode generation and Data submission
The Confirmed COI sequences were uploaded to the NCBI (National Centre of Biotechnology Information with the accession number of PP396739 and also to the BOLD v3 (Barcode of Life Data Systems). A DNA barcode record was developed as per BOLD conventions that included the information about the voucher specimens, collection data and trace files.
Phylogenetic analysis
Multiple sequence alignments were performed with MUSCLE implemented in MEGA v11.13 Evolutionary relationships were inferred using the Neighbor-Joining (NJ) method14 with 500 bootstrap replications.15 Evolutionary distances were calculated using the Maximum Composite Likelihood (MCL) model.16 The analysis included 13 representative sequences from GenBank for related Scarabaeinae taxa (e.g., Onthophagus, Dichotomius spp.). Ambiguous positions were removed by pairwise deletion, resulting in a final alignment of 573 positions.
Observation and Results
Isolation of genomic DNA of C. molossus
Genomic DNA of C molossus was successfully isolated from leg tissue using standardized method. Agarose gel electrophoresis showed intact high-molecular-weight DNA with no evidence of degradation, indicating good-quality DNA to perform PCR amplification (Figure 1).
PCR amplification and purification of COI gene
The mitochondrial COI bar code region (~658 bp) was amplified successfully using the universal primers LCO1490 and HCO2198. The presence of a distinct band with an appropriate size was detected by agarose gel electrophoresis and no observable nonspecific products were detected. There was no amplification with negative controls, and this confirms that there was no contamination (Figure 2). PCR products were then purified with GeneJET PCR purification kit. Electrophoresis showed a sharp and strong band of expected size, after purification indicating the elimination of primer dimers and residual reagents.
 |
Figure 1: Agarose gel electrophoresis of genomic DNA isolated from C. molossus (DNA quality checking). |
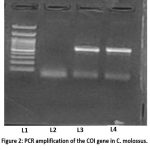 |
Figure 2: PCR amplification of the COI gene in C. molossus. Lane 1: 100 bp DNA ladder; Lane 2: negative control; Lane 3: genomic DNA sample; Lane 4: positive control. |
Sequence generation and submission
Purified PCR products of C. molossus were sequenced bidirectionally. Consensus sequences were assembled and manually curated in CodonCode Aligner. The presence of species-level identification with C. molossus was confirmed by BLASTn searches into the NCBI database indicating more than 99 percent identity with consensus sequence. This curated COI sequence was deposited in NCBI (Genbank) with the accession number of PP396739. Metadata voucher ID, collecting locality and the details of the specimen were also documented through Barcode of Life Data Systems (BOLD v3).
Following FASTA file of published COI gene nucleotide sequences of the C. molossus submitted to the NCBI
ORIGIN
1 acatctttaa gaattttaat tcgaatagaa ttagggtctc ctggaacctt aattggtgat
61 gatcaaattt ataatgtaat tgtaactgca catgcattta ttataatttt ttttatagtt
121 atacctattt taattggtgg atttggaaat tgacttgttc cattaatatt aggggcccca
181 gatatagcat ttccacgaat aaataatata agattttgac ttttacctcc atctttaact
241 cttttattaa taagaagaat agtaraaaag ggggcaggta caggatgaac agtatacccc
301 cctctttctt caaatattgc tcatggaggg gcatctgtag atttagctat ttttaractt
361 catcttgcag gaatttcttc aattttaggt gctattaatt ttattactac agttattaat
421 atacgatcaa caggaataac atttgatcga ataccattat ttgcttgagc tgttgctatt
481 actgcattac tacttttact ttctttacct gtattagcag gcgctattac tatacttcta
541 acagatcgaa atttaaatac ttcctttttt gatcctatag gaggtggaga tccaatttta
601 taccaacatt tattttgatt ttttgg
//
DNA barcode generation
A DNA barcode was generated for C. molossus using the BOLD Systems v3 platform (Figure 3). Barcode Index Number (BIN) was assigned for the sequence and digitally archived, offering permanent molecular identifier for the species. The barcode record contained sequence data, voucher information, raw trace files and metadata of collection.
 |
Figure 3: DNA barcode generated for C. molossus in the BOLD Systems v3 database. |
Phylogenetic analysis
The resulting Neighbor-Joining (NJ) tree of MEGA11 grouped C. molossus (PP396739) together with conspecific sequences retrieved via the GenBank (e.g., JQ855867, JQ855861). The bootstrap values were very high, C. molossus clade was strongly supported (>90). At elevated taxonomic ranks, C. molossus was found to be clustered with other species of Scarabaeinae such as Dichotomius mamillatus, Onthophagus ovatus and Onthophagus turbatus. The topology was compatible with classical morphological segregations of Scarabaeinae. Divergence of sequences between C. molossus and its nearest relatives was 0.811.4 percent, and species divergence over genera was more than 8 percent, which reinforces the power of COI as a discriminative marker. The results of the phylogenetic analysis of the dung beetle are shown in figure 4.
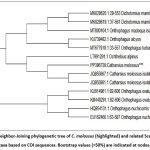 |
Figure 4: Neighbor-Joining phylogenetic tree of C. molossus (highlighted) and related Scarabaeinae taxa based on COI sequences. Bootstrap values (>50%) are indicated at nodes. |
Discussion
DNA barcoding has emerged as an important approach for the bio-surveillance of insect pests, enabling the quick identification of specimens irrespective of their developmental stage or preservation state. One of its most promising applications, especially when combined with sequence databases and cost-effective high-throughput sequencing, lies in biodiversity assessment. This approach can enhance the efficiency and standardization of ecological surveys, particularly for taxa that are diverse and challenging to identify using traditional morphological methods.
DNA quality and amplification success
Isolation of pure genomic DNA of C. molossus and reproducible amplification of COI barcode region show the strength of molecular methods in identifying species in Scarabaeinae beetles. The amplification of the isolated DNA using the universal primers LCO1490 and HCO219817 was clear and consistent with the primers used in other insect taxa.10,12 The same success rate has been observed with scarabaeids in the Western Ghats, where the success rate also corroborates the consistency of COI primers in different genera of dung beetles.6
A lack of nonspecific products or contamination of PCR reactions also highlights the reliability of the method and is essential in areas such as India where dung beetle molecular data are sparse. Sulistyani et al.18 also introduced recent research on Helopeltis spp., which identified the efficiency of COI in sorting closely related species, and indicated that the marker can be used across taxonomically diverse insect groups.
Sequence identity and species confirmation
The generated sequence of the COI (626 bp) was found to be more than 99% similar to C. molossus sequences found in Genbank, which validated correct species identification. This sequence identity is comparable to the thresholds typically employed in insect DNA barcoding, at which >98% similarity usually indicates conspecificity.10,19
Similar results were also observed in Bangladesh by Das et al,20 who were able to prove that COI was a reliable discriminative factor among cryptic Aulacophora species. Similarly, Tahir et al21 confirmed COI as an accurate tool in identifying pests in rice eco systems. This combination of findings confirms that COI could be effectively used in Indian dung beetle fauna, where morphological traditional taxonomy frequently fails to sort out species delimits.
DNA barcode generation and submission to global repositories
The submission of DNA sequences of C. molossus to both NCBI (Accession PP396739) and BOLD Systems strengthen the molecular database of Indian Scarabaeinae. Existing coverage of Indian dung beetles in world repositories is poor, and less than 10 percent of described species are publicly available with barcodes.5,6 Having a barcode record, which is standardized, this study will be used to make a reference point in future ecological observation and species discovery.
Insect monitoring worldwide has already been sped up through BOLD-based barcoding.18,22 As an example, metabarcoding work has demonstrated that dung beetles can serve as an iDNA sampler of mammalian biodiversity.21 Inclusion of C. molossus sequences, therefore, does not only confirm the taxonomy of insects, but can also indirectly reinforce biodiversity measuring tools in tropical ecosystems.
Phylogenetic analysis
The phylogenetic tree positioned C. molossus in a highly supported clade with conspecific sequences (JQ855867, JQ855861) and a bootstrap of more than 90. The observation indicates a high rate of genetic cohesion in the species, which is similar to previous morphological classification.8
In addition, Onthophagus and Dichotomius spp. are clustered and there is a tribal relationship established within Scarabaeinae.8 The intergeneric divergence was observed to be greater than 8% which is consistent with the values in other scarabaeids6,23 and shows that using the marker to determine genus-level taxonomy is informative. More recent phylogenomic studies have established that COI topologies tend to show consistent results with multi-gene datasets,24 which point to its continued use in single-marker barcoding studies.
Ecological and taxonomical implications
Dung beetles contribute to the important functions of nutrient cycling, secondary seed dispersal, and soil aeration.24 This is more important to evaluate their ecological functions and conservation status, as well as to infer their ecological roles. Nevertheless, the morphological diagnosis is often obstructed due to phenotypic plasticity and cryptic diversity.6,25 DNA barcoding overcomes these difficulties and offers a unifying structure to the species delimitation, ecological scanning and conservation planning.
Having C. molossus in the global barcode databases guarantees the ability of future biodiversity studies in India to depend on molecular reference data. This is especially applicable in the face of the increasing focus on metabarcoding methods of community-level monitoring.21,26
Limitations and future directions
Although the present study provides a first DNA-barcode reference for C. molossus from Southern India, some limitations have to be acknowledged. First, only the COI gene was analyzed even if it is effective, that may not be enough to resolve deeper phylogenetic relationships because of incomplete lineage sorting or mitochondrial introgression.27 More resolution may be achieved via multi-gene approaches that use nuclear loci (e.g., ITS, 28S rDNA) or mitogenomic data.27
Second, the study only used one geographic population. A more extensive sampling program over the range of the species is required to evaluate intraspecific variation and potential cryptic diversity. In order to obtain a more comprehensive picture of the diversity of C. molossus, DNA barcoding should in future be combined with morphometric analysis and ecological niche modelling.
Conclusion
In conclusion, this study demonstrates that COI-based DNA barcoding provides a reliable, efficient, and scalable tool for the molecular identification of C. molossus. The findings contribute valuable genetic data to the existing reference library for Indian dung beetles. The inferred phylogenetic placement is consistent with established taxonomic classifications, further validating the utility of the COI marker. Overall, the expanded barcode dataset strengthens ongoing efforts in biodiversity assessment, monitoring, and conservation.
Acknowledgement
The authors extend their heartfelt thanks to the Central Instrumentation Centre, Holy Cross College, Tiruchirappalli, for their helps in conducting our experiments.
Funding Sources
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflict of Interest
The authors do not have any conflict of interest.
Data Availability Statement
This statement does not apply to this article.
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Clinical Trial Registration
This research does not involve any clinical trials.
Permission to reproduce material from other sources
Not Applicable
Author Contributions Statement
Elavarasi Siddhaiyan: Supervision, Conceptualization, Methodology review and editing.
Priyadharshni Saravanan: Investigation and formal analysis.
Horne Iona Averal: Formal analysis, review and editing.
Revathi Ganesan: Validation, Formal analysis, and Investigation.
Saravanan Kaliyaperumal: Conceptualization, Methodology, Writing – original draft.
References
- Davis ALV, Scholtz CH. Historical vs. ecological factors influencing global patterns of scarabaeine dung beetle diversity. Diversity and Distributions. 2001; 7: 161–174. https://doi.org/10.1111/j.1472-4642.2001.00102.x
CrossRef - Simmons, L. W., & Ridsdill-Smith, T.J. (2011). Ecology and Evolution of Dung Beetles. Wiley-Blackwell, 368p.
CrossRef - Eleanor M. Slade, Darren J. Mann, Owen T. Lewis, Biodiversity and ecosystem function of tropical forest dung beetles under contrasting logging regimes, Biological Conservation. 2011; 144 (1): 166-174. https://doi.org/10.1016/j.biocon.2010.08.011.
CrossRef - McGeoch MA, Van Rensburg BJ, Botes A. The verification and application of bioindicators: a case study of dung beetles in a savanna ecosystem. Journal of Applied Ecology. 2002; 39(4): 661–672. doi: https://doi.org/10.1046/j.1365-2664.2002.00743.x
CrossRef - Nichols, E. Spector, S. Louzada, J. Larsen, T. Amezquita, S. Favila, M.E. Ecological functions and ecosystem services provided by Scarabaeinae dung beetles. Biological Conservation. 2008; 141(6): 1461-1474. https://doi.org/10.1016/j.biocon.2008.04.011.
CrossRef - Asha G, Palatty S. DNA barcode and phylogenetic analysis of dung beetles (Coleoptera: Scarabaeidae) from the Western Ghats biodiversity hotspot, India. Indian Journal of Tropical Insect Science. 2020; 41(2): 1419-1425. https://doi.org/10.1007/s42690-020-00336-z
CrossRef - Philips TK. Phylogeny of the Oniticellini and Onthophagini dung beetles (Scarabaeidae, Scarabaeinae) from morphological evidence. Zookeys. 2016; 579: 9-57. doi: https://doi.org/10.3897/zookeys.579.6183.
CrossRef - Tarasov S, Génier F. Innovative Bayesian and parsimony phylogeny of dung beetles (Coleoptera: Scarabaeinae) enhanced by ontology-based partitioning of morphological characters. PLoS One. 2015; 10(3): e0116671. doi: https://doi.org/10.1371/journal.pone.0116671
CrossRef - Sabu TK, Nithya S. Vinod KV. Faunal survey, endemism and possible species loss of Scarabaeinae (Coleoptera: Scarabaeidae) in the western slopes of the moist South Western Ghats, South India. Zootaxa. 2011; 2830(1):29–38. https://doi.org/10.11646/zootaxa.2830.1.3
CrossRef - Hebert PDN, Cywinska A, Ball SL, Jeremy R deWaard. Biological identification through DNA barcodes. Proceedings of the Royal Society B: Biological Sciences. 2003; 270(1512):313–21. https://doi.org/10.1098/rspb.2002.2218
CrossRef - Hebert PD, Penton EH, Burns JM, Janzen DH, Hallwachs W. Ten species in one: DNA barcoding reveals cryptic species in the neotropical skipper butterfly Astraptes fulgerator. Proc Natl Acad Sci. 2004; 101(41):14812-7. doi: 10.1073/pnas.0406166101.
CrossRef - Virgilio M, Backeljau T, Nevado B. et al.Comparative performances of DNA barcoding across insect orders. BMC Bioinformatics. 2010; 11: 206. https://doi.org/10.1186/1471-2105-11-206
CrossRef - Tamura K, Stecher G, Kumar S. MEGA11: Molecular Evolutionary Genetics Analysis Version 11. Mol Biol Evol. 2021; 25;38(7): 3022-3027. doi: 10.1093/molbev/msab120.
CrossRef - Saitou N, Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Molecular Biology and Evolution. 1987; 4(4): 406– 425, https://doi.org/10.1093/oxfordjournals.molbev.a040454
CrossRef - Felsenstein J. Confidence limits on phylogenies: an approach using the bootstrap. Evolution. 1985; 39(4): 783–791. https://doi.org/10.1111/j.1558-5646.1985.tb00420.x
CrossRef - Tamura K, Nei M, Kumar S. Prospects for inferring very large phylogenies using the neighbor-joining method. 2004; 101(30): 11030–11035. https://doi.org/10.1073/pnas.0404206101
CrossRef - Folmer O, Black M, Hoeh W, Lutz R, Vrijenhoek R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Molecular Marine Biology and Biotechnology. 1994; 3 (5): 294‑299
- Sulistyani P, Sari ABT, Rahayu DS. DNA barcoding for identification of Helopeltis from Kaliwining experimental station using mitochondrial COI gene. Coffee and Cocoa Research Journal. 2022; 38(3): 179–189. https://doi.org/10.22302/iccri.jur.pelitaperkebunan.v38i3.521
CrossRef - Ratnasingham S, Hebert PDN. A DNA-based registry for all animal species: The Barcode Index Number (BIN) system. PLoS ONE. 2013; 8(7): e66213. https://doi.org/10.1371/journal.pone.0066213.
CrossRef - Das SR, Tithi JF, Sarker S, Rain FF, Md Aslam AF. Diagnosis of three Pumkin Leaf Beetle (Aulacophora: Chrysomelidae; Coleoptera) pests through mtDNA-COI barcode molecular approach. Bangladesh Journal of Zoology, 2021. 48(2); 399–411. doi: https://doi.org/10.3329/bjz.v48i2.52379
CrossRef - Tahir HM, Noor A, Mehmood S, Sherawat SM, Qazi MA. Evaluating the accuracy of morphological identification of insect pests of rice crops using DNA barcoding. Mitochondrial DNA Part B Resour. 2018; 3(2): 1220-1224. doi: https://doi.org/10.1080/23802359.2018.1532334
CrossRef - Drinkwater R, Williamson J, Clare EL, Chung AYC, Rossiter SJ, Slade E. Dung beetles as samplers of mammals in Malaysian Borneo: A test of high‐throughput meta barcoding of iDNA. Peer J. 2021; 9: e11897. https://doi.org/10.7717/peerj.11897
CrossRef - Zothansanga C. A review of scarab beetles (Coleoptera: Scarabaeidae) diversity in India. Science vision, 2021; (2): 43-49. https://doi.org/10.33493/scivis.21.02.03
CrossRef - Filipović I, Hereward JP, Rašić G, Devine GJ, Furlong MJ, Etebari K. The complete mitochondrial genome sequence of Oryctes rhinoceros(Coleoptera: Scarabaeidae) based on long-read nanopore sequencing. Peer J. 2021; 13;9: e10552. doi: 10.7717/peerj.10552.
CrossRef - Jake E Bicknell, Simon P Phelps, Richard G Davies, Darren J Mann, Matthew J Struebig, Zoe G Davies. Dung beetles as indicators for rapid impact assessments: Evaluating best practice forestry in the neotropics, Ecological Indicators. 2014; 43: 154-161. https://doi.org/10.1016/j.ecolind.2014.02.030.
CrossRef - Moritz C, Cicero C. DNA barcoding: promise and pitfalls. PLoS Biol. 2004; 2(10):e354. doi: 10.1371/journal.pbio.0020354.
CrossRef - Boykin LM, Armstrong K, Kubatko L, De Barro P. DNA barcoding invasive insects: Database roadblocks. Invertebrate Systematics. 2012; 26(6): 506–514. https://doi.org/10.1071/IS12025.
CrossRef
Accepted on: 21-11-2025
Second Review by: Dr. Vivek Anand
Final Approval by: Dr. Wagih Ghannam









