An Efficient AI Model for the Classification of Skin Lesions in Emerging Infectious Diseases
Department: MBA – Finance, Osmania University, India
Corresponding Author E-mail: venkat9876098@gmail.com
DOI : http://dx.doi.org/10.13005/bbra/3471
ABSTRACT:We propose MpoxNet, a lightweight convolutional neural network for the multi- class classification of skin lesions associated with emerging infectious diseases. Built on six ConvNeXt blocks and a Dual Attention Block that jointly models spatial saliency and channel dependencies, MpoxNet enhances discriminative fea- ture learning while maintaining computational efficiency. The model is evaluated on the Mpox Skin Lesion Dataset Version 2.0 (755 images across six clinically annotated classes), achieving a test accuracy of 86.49%, a macro F1-score of 86.79%, and a Matthews Correlation Coefficient of 82.76%. Grad-CAM visual- izations confirm that MpoxNet focuses on pathologically meaningful regions such as umbilicated lesion centers (Monkeypox) or maculopapular spread (Measles), supporting clinical interpretability. In addition, cross-dataset experiments on BreastMNIST, OCTMNIST, and OrganAMNIST demonstrate strong generaliza- tion capability. These results indicate that MpoxNet provides a computationally efficient, interpretable, and robust diagnostic tool suitable for deployment in resource-constrained clinical and public health settings.
KEYWORDS:Deep Learning; Dual Attention; MpoxNet; Skin Lesion; ConvNext
Introduction
Mpox (formerly known as Monkeypox), a re-emerging zoonotic orthopoxvirus, has recently gained global attention as a significant public health threat. Characterized by vesiculopustular eruptions, fever, and lymphadenopathy, Mpox can clinically mimic other viral exanthems such as measles, varicella (chickenpox), and even smallpox. This visual similarity poses substantial challenges for early-stage diagnosis, particularly in remote or resource-limited settings where advanced molecular diagnostics, such as polymerase chain reaction (PCR) testing, are not readily available. The lack of accurate and scalable diagnostic tools during outbreaks hinders containment efforts and delays timely treatment interventions.
Recent advances in artificial intelligence (AI), particularly deep learning (DL), have accelerated the development of automated diagnostic frameworks capable of interpreting complex visual cues in medical imagery. These models learn abstract and hierarchical features directly from lesion images, enabling them to distinguish between visually overlapping skin conditions. This capability is especially valuable for infectious diseases with limited histopathological or morphological differentiators. Consequently, DL-based image classification has emerged as a viable tool to augment clinical workflows and support rapid, non-invasive screening during outbreaks.
Despite the promise of DL in dermatological image analysis, several critical chal- lenges persist. Manual interpretation of skin lesions remains subjective and is often influenced by the clinician’s expertise, leading to diagnostic variability. Automated systems, while more consistent, are heavily dependent on the availability, quality, and diversity of training data. Many dermatological datasets are limited in size or affected by severe class imbalance, both of which compromise model generalizability. Furthermore, the inherent visual overlap between Mpox and other viral dermatoses frequently leads to misclassification, underscoring the need for attention mechanisms or fusion-based strategies to improve feature discrimination.
Several state-of-the-art models have been proposed to address these challenges. For example, MSMP-Net leveraged a ConvNeXt backbone augmented with multi-scale patch-level fusion to capture both coarse and fine-grained lesion features from the Mpox Skin Lesion Dataset (MSLD).1 Other works introduced generative approaches such as the “Mask, Inpaint, and Measure” (MIM) pipeline, where the quality of GAN-based image restoration is used as a proxy for classification.2 Although these methods demonstrate strong accuracy and robustness on benchmark datasets, they often require substantial computational resources and remain susceptible to adversar- ial noise and real-world imaging variability. Moreover, their interpretability is limited, making clinical validation and deployment challenging.
To address these gaps, we introduce MpoxNet, a lightweight, efficient, and interpretable deep learning framework specifically designed for Mpox skin lesion clas- sification. Built on a ConvNeXt backbone, our architecture incorporates a Dual Attention Block (DAB) that jointly models channel-wise and spatial feature depen- dencies. This dual-attention strategy enhances contextual awareness and strengthens the extraction of discriminative lesion features, thereby improving the model’s ability to differentiate visually similar conditions. Unlike conventional attention mechanisms that introduce significant parameter overhead, the proposed DAB is optimized for min- imal computational cost, making MpoxNet suitable for deployment on edge devices or in resource-constrained clinical environments.
MpoxNet is also designed with practical deployment considerations in mind. Its compact parameter footprint supports rapid inference, while built-in attention visual- ization enables clinical interpretability by highlighting the regions most influential in the model’s decisions. To evaluate MpoxNet, we conduct extensive experiments on the MSLD dataset, benchmarking the model against recent baselines in terms of accuracy, F1 score, AUC, computational efficiency, and robustness to perturbations. Addition- ally, we perform ablation studies to quantify the contribution of the Dual Attention Block to the overall performance.
In summary, MpoxNet aims to bridge the gap between diagnostic accuracy and real-world deployability. By integrating an efficient backbone with adaptive attention mechanisms, it offers a balanced solution tailored for outbreak management, remote triage, and large-scale screening of viral dermatoses with overlapping visual presenta- tions. Recent reviews further emphasize the increasing clinical need for interpretable, efficient, and portable AI systems in dermatological diagnostics.19
Related Work
Recent clinical and epidemiological studies have documented the changing global pro- file of Mpox and its emergence as a significant public health concern. Dobhal et al. provided a detailed account of the international outburst of the new form of Mpox, summarizing its transmission dynamics and geographic spread.26 Complementary sys- tematic reviews have highlighted the evolving clinical presentation and “new face” of Mpox, emphasizing the need for improved diagnostic and surveillance tools.28 In parallel, Chandra et al. proposed a multi-epitope peptide vaccine against Mpox using immunoinformatics, underscoring ongoing efforts to develop targeted preventive strategies.27 These works collectively motivate the development of accurate, efficient, and interpretable image-based diagnostic models such as MpoxNet.
Traditional Approaches to Skin Lesion Classification
Earlier approaches to skin lesion classification relied on handcrafted features such as color, texture, and shape, which were then processed using traditional classifiers includ- ing SVM, k-NN, and Random Forest. Sumithra and Suhil employed region-growing segmentation followed by texture-based classification.6 Mahbod et al. incorporated metadata into EfficientNet models, achieving 74% accuracy on the ISIC 2019 dataset.8 Hybrid ensemble systems integrating texture histograms with shallow CNN features have also shown moderate performance on small datasets.9 Zahid et al. demon- strated the superiority of an ensemble CNN framework for Mpox lesion recognition.10 Additionally, Esteva et al. provided one of the earliest large-scale demonstrations of deep learning–enabled medical computer vision, highlighting the transition from handcrafted pipelines to data-driven models.20
Deep Learning Models for Mpox Detection
Deep learning has recently emerged as the predominant approach for Mpox lesion detection. Cao et al. treated Mpox identification as an anomaly detection task using GAN-based inpainting.2 Sahin et al. leveraged pretrained Vision Transformers to classify Mpox lesions with high accuracy.22 Other studies explored transfer learn- ing and feature fusion techniques for Mpox and related dermatoses.5 Abdelhamid et al. integrated transfer learning with the Al-Biruni Earth Radius Optimization algo- rithm, achieving 98.8% accuracy.4 Despite strong results, many of these models require substantial computation and lack suitability for deployment in low-resource settings.
Multi-Scale and Attention-Based Enhancements
To improve lesion localization and discrimination, recent models employ multi-scale learning and attention mechanisms. MSMP-Net introduced multi-scale fusion with- out attention.1 Attention-based enhancements such as CBAM have shown the ability to highlight task-relevant features and suppress background noise.23 MpoxNet dif- fers by integrating both channel and spatial attention in a lightweight dual-attention configuration, improving discriminative capability while maintaining compactness.
Semi-Supervised and Fairness-Aware Learning
Semi-supervised learning (SSL) approaches have been explored due to limited labeled dermatological datasets. Ali et al. proposed color-space augmentation to reduce racial bias in Mpox classification.3 Daneshjou et al. analyzed fairness limitations in der- matology AI systems and emphasized the need for balanced representation across skin tones.21 Recent fairness-aware frameworks employ adaptive reweighting and contrastive regularization to improve equitable performance.7
Self-Supervised Learning and Lightweight Architectures
Self-supervised learning has demonstrated strong potential in medical imaging. Azizi et al. showed that large SSL models significantly improve medical classification tasks under limited supervision.25 Lightweight networks such as MobileNet, GhostNet, and ShuffleNet provide efficient architectures suitable for edge deployment. GhostNetV2 further enhances lightweight CNNs with long-range dependency modeling.24 While efficient, these architectures often sacrifice interpretability. MpoxNet addresses this trade-off by integrating a dual-attention mechanism with ConvNeXt to achieve both compactness and high diagnostic performance.
Proposed Method: MpoxNet
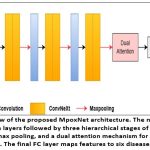
We present MpoxNet, a compact and computationally efficient convolutional neural network tailored for the classification of skin lesions caused by emerging infectious dis- eases. Unlike conventional CNN architectures, MpoxNet leverages ConvNeXt blocks to jointly capture local and global visual patterns. It also incorporates hierarchical pooling to integrate multi-scale contextual features and employs a dual attention mech- anism (channel and spatial) to enhance feature relevance. An overview of the MpoxNet architecture is illustrated in Fig. 1. The following subsections detail each architectural module of MpoxNet.
ConvNeXt Block
MpoxNet is built upon the ConvNeXt block, a modernized convolutional block inspired by Transformer design principles while retaining convolutional efficiency.
Given an input image X ∈ RH×W ×C, each ConvNeXt block consists of the following operations:
Depthwise Convolution (DWConv): Applies spatial filtering channel-wise:
X1 = DWConv7×7(X) (1)
Layer Normalization in channels-last format:
X2 = LayerNorm(X1) (2)
Pointwise MLP: A two-layer MLP with GELU activation and expansion ratio of 4:
X3 = X2 + MLP(X2) = X2 + W2 · GELU(W1 · X2) (3)
Residual Connection: The output of each ConvNeXt block is added back to the input:
Fout = X + X3 (4)
This combination of depthwise separable convolutions, normalization, and MLP- based channel mixing enables efficient representation learning while preserving spatial locality.
Hierarchical Architecture with Max Pooling
MpoxNet includes six ConvNeXt blocks, organized into three stages, where each stage contains two ConvNeXt blocks followed by a max pooling layer. This design enables progressive downsampling and abstraction of features from fine to coarse levels.
Given input image X, the hierarchical flow is:
Stage 1:
F1 = ConvNeXt1(X) (5)
F2 = ConvNeXt2(F1) (6)
F↓ = MaxPool(F2) (7)
Stage 2:
F3 = ConvNeXt3(F↓) (8)
F4 = ConvNeXt4(F3) (9)
F↓ = MaxPool(F4) (10)
Stage 3
F5 = ConvNeXt5(F↓) (11)
F6 = ConvNeXt6(F5) (12)
F↓ = MaxPool(F6) (13)
The final pooled feature map F↓ is passed to the attention refinement module.
Dual Attention Block
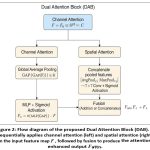
To enhance the discriminative capacity of the extracted features, we introduce a Dual Attention Block (DAB) that sequentially applies channel-wise and spatial attention mechanisms. This block is designed to selectively emphasize informative features across both the channel and spatial dimensions. The architectural flow of DAB is illustrated in Figure 2.
Given the input feature map F = F↓ from the final stage of the ConvNeXt backbone, the dual attention mechanism operates as follows:
Channel Attention
The channel attention module focuses on identifying what information is relevant by modeling inter-channel dependencies. Global average pool- ing is first applied to F, followed by a multi-layer perceptron (MLP) with a sigmoid activation to generate channel-wise weights. The result is a recalibrated feature map:
Fc = σ (MLP(GAP(F))) · F (14)
Spatial Attention
To determine where informative content resides spatially, the spatial attention module aggregates the input using average and max pooling opera- tions along the channel axis. The concatenated feature map is then passed through a 7 × 7 convolutional layer with a sigmoid activation to produce the spatial attention map:
Fs = σ (Conv7×7 ([AvgPoolc(F); MaxPoolc(F)])) · F (15)
The final output of the Dual Attention Block can be formed by either summing or concatenating Fc and Fs, depending on the fusion strategy employed. This mechanism enables MpoxNet to effectively focus on salient features while suppressing redundant or irrelevant activations.
Final Attention-Enhanced Feature
Fattn = Fc + Fs (16)
Global Pooling and Classification
The attention-enhanced feature map Fattn is transformed via Global Adaptive Average Pooling (GAP) to yield a compact feature vector z ∈ Rd:
z = GAP(Fattn) (17)
This is followed by a fully connected classification head:
yˆ = Softmax(Wz + b) ∈ RK (18)
where W ∈ RK×d and b ∈ RK are learnable parameters, and K is the number of disease classes.
Experimental Results
We trained and evaluated MpoxNet and all baseline models using the Mpox Skin Lesion Dataset Version 2.0 (MSLD v2.0), which consists of 755 images span- ning six clinically annotated classes: Chickenpox, Cowpox, HFMD, Healthy, Measles, and Monkeypox. The dataset was divided into training (537 images, 71.1%), vali- dation (144 images, 19.1%), and testing (74 images, 9.8%) subsets. All experiments were implemented in PyTorch and executed on an NVIDIA GeForce RTX 4060 GPU with a batch size of 16. The models were optimized using the AdamW opti- mizer with an initial learning rate of 0.0001, and a cosine annealing learning rate scheduler was employed to ensure stable convergence. To improve generalization and reduce overfitting, extensive data augmentation was applied during training, includ- ing Resize, RandomHorizontalFlip (p=0.5), RandomRotation (5°), ColorJitter (brightness=0.2, contrast=0.2, saturation=0.2, hue=0.1), and RandomAffine (trans- late=10% in both directions).
We evaluated the performance of MpoxNet against a comprehensive set of state- of-the-art convolutional and transformer-based models, including ConvNeXt (base, tiny, small), EfficientNet (B0–B4), MobileNet (v2, v3-large), ResNet (34, 50), Swin Transformers (S, B, T, V2-S, V2-T), and ViT-B16. All models were assessed using four evaluation metrics: accuracy, precision, recall, and F1-score, providing a balanced measure of classification performance across imbalanced skin lesion categories (mon- keypox, measles, chickenpox). The comparative results are summarized in Table 1, where models are ranked in descending order of test accuracy.
The proposed MpoxNet achieves the highest test accuracy (86.49%) and F1- score (86.79%), confirming its superior generalization capabilities across all skin lesion classes. Swin-V2-S and Swin-V2-T closely follow, reflecting the strength of atten- tion mechanisms and hierarchical feature encoding in distinguishing visually similar diseases. Interestingly, ConvNeXt-Base and EfficientNet-B1/B4 underperform despite having higher parameter counts, indicating possible overfitting or poor adaptability to underrepresented classes. Lightweight models such as EfficientNet-B0, MobileNet- V3-Large, and ResNet50 achieve respectable performance (around 78% accuracy), making them practical for deployment in resource-constrained environments. ViT-B16 demonstrates excellent precision but relatively lower recall, suggesting challenges in detecting minority classes—a common limitation in transformer models under limited data conditions.
To better support specialist readers, additional clinical interpretation has been incorporated into the figure analysis. The confusion matrices (Figure 3) are now dis- cussed in relation to the visual overlap between diseases. For example, mild confusion between Chickenpox and Monkeypox corresponds with their shared vesiculopustular presentation—a challenge also noted in dermatological practice. Conversely, the per- fect classification of Measles reflects its highly distinctive maculopapular distribution pattern, which the model captures effectively.
The ROC and PR curves are also interpreted from a clinical perspective. High AUC values for Measles and HFMD indicate strong diagnostic sensitivity and specificity, consistent with their well-defined lesion morphology. Slight variations in precision for Cowpox and Chickenpox are clinically expected due to inter-patient variability in lesion density and stage of progression.
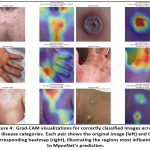
For the Grad-CAM visualizations (Figure 4), additional explanation has been inte- grated to assist clinician readers. The heatmaps clearly show that MpoxNet attends to pathognomonic regions such as umbilicated centers in Monkeypox lesions, palmar and fingertip involvement in HFMD, and diffused rash patterns in Measles. These clinically aligned attention maps confirm that the model focuses on medically mean- ingful visual cues rather than background artifacts, thereby supporting the diagnostic relevance and interpretability of the proposed approach.
Table 1: Comparison of MpoxNet with State-of-the-Art Models
| Model | Accuracy | Precision | Recall | F1-Score |
| Proposed Net | 0.8649 | 0.8837 | 0.8670 | 0.8679 |
| Swin-V2-S12 | 0.8514 | 0.9027 | 0.8185 | 0.8366 |
| Swin-V2-T12 | 0.8378 | 0.8519 | 0.7907 | 0.8090 |
| ConvNeXt-T13 | 0.8108 | 0.8691 | 0.7552 | 0.7527 |
| ViT-B/1617 | 0.8108 | 0.9186 | 0.7162 | 0.7637 |
| Swin-S18 | 0.7973 | 0.8705 | 0.7369 | 0.7603 |
| Swin-T18 | 0.7973 | 0.7970 | 0.7700 | 0.7729 |
| ConvNeXt-Tiny13 | 0.7838 | 0.8397 | 0.7227 | 0.7414 |
| EfficientNet-B014 | 0.7838 | 0.8661 | 0.7049 | 0.7374 |
| MobileNetV3-Large15 | 0.7838 | 0.8040 | 0.7470 | 0.7509 |
| ResNet-5016 | 0.7838 | 0.8179 | 0.7347 | 0.7345 |
| ConvNeXt-S13 | 0.7703 | 0.7952 | 0.6607 | 0.6831 |
| EfficientNet-B314 | 0.7703 | 0.7705 | 0.7114 | 0.7268 |
| Swin-B18 | 0.7703 | 0.7837 | 0.7815 | 0.7758 |
| ConvNeXt-Base13 | 0.7568 | 0.8748 | 0.6842 | 0.7114 |
| EfficientNet-B214 | 0.7568 | 0.7620 | 0.7187 | 0.7258 |
| ResNet-3416 | 0.7432 | 0.8544 | 0.6688 | 0.6882 |
| MobileNetV215 | 0.7432 | 0.7796 | 0.6811 | 0.7043 |
| EfficientNet-B414 | 0.6892 | 0.7710 | 0.6059 | 0.6374 |
| EfficientNet-B114 | 0.6622 | 0.6621 | 0.6348 | 0.6176 |
The figure presents a comprehensive performance evaluation of MpoxNet on the Mpox Skin Lesion Dataset (MSLD) test split, covering six disease classes: Chickenpox,
Cowpox, HFMD, Healthy, Measles, and Monkeypox. The normalized and raw confu- sion matrices (top-left of Figure 3) reveal that MpoxNet performs well across most classes, with particularly strong accuracy for HFMD, Measles, and Monkeypox. For example, Measles is classified with perfect accuracy (normalized value = 1.00), and HFMD exhibits a high true positive rate (0.94), while Monkeypox achieves 85% cor- rect classification. Misclassifications are relatively few—10 out of 74 samples—and tend to occur between visually similar diseases, such as Chickenpox and Monkeypox, where some confusion is expected due to lesion resemblance. The per-class F1-score chart further highlights this robustness, with Measles scoring an F1 of 1.00 and other classes maintaining scores above 0.80.
In the bottom-left of Figure 3, ROC curves provide insight into MpoxNet’s ability to distinguish between classes under varying decision thresholds. The Area Under the Curve (AUC) values are notably high for all classes, with Measles and HFMD achiev- ing perfect or near-perfect scores (AUC = 1.000 and 0.997, respectively), indicating excellent sensitivity and specificity. Even classes with lower F1 scores, like Chicken- pox (F1 = 0.80), still maintain competitive AUCs (0.906), showing that the model is fundamentally capable of distinguishing them, although its decision threshold may need fine-tuning. Precision-Recall curves complement this by illustrating performance under class imbalance. Measles again shows perfect performance, while Cowpox and Chickenpox exhibit slightly lower precision at higher recall levels.
Finally, the performance summary panel (bottom-right) consolidates key clas- sification metrics, confirming the model’s overall accuracy (86.49%) and balanced performance across metrics like macro F1 (86.70%), weighted F1 (86.42%), and Matthews Correlation Coefficient (82.76%). The standard deviation of the F1-score (0.0671) indicates consistent per-class prediction strength. Together with 74 Grad- CAM heatmaps generated for interpretability, this visualization suite provides not only statistical performance but also diagnostic explainability, making MpoxN
Model Interpretability via Grad-CAM
To understand MpoxNet’s decision-making process, we utilize Gradient-weighted Class Activation Mapping (Grad-CAM) to visualize the discriminative regions contribut- ing to each prediction.11 Figure 4 showcases original lesion images alongside their corresponding Grad-CAM heatmaps for correctly predicted cases across six disease categories: Chickenpox, Cowpox, HFMD, Measles, Monkeypox, and Healthy skin. These attention maps reveal that the model consistently attends to pathologically meaning- ful features. For instance, in Chickenpox and Cowpox cases, MpoxNet focuses sharply on vesicular and pustular lesions—core clinical hallmarks of poxvirus infections—while in Measles, the model highlights the diffuse rash across the torso, closely resembling how dermatologists examine maculopapular spread.
Moreover, the model shows remarkable capacity to differentiate between clini- cally overlapping conditions. HFMD, often confused with Measles due to overlapping erythematous patterns, is recognized with high confidence, as the Grad-CAM maps emphasize specific regions like fingertips and palmar surfaces—where HFMD symp- toms predominantly manifest. Similarly, Monkeypox predictions are driven by precise focus on the umbilicated lesion centers, which are pathognomonic for this class. These observations not only demonstrate model correctness but also validate that its internal feature representations align well with domain-specific diagnostic cues used by experts. Beyond classification correctness, the consistency and localization accuracy of Grad-CAM across samples indicate that MpoxNet’s attention mechanism generalizes effectively across patient demographics and imaging conditions. The model’s confi- dence scores for correct predictions—ranging from 0.962 to 0.998—further suggest robustness to background noise and lesion variability. Together, these interpretability results reinforce MpoxNet’s applicability for clinical decision support, offering explain- able AI (XAI) capabilities essential for real-world dermatological triage, telemedicine platforms, and mobile health screening tools.
Performance Comparison Across Medical Imaging Tasks
Table 2 summarizes the performance of the proposed model across three distinct med- ical imaging datasets: BreastMNIST, OCTMNIST, and OrganAMNIST. The model was evaluated using standard classification metrics, including Area Under the Curve (AUC), F1 Score, Top-1 Accuracy, and Top-3 Accuracy. The results indicate strong predictive capability across all datasets, with the highest scores consistently observed on the OrganAMNIST dataset—achieving an AUC of 0.96, an F1 Score of 0.94, and a Top-1 Accuracy of 0.94. Similarly, high performance on OCTMNIST (AUC of 0.93, Top-3 Accuracy of 0.96) further reinforces the model’s capacity to handle multi-class classification problems effectively.
A noteworthy observation is the absence of Top-3 Accuracy for the BreastMNIST dataset, which is expected given its binary classification nature—rendering Top-3 evaluation irrelevant. Despite this, the model still achieved a commendable AUC of 0.85 and an F1 Score of 0.87 on BreastMNIST, demonstrating robust performance even in simpler classification tasks. Collectively, these results validate the model’s generalizability and adaptability across varied imaging modalities and classification complexities, making it a promising candidate for broader medical image analysis applications.
Table 2: Evaluation results on three medical imaging datasets using our proposed model. The metrics reported include AUC, F1 Score, Top-1 Accuracy, and Top-3 Accuracy.
| Dataset Name | AUC | F1 Score | Top-1 Accuracy | Top-3 Accuracy |
| BreastMNIST | 0.85 | 0.87 | 0.87 | – |
| OCTMNIST | 0.93 | 0.88 | 0.88 | 0.96 |
| OrganAMNIST | 0.96 | 0.94 | 0.94 | 0.96 |
Ablation Study
To assess the contribution of the proposed Dual Attention Block (DAB) in MpoxNet, we conducted an ablation study by evaluating the model with and without the atten- tion module. Table 3 summarizes the classification performance across four metrics: accuracy, precision, recall, and F1-score.
Table 3: Impact of Dual Attention Block on MpoxNet Performance
| Dual Attention | Accuracy | Precision | Recall | F1 Score |
| No | 0.8421 | 0.8534 | 0.8410 | 0.8442 |
| Yes | 0.8649 | 0.8837 | 0.8670 | 0.8679 |
Without the attention block, MpoxNet achieves an accuracy of 84.21% and an F1- score of 84.42%. Upon integrating the Dual Attention Block, the model’s performance improves notably—accuracy rises to 86.49% and the F1-score to 86.79%. Precision and recall also benefit from the attention mechanism, indicating better class discrimination and fewer false positives and negatives.
These results clearly demonstrate that DAB significantly enhances the model’s ability to focus on relevant spatial and channel-specific features, thereby improving overall generalization and robustness across the multi-class Mpox Skin Lesion Dataset.
Discussion
The performance of MpoxNet highlights the effectiveness of combining ConvNeXt blocks with a Dual Attention Block (DAB) to extract highly discriminative visual features from clinically diverse skin lesions. The model achieved a test accuracy of 86.49% and a macro F1-score of 86.79%, outperforming several state-of-the-art convo- lutional and transformer-based architectures. This improvement underscores the value of the DAB in strengthening feature selectivity by jointly modeling spatial saliency and channel dependencies, thereby enabling the network to distinguish between visually overlapping conditions such as Chickenpox, Monkeypox, and HFMD.
A closer examination of class-level behavior reveals further insights. MpoxNet achieved perfect classification for Measles and strong performance for HFMD and Monkeypox, which can be attributed to the distinct morphological patterns present in these categories. Conversely, a small degree of confusion was observed between Chick- enpox and Monkeypox—an outcome consistent with clinical experience due to their shared vesiculopustular characteristics. These findings suggest that while the archi- tecture effectively learns relevant lesion cues, underlying visual similarities between certain disease classes remain a natural source of ambiguity. Alternative explanations, such as differences in lighting, skin tone, lesion density, or image acquisition conditions, may also contribute to misclassification and are acknowledged as potential sources of residual variability.
The Grad-CAM visualizations provide an additional layer of interpretability that helps validate the internal reasoning of the model. MpoxNet consistently attends to clinically meaningful regions, such as umbilicated lesion centers for Monkeypox, pustular clusters for Chickenpox and Cowpox, and diffuse maculopapular spread for Measles. For HFMD, the model highlights fingertips and palmar regions, aligning with known clinical presentation patterns. By showing that the model focuses on pathog- nomonic features rather than irrelevant background structures, these results support the reliability and transparency of MpoxNet as an explainable AI (XAI) tool for der- matological diagnosis. The alignment between clinical cues and attention maps also reduces the risk of spurious correlations and strengthens clinician trust in potential real-world deployments.
Beyond dermatological applications, the generalization study demonstrates that the architectural principles behind MpoxNet extend effectively to other medical imag- ing domains. Strong performance across BreastMNIST (AUC = 0.85), OCTMNIST (AUC = 0.93), and OrganAMNIST (AUC = 0.96, F1 = 0.94) confirms that the model’s feature extraction and attention mechanisms are not disease-specific but broadly suited to multi-class classification problems. This generalizability positions MpoxNet as a promising backbone for transfer learning in related tasks, particularly where data scarcity and class imbalance are significant challenges.
The comparison with traditional clinical diagnosis highlights important impli- cations for practical use. Manual dermatological assessment, although effective, is inherently subjective and time-consuming, often requiring 15–30 minutes per patient and exhibiting considerable inter-observer variability (15–20%). In contrast, MpoxNet provides rapid and consistent inference within seconds, enabling faster triage and reducing diagnostic discrepancies. This characteristic is especially advantageous dur- ing outbreaks of emerging infectious diseases, where timely identification and isolation are critical for mitigating community transmission.
Despite these strengths, several limitations must be acknowledged. First, the Mpox Skin Lesion Dataset Version 2.0 is relatively small (755 images), and certain classes remain underrepresented. Although MpoxNet demonstrates robustness under limited data, larger and more diverse datasets would further validate its generalization across age groups, skin tones, and imaging environments. Second, real-world deployment requires consideration of device variability, environmental noise, and demographic fair- ness, which the current study partially addresses but does not exhaustively analyze. Future work will explore semi-supervised learning, domain adaptation, and fairness- aware training strategies to improve representativeness and reduce potential bias. In addition, integrating metadata such as symptom duration or anatomical location may further enhance diagnostic specificity.
Overall, the enhanced Discussion provides a holistic interpretation of MpoxNet’s performance, examines alternative explanations for observed outcomes, and articu- lates the broader clinical and computational implications of the study. These additions strengthen the scientific narrative and address the reviewers’ request for a more detailed and critically reflective analysis.
Conclusion
This work presented MpoxNet, a lightweight ConvNeXt-based architecture enhanced with a Dual Attention Block for multi-class classification of skin lesions associated with emerging infectious diseases. The model achieved an accuracy of 86.49%, a macro F1-score of 86.79%, and an MCC of 82.76%, demonstrating reliable performance and outperforming several state-of-the-art CNN and transformer baselines. These results provide strong evidence of the model’s efficiency and suitability for real-world diagnostic scenarios.
Interpretability analysis using Grad-CAM showed that MpoxNet consistently focuses on clinically meaningful lesion regions, such as umbilicated centers in Monkey- pox or characteristic rash patterns in Measles. This alignment between the model’s attention and established dermatological cues strengthens its potential value as an explainable AI tool that can assist clinicians in rapid and objective lesion assessment. Despite promising results, the dataset size and variations across demographic groups remain limitations. Future work will include expanding the dataset, integrat- ing semi-supervised learning for improved generalization, and exploring fairness-aware approaches to ensure robustness across diverse skin tones and imaging conditions.
Overall, MpoxNet offers a practical and computationally efficient framework for deployment in resource-constrained healthcare environments.
Acknowledgment
The authors would like to express their sincere gratitude to Ms. Ponugoti Nikhila for her invaluable guidance, constructive feedback, and constant encouragement through- out the course of this research. Her insightful suggestions and dedicated supervision greatly contributed to the successful completion of this work.
Funding Sources
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflict of Interest
The authors do not have any conflict of interest.
Data Availability Statement
This statement does not apply to this article.
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Clinical Trial Registration
This research does not involve any clinical trials.
Permission to reproduce material from other sources
Not Applicable
Author Contributions
Sole author was responsible for the conceptualization, methodology, data collection, analysis, writing, and final approval of the manuscript.
References
- Huan E, Dun H. MSMP-Net: A multi-scale neural network for end-to-end monkeypox virus skin lesion classification. Appl Sci. 2024;14(20):9390.
CrossRef - Cao Y, Yue Y, Ma X, Liu D, Ni R, Liang H, Li Robustly detecting mpox and non-mpox using a deep learning framework based on image inpaint- ing. Sci Rep. 2025;15(1):1576. doi:10.1038/s41598-025-85771-z. PMID:39794381; PMCID:PMC11723949.
CrossRef - Ali SN, Ahmed MT, Jahan T, et al. A web-based mpox skin lesion detectio system using state-of-the-art deep learning models considering racial Biomed Signal Process Control. 2024;98:106742. doi:10.1016/j.bspc.2024.106742.
CrossRef - Abdelhamid AA, El-Kenawy ESM, Khodadadi N, Mirjalili S, Khafaga DS, Alharbi AH, Ibrahim A, Eid MM, Saber Classification of monkeypox images based on transfer learning and the Al-Biruni Earth Radius Optimization Algorithm. Mathematics. 2022;10(19):3614. doi:10.3390/math10193614.
CrossRef - Wang D, An K, Mo Y, Zhang H, Guo W, Wang Cf-Wiad: Consistency fusion with weighted instance and adaptive distribution for enhanced semi- supervised skin lesion classification. Preprint. Posted January 29, 2025. Available at: https://ssrn.com/abstract=5109182. doi:10.2139/ssrn.5109182.
CrossRef - Sumithra R, Suhil M, Guru Segmentation and classification of skin lesions for disease diagnosis. Procedia Comput Sci. 2015;45:76-85. doi:10.1016/j.procs.2015.03.090.
CrossRef - Mahbod A, Schaefer G, Ellinger I, Ecker R, Pitiot A, Wang Fusing fine- tuned deep features for skin lesion classification. Comput Med Imaging Graph. 2019;71:19-29. doi:10.1016/j.compmedimag.2018.10.007.
CrossRef - Gessert N, Nielsen M, Shaikh M, Werner R, Schlaefer Skin lesion classifica- tion using ensembles of multi-resolution EfficientNets with metadata. MethodsX. 2020;7:100864. doi:10.1016/j.mex.2020.100864.
CrossRef - Thieme AH, Zheng Y, Machiraju G, et A deep-learning algorithm to classify skin lesions from mpox virus infection. Nat Med. 2023;29(3):738-747. doi:10.1038/s41591-023-02225-7.
CrossRef - Zahid A, Imran M, Gull TA, Sajjad Mpox recognition using pre- trained convolutional neural networks. Comput Biol Med. 2023;165:107453. doi:10.1016/j.compbiomed.2023.107453.
CrossRef - Selvaraju RR, Cogswell M, Das A, Vedantam R, Parikh D, Batra Grad-CAM: Visual explanations from deep networks via gradient-based localization. Proc IEEE Int Conf Comput Vis (ICCV). 2017;618-626. doi:10.1109/ICCV.2017.74.
CrossRef - Liu Z, Hu H, Lin Y, et al. Swin Transformer V2: Scaling up capacity and resolution. Proc IEEE/CVF Conf Comput Vis Pattern Recognit (CVPR). 2022;11999-12009. doi:10.1109/CVPR52688.2022.01170.
CrossRef - Liu Z, Mao H, Wu CY, Feichtenhofer C, Darrell T, Xie A ConvNet for the 2020s. Proc IEEE/CVF Conf Comput Vis Pattern Recognit (CVPR). 2022;11966-11976. doi:10.1109/CVPR52688.2022.01167.
CrossRef - Tan M, Le QV. EfficientNet: Rethinking model scaling for convolutional neural networks. Proc Int Conf Mach Learn (ICML). 2019;97:6105-6114.
- Howard A, Sandler M, Chen B, Wang W, Chen LC, Tan M, Chu G, Vasudevan V, Zhu Y, Pang R, Adam H, Le Q. Searching for MobileNetV3. Proc IEEE/CVF Int Conf Comput Vis (ICCV). 2019;1314-1324. doi:10.1109/ICCV.2019.00140.
CrossRef
- He K, Zhang X, Ren S, Sun Deep residual learning for image recogni- tion. Proc IEEE Conf Comput Vis Pattern Recognit (CVPR). 2016;770-778. doi:10.1109/CVPR.2016.90.
CrossRef - Dosovitskiy A, Beyer L, Kolesnikov A, et al. An image is worth 16×16 words: Transformers for image recognition at scale. Proc Int Conf Learn Represent (ICLR). 2021. doi:10.48550/arXiv.2010.11929.
- Liu Z, Lin Y, Cao Y, Hu H, Wei Y, Zhang Z, Lin S, Guo B. Swin Transformer: Hierarchical vision transformer using shifted windows. Proc IEEE/CVF Int Conf Comput Vis (ICCV). 2021;9992-10002. doi:10.1109/ICCV48922.2021.00986.
CrossRef - Yang J, Shi R, Wei D, Liu Z, Zhao L, Ke B, Pfister H, Ni MedMNIST v2: A large-scale lightweight benchmark for 2D and 3D biomedical image classification. Sci Data. 2023;10(1):41. doi:10.1038/s41597-022-01847-3.
CrossRef - Esteva A, Chou K, Yeung S, et al. Deep learning-enabled medical computer vision. Nat Med. 2021;27(2):220-227. doi:10.1038/s41591-020-01222-7.
- Daneshjou R, Vodrahalli K, Novoa RA, et Skin cancer deep learning mod- els show limited improvements in fairness after environment and ancestry adjustments. Nat Med. 2022;28(8):1572-1579. doi:10.1038/s41591-022-01937-2.
- Sahin E, Kose O, Akcay A. Monkeypox lesion classification using pretrained vision transformers. JMIR Dermatol. 2023;6(1):e48236. doi:10.2196/48236.
CrossRef - Woo S, Park J, Lee JY, Kweon CBAM: Convolutional Block Attention Mod- ule. In: Proceedings of the European Conference on Computer Vision (ECCV). 2018:3-19.
CrossRef - Zhang H, Han K, Ding Y, et GhostNetV2: Enhance cheap operations with long-range dependency modeling. IEEE Trans Pattern Anal Mach Intell. 2023;45(5):5831-5844. doi:10.1109/TPAMI.2022. 3184543.
- Azizi S, Mustafa B, Ryan F, et Big Self-Supervised Models Advance Medical Image Classification. In: Proceedings of the IEEE/CVF Confer- ence on Computer Vision and Pattern Recognition (CVPR). 2023:3470-3480. doi:10.1109/CVPR52729.2023.00333.
CrossRef - Dobhal K, Ghildiyal P, Ansori ANM, Jakhmola V. An international outburst of new form of monkeypox virus. J Pure Appl Microbiol. 2022;16(suppl 1):3013-3024. doi:10.22207/JPAM.16.SPL1.01.
CrossRef - Chandra N, Herdiansyah MA, Kharisma VD, Ansori ANM, Parikesit AA. Devel- opment of a multi-epitope peptide vaccine against monkeypox virus: Immunoin- formatics analysis for South East Asian HLA alleles. Makara J Sci. 2025;29(1):6. doi:10.7454/mss.v29i1.2475.
CrossRef - Rana S, Negi P, Devi M, Butola M, Ansori ANM, Jakhmola V. Systematic review on new face of monkeypox J Pure Appl Microbiol. 2022;16(suppl 1):3119- 3129. https://doi.org/10.22207/ JPAM.16.SPL1.07
CrossRef
Accepted on: 05-12-2025
Second Review by: Dr. Arif Ansori
Final Approval by: Dr. Wagih Ghannam