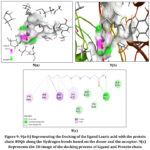
An In-Silico Molecular Docking Analysis of Phytochemicals from Carica papaya, Moringa oleifera, and Tinospora cordifolia as Potential Dengue Virus Inhibitors
Department of Pharmacology, Vignan Institute of Pharmaceutical Technology, Duvvada, Visakhapatnam, Andhra Pradesh, India.
Corresponding Author E-mail: saisudheer9090@gmail.com
DOI : http://dx.doi.org/10.13005/bbra/3505
ABSTRACT:The search for effective antiviral agents against dengue virus (DENV) remains a global priority due to the absence of specific therapeutics. The present study evaluates selected phytochemicals from Carica papaya, Moringa oleifera, and Tinospora cordifolia using an in-silico molecular docking approach to identify compounds with potential dengue-related inhibitory activity. Phytochemicals were screened based on drug-likeness and ADME properties, and their disease-associated gene targets were identified through database mining. Venn analysis identified plasminogen (PLG), a host protein implicated in dengue-associated thrombocytopenia and vascular complications, as a common target linked to both dengue pathophysiology and the selected phytochemicals. The human plasminogen protein (PDB ID: 8UQ6) was selected for molecular docking using PyRx, and interaction analyses were performed with BIOVIA Discovery Studio. Among the evaluated compounds, carpaine from Carica papaya exhibited the highest binding affinity (−9.2 kcal/mol), followed by hesperetin from Moringa oleifera (−8.3 kcal/mol), whereas phytochemicals from Tinospora cordifolia showed comparatively lower affinities. These findings suggest that Carica papaya and Moringa oleifera contain phytochemicals capable of interacting with dengue-associated host molecular pathways. However, as molecular docking provides predictive insights only, these results are limited to in-silico observations and require further in vitro, in vivo, and clinical validation.
KEYWORDS:Carica papaya; Dengue virus; Moringa oleifera; Molecular docking; Plasminogen; Tinospora cordifolia
Introduction
Dengue virus
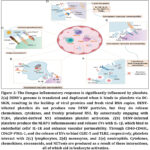
Dengue is a globally prevalent arboviral disease transmitted by Aedes mosquitoes, with clinical manifestations ranging from asymptomatic infection to severe dengue characterized by hemorrhage, organ dysfunction, and vascular leakage.1 Increasing evidence highlights the central role of platelets in dengue pathogenesis. Dengue virus (DENV)–infected platelets actively participate in inflammatory and immunological responses by releasing mediators such as RANTES, macrophage migration inhibitory factor (MIF), and interleukin-1β (IL-1β).2,3 Elevated levels of these mediators in severe dengue promote leukocyte activation, platelet–leukocyte aggregation, and endothelial dysfunction, ultimately contributing to vascular leakage.4–9 Thrombocytopenia is a hallmark hematological feature of dengue and correlates strongly with disease severity.12,13 It is associated with enhanced platelet activation, increased platelet leukocyte interactions, and accelerated clearance of activated or apoptotic platelets by monocytes and macrophages.4,14,15 These processes collectively underscore the importance of platelet dysfunction in dengue-associated complications.
History and Epidemiology
Historical records indicate that dengue-like illness was first documented in China between 265 and 420 AD, with later outbreaks reported in the West Indies, Central America, and parts of Africa and Asia between the 17th and 18th centuries.47 The four classical dengue virus serotypes (DENV-1 to DENV-4) are thought to have evolved through both sylvatic transmission cycles involving non-human primates and mosquitoes, and urban transmission between humans and Aedes mosquitoes, facilitated by human migration and trade.45,47,48 While some evidence suggests an African origin with global dissemination via historical trade routes, other studies propose emergence from sylvatic cycles in Southeast Asia.45,47,48 A genetically distinct fifth serotype (DENV-5) was later identified in Malaysia in 2007, with a probable sylvatic origin.46,50,51
Currently, dengue is endemic in more than 100 countries across tropical and subtropical regions, with a marked increase in incidence over recent decades.46,52 Southeast Asia remains a hyperendemic region, with all serotypes co-circulating and a large population at risk.53,54 Countries such as India and Sri Lanka experience recurrent outbreaks, with regional variation in circulating serotypes and significant disease burden.53–56
Clinical Aspects
DENV infection presents with a broad clinical spectrum ranging from mild dengue to severe disease. Following an incubation period of 4–10 days, patients typically develop acute febrile illness accompanied by myalgia, headache, and malaise.1,16 Disease progression in severe cases is marked by leukopenia, thrombocytopenia, systemic inflammation, and increased plasma leakage, which may lead to hypotension, bleeding, and organ failure.1,17 Platelet activation and apoptosis are prominent during dengue infection and are more pronounced in severe disease. Activated platelets exhibit surface expression of P-selectin, CD63, and phosphatidylserine, facilitating interactions with leukocytes and endothelial cells and promoting inflammatory and thrombo-inflammatory responses.4,15,32 These platelet-mediated processes play a key role in disease severity.
Cycle of Replication
DENV infects multiple host cell types, including monocytes, macrophages, endothelial cells, and platelets, through receptors such as DC-SIGN, heparan sulfate proteoglycans, and mannose receptors.18–21 Viral entry is mediated by the envelope glycoprotein, followed by endocytosis and translation of viral RNA into structural and non-structural proteins.22,23 In platelets, DENV binding results in viral RNA accumulation and NS1 expression; however, productive viral replication does not occur. Instead, platelets undergo an abortive infection that triggers inflammatory signaling, platelet activation, and cytokine release, contributing to vascular dysfunction without the production of infectious virions.2,19,20
Mechanisms of Thrombocytopenia
Dengue-associated thrombocytopenia results from a combination of reduced platelet production and enhanced peripheral platelet clearance.24,25 Early in infection, transient suppression of megakaryopoiesis may occur due to bone marrow dysfunction, while disease progression is dominated by peripheral mechanisms.26–28 Peripheral platelet clearance is driven by platelet activation, apoptosis, immune-mediated opsonization, complement activation, and increased platelet adhesion to leukocytes and endothelial cells.15,27,32–34 Additional mechanisms, including von Willebrand factor–mediated platelet desialylation and antibody-dependent platelet destruction, further exacerbate platelet loss and correlate with disease severity.36–41 Collectively, these processes highlight the multifactorial nature of dengue-associated thrombocytopenia and support the exploration of host-targeted molecular strategies aimed at modulating platelet-related pathways involved in dengue pathophysiology.44
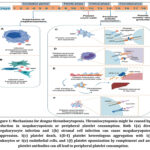 |
Figure 1: Mechanisms for dengue thrombocytopenia. Thrombocytopenia might be caused by a reduction in megakaryopoiesis or peripheral platelet consumption.
|
Platelet counts in dengue reach their lowest levels around defervescence and disease progression, even though recovery from acute bone marrow suppression and restoration of hematopoiesis occur before the critical phase, with platelet count normalization seen only during convalescence.27,32 This temporal mismatch suggests that peripheral platelet clearance plays a major role in dengue-associated thrombocytopenia.15,21 During infection, platelets exhibit marked activation and apoptotic features, including Translocation of P-selectin and CD63, activation of integrin αIIbβ3, exposure to phosphatidylserine, and mitochondrial dysfunction.15,21 These changes are more pronounced in severe dengue and are closely linked to enhanced monocyte and macrophage mediated phagocytosis of apoptotic platelets via phosphatidylserine recognition, and platelet-monocyte interactions are facilitated by surface P-selectin and platelet activation.4,15,32
Thrombocytopenia in dengue is strongly associated with enhanced platelet interactions with monocytes, other leukocytes, and endothelial cells, leading to increased platelet activation and clearance.4,10,33,34 Elevated platelet–monocyte and platelet–neutrophil aggregates, platelet adhesion to endothelial cells in organs such as the liver, spleen, and heart, and increased platelet deposition in the microvasculature contribute to disease severity.4,33–35 DENV infection also promotes endothelial release of von Willebrand factor (vWF), increasing platelet binding, desialylation via GPIbα, and subsequent hepatic clearance.15,36,37 Antibody-mediated mechanisms further exacerbate thrombocytopenia, as platelet-associated IgG and IgM—particularly anti-NS1 antibodies—cross-react with platelet integrins, enhancing opsonization, complement activation, and platelet destruction.38–41 Additionally, direct binding of NS1 to platelets activates the complement system, with increased complement activity correlating inversely with platelet counts.2,42,43 Collectively, dengue-associated thrombocytopenia results from a combination of bone marrow suppression and peripheral mechanisms including platelet activation, immune-mediated clearance, complement lysis, aggregation, and platelet–cell interactions.15,44
Carica papaya
Carica papaya L. is a fast-growing herbaceous plant of the family Caricaceae, widely cultivated in tropical and subtropical regions. The plant is characterized by a soft, unbranched stem, palmately lobed leaves, and a large fleshy fruit containing numerous seeds. C. papaya is rich in bioactive compounds, including alkaloids (notably carpaine), flavonoids, phenolic acids, and proteolytic enzymes such as papain.62 These phytochemicals are reported to exhibit antioxidant, anti-inflammatory, and platelet-modulating properties, which have attracted attention for their potential role in dengue-associated thrombocytopenia. Owing to its diverse pharmacological profile and traditional use in managing febrile illnesses, C. papaya represents a promising source of host-targeted compounds for dengue-related therapeutic exploration.
Moringa oleifera
Moringa oleifera Lam., commonly known as the drumstick tree, is a rapidly growing deciduous plant belonging to the family Moringaceae and distributed across tropical and subtropical regions. The plant contains a wide spectrum of bioactive constituents, including flavonoids (quercetin, kaempferol), isothiocyanates, phenolic acids, vitamins, and essential amino acids.64 These compounds are known for their antioxidant, anti-inflammatory, hematopoietic, and immunomodulatory activities. Given the involvement of oxidative stress and immune dysregulation in dengue pathogenesis, M. oleifera phytochemicals are of particular interest as potential modulators of host inflammatory and platelet-related pathways relevant to dengue infection.
Tinospora cordifolia
Tinospora cordifolia (Willd.) Miers is a medicinal climber of the family Menispermaceae, widely distributed in tropical regions of India. The plant is characterized by succulent climbing stems with aerial roots and heart-shaped leaves. It possesses a rich phytochemical profile comprising alkaloids (berberine, palmatine), diterpenoids, glycosides (tinosporaside, cordifolide), flavonoids, and immunomodulatory polysaccharides.63 Traditionally known as “Amrita,” T. cordifolia has been reported to exhibit antipyretic, anti-inflammatory, antioxidant, and immunomodulatory properties. These biological activities support its selection for in-silico evaluation as a potential source of host-directed compounds in dengue-associated inflammatory and immune responses.
In-silico approaches such as molecular docking offer a rapid and cost-effective strategy to predict molecular interactions between viral or host targets and bioactive compounds prior to experimental validation. Medicinal plants such as Carica papaya, Moringa oleifera, and Tinospora cordifolia have been traditionally used in dengue management, particularly for thrombocytopenia and immune modulation. However, limited information is available regarding their molecular interactions with dengue-associated targets. Therefore, the present study aims to computationally evaluate selected phytochemicals from these plants against dengue-related gene targets to identify potential inhibitors and provide a rational basis for further experimental investigation[60,61].
Given the involvement of plasminogen in platelet dysfunction during dengue infection, targeting PLG through phytochemical-based molecular docking offers a rational in-silico strategy
Materials and Methods
Identification of target genes of both Disease and Phytochemicals
The Disease target genes of Dengue virus are identified and taken from the Mala cards data base website. The plant phytochemical compounds present in Carica papaya, Moringa Oleifera and Tinospora Cordifolia are identified from the website IMPPAT and they further scrutinized based on the Drug likeliness and ADME analysis. The Gene targets of these phytochemical compounds are identified and taken from the Swiss Target predictor website.
Identification of common genes
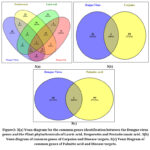
Using the VENNY 2.0 website, the disease genes from the mala cards and the phytochemical target genes from the Swiss target predictor are used for the Venn analysis to determine the shared target genes.
Identification of proteins and Ligands
The Protein Data Bank website (PDB) is used to identify the proteins associated with the common target genes that were retrieved from Venny. Proteins are chosen for further processing based on their characteristics, such as Organism, Mutations, and Resolution, among others. For molecular docking research, proteins found in humans with resolutions greater than 2.0 and no mutations are often recommended. The phytochemical compounds used in docking research are called ligands, and they may be found on the Pubchem website.
Molecular docking
The molecular docking of these identified proteins are done using the PyRx-0.8 software and the results of binding affinity are obtained from it. The Visualization of the docking study is done using the Biovia discovery studio 2025 software. And at sometimes CB-Dock an online blind docking tool which is used to identify the cur pockets in proteins which are suitable for Flex and Rigid docking.57,58,59 It is important to emphasize that molecular docking provides predictive insights into binding affinity and interaction patterns, but does not confirm biological efficacy.
Results
Dengue virus Genes
These Dengue virus genes are identified from the Mala card data base website from the Gene cards.
Plant Phytochemicals
These Phytochemicals are obtained from the IMPPAT website and later Scrutinized based on drug likeliness score and ADME analysis data.
Table 1: Selected Plant phytochemicals of Carica papaya, Moringa oleifera and Tinospora cordifolia.
| Carica papaya | ||||||||||||||||
| S.no | Name of phytochemical | IMPPAT identifier (IMPHY no) | Drug Likeness | ADME properties | ||||||||||||
| Lipins rule | Ghose rule | Drug like score (%) | GI Absorption | BBB crossing | Bio-availability | |||||||||||
| 1 | Carpaine | 003831 | Pass (1) | Pass (1) | 0.44 | High | No | 0.55 | ||||||||
| 2 | Palmitic acid | 007327 | Pass (1) | Pass (0) | 0.41 | High | Yes | 0.85 | ||||||||
| 3 | Lauric acid | 003016 | Pass (0) | Pass (0) | 0.54 | High | Yes | 0.85 | ||||||||
| 4 | Myristic acid | 000060 | Pass (0) | Pass (0) | 0.29 | High | Yes | 0.85 | ||||||||
| Moringa oleifera | ||||||||||||||||
| S.no | Name of phytochemical | IMPPAT identifier (IMPHY no) | Drug Likeness | ADME properties | ||||||||||||
| Lipins rule | Ghose rule | Drug like score (%) | GI Absorption | BBB crossing | Bio-availability | |||||||||||
| 1 | Palmitic acid | 007327 | Pass (1) | Pass (0) | 0.41 | High | Yes | 0.85 | ||||||||
| 2 | Hesperetin | 006750 | Pass (0) | Pass (0) | 0.79 | High | No | 0.55 | ||||||||
| 3 | Apigenin | 004661 | Pass (0) | Pass (0) | 0.63 | High | No | 0.55 | ||||||||
| 4 | Niazirinin | 001105 | Pass (0) | Pass (0) | 0.78 | High | No | 0.55 | ||||||||
| Tinospora Cordifolia | ||||||||||||||||
| S.no | Name of phytochemical | IMPPAT identifier (IMPHY no) | Drug Likeness | ADME properties | ||||||||||||
| Lipins rule | Ghose rule | Drug like score (%) | GI Absorption | BBB crossing | Bio-availability | |||||||||||
| 1 | Palmitic acid | 007327 | Pass (0) | Pass (0) | 0.41 | High | Yes | 0.85 | ||||||||
| 2 | Pentadecanoic acid | 002667 | Pass (1) | Pass (0) | 0.45 | High | Yes | 0.85 | ||||||||
| 3 | Myristic acid | 000060 | Pass (0) | Pass (0) | 0.29 | High | Yes | 0.85 | ||||||||
Identification of common genes
The common genes of the disease targets and the selected phytochemical compounds of Carica papaya, Moringa oleifera and Tinospora cordifolia is identified by drawing VENN diagrams for the disease genes and the plant phytochemical targets. The Venn diagram was drawn in the website VENNY 2.0 between disease genes and the selected plant phytochemical target compounds. Venn diagram drawn between disease genes and Carpaine, Palmitic acid, Lauric acid, Hesperetin and Pentadecanoic acid.
Table 2: Identified common genes of the plant phytochemicals.
| S.no | Medicinal Plant | Phytochemical | Number of Identified genes | Name of the genes | Gene code |
| 1 | Carica papaya | Carpaine | 1 | Plasminogen | PLG |
| Palmitic acid | 1 | Plasminogen | PLG | ||
| Lauric acid | 1 | Plasminogen | PLG | ||
| Myristic acid | 1 | Plasminogen | PLG | ||
| 2 | Moringa Oleifera | Palmitic acid | 1 | Plasminogen | PLG |
| Hesperetin | 1 | Plasminogen | PLG | ||
| Apigenin | 1 | Plasminogen | PLG | ||
| Niazirinin | 1
|
Heat Shock Protein Family A (Hsp70)
Member 5 |
HSPA5 | ||
|
3 |
Tinospora cordifolia | Palmitic acid | 1 | Plasminogen | PLG |
| Pentadecanoic acid | 1 | Plasminogen | PLG | ||
| Myristic acid | 1 | Plasminogen | PLG |
Protein Selection
Plasminogen (PLG) as the common gene target among all the three plants and Protein are identified according to their characters and 8UQ6 with (pdb_00008uq6) id The Human plasminogen protein which bind to the streptococcal surface enolase is selected for further molecular docking studies. This protein has a Resolution of 3.60A with no mutations in Humans. Without mutations and a high resolution value it is easy to use for the studies without any disturbances. The protein is cleaned by removing the Hetero and Water atoms present if any and after used for molecular docking process.
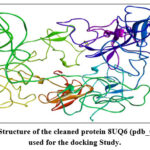 |
Figure 4: Structure of the cleaned protein 8UQ6 (pdb_00008uq6) used for the docking Study.
|
Molecular Docking
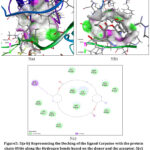
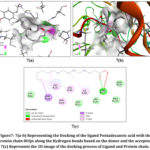
The Molecular docking process of the obtained Phytochemical (ligand) compounds Carpaine, Lauric acid, Palmitic acid, Hesperetin and Pentadecanoic acid as they have a similar gene targets PLG and the obtained protein 8UQ6 is commonly used for the 3 ligands. The molecular docking done by using the software’s PyRx and the Biovia Discovery studio 2025. The PyRx software provides the docking data and the binding affinity of the ligand and protein, where the discovery studio is used to visualize the actual site of docking of the Ligand inside the protein chain. The protein 8UQ6 got bind with the 5 ligand compounds. It is important to emphasize that molecular docking provides predictive insights into binding affinity and interaction patterns, but does not confirm biological efficacy.
Table 3: Binding Affinity of the compounds
| S,no | Phytochemical (Ligand) | Pubchem Id | Binding compounds | Binding Affinity
(kcal/mol) |
| 1 | Carpaine | 442630 | 8UQ6_442630_uff_E=332.50 | -9.2 |
| 2 | Hesperetin | 72281 | 8UQ6_72281_uff_E=283.48 | -8.3 |
| 3 | Pentadecanoic acid | 13849 | 8UQ6_13849_uff_E=53.38 | -5.5 |
| 4 | Palmitic acid | 985 | 8UQ6_985_uff_E=57.46 | -5.4 |
| 5 | Lauric acid | 3893 | 8UQ6_3893_uff_E=43.33 | -5.2 |
Visualization Analysis
The Visualization analysis for the 5 different ligands with the 1 targeted protein is done by using the Biovia Discovery studio 2025 software.
Discussion
The present in-silico investigation explored the interaction potential of phytochemicals from Carica papaya, Moringa oleifera, and Tinospora cordifolia with host factors relevant to dengue pathogenesis. Network-based target analysis identified plasminogen (PLG) as a shared molecular target associated with dengue-related pathways and the selected phytochemicals, highlighting its potential relevance in host-directed therapeutic strategies.
Plasminogen is a key regulator of fibrinolysis, inflammation, and vascular integrity, processes that are markedly dysregulated during dengue infection and contribute to thrombocytopenia and plasma leakage. Modulation of PLG activity may therefore influence platelet stability and endothelial function, both of which are central to dengue disease severity. Based on structural quality and biological relevance, human plasminogen (PDB ID: 8UQ6) was selected for molecular docking to evaluate ligand–protein interactions.
Docking analysis demonstrated that carpaine, an alkaloid from Carica papaya, exhibited the highest binding affinity toward PLG, indicating a stable and energetically favorable interaction. This observation provides a plausible molecular basis for previous clinical and experimental reports describing the platelet-supportive effects of C. papaya in dengue-associated thrombocytopenia. Hesperetin from Moringa oleifera also showed a strong binding interaction, suggesting that flavonoid-mediated modulation of host inflammatory or fibrinolytic pathways may contribute to its reported therapeutic potential. Although Tinospora cordifolia is widely recognized for its immunomodulatory and antiviral properties, its phytochemicals displayed comparatively weaker binding affinities toward PLG in the present model, indicating that its effects may be mediated through alternative molecular targets or pathways not captured in this analysis.
Despite these promising findings, the study has inherent limitations. Molecular docking provides predictive insights into ligand–protein interactions but does not confirm biological activity or therapeutic efficacy. The identification of a single common host target does not fully capture the complexity of dengue pathogenesis, which involves multiple viral and host-mediated mechanisms. Furthermore, docking studies do not account for pharmacokinetic behavior, bioavailability, metabolism, or cellular context. Consequently, the results should be interpreted as preliminary evidence supporting further in vitro, in vivo, and clinical investigations to validate the role of these phytochemicals in dengue management.
Conclusion
The present in-silico study investigated phytochemicals from Carica papaya, Moringa oleifera, and Tinospora cordifolia for their potential interactions with a dengue-associated host target. Molecular docking analysis identified plasminogen as a common target, with carpaine from Carica papaya exhibiting the highest binding affinity, followed by hesperetin from Moringa oleifera. These interactions suggest a favorable molecular basis for the reported platelet-supportive and anti-inflammatory properties of these plants in dengue-associated complications.
While phytochemicals from Tinospora cordifolia showed comparatively weaker interactions with plasminogen in this model, their established immunomodulatory effects may involve alternative molecular pathways. Overall, the findings highlight Carica papaya and Moringa oleifera as promising sources of host-directed compounds relevant to dengue pathophysiology and provide a rational computational framework to support further experimental exploration.
Future studies should prioritize experimental validation of the identified phytochemicals using in vitro assays, including antiviral activity and platelet function models, followed by evaluation in suitable in vivo dengue models. Detailed pharmacokinetic, toxicological, and bioavailability studies will be essential to assess the translational potential of carpaine and hesperetin. Additionally, exploring multi-target host pathways and synergistic phytochemical combinations may provide deeper insight into their therapeutic relevance. Such investigations could facilitate the development of evidence-based, phytochemical-derived adjunct strategies for dengue management.
The promising in silico interactions observed for carpaine and hesperetin warrant further experimental validation through in vitro and in vivo models to confirm their biological relevance and therapeutic potential in dengue-associated complications.
Acknowledgement
The authors sincerely thank Mrs. V. Jhansi Lakshmi for guidance and Vignan Institute of Pharmaceutical Technology for providing necessary facilities.
Funding Sources
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflict of Interest
The authors do not have any conflict of interest.
Data Availability Statement
This statement does not apply to this article.
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Clinical Trial Registration
This research does not involve any clinical trials.
Permission to reproduce material from other sources
Not Applicable
Author Contributions
Jhansi Lakshmi Varanasi: Guidance and Supervision.
Naga Bharathi Marni: Methodology and Conceptualization.
Sai Sudheer Thatavarthi: Data Collection, Analysis, Molecular Docking, Visualization, Writing – Original Draft.
Mounika Matcha: Review & Editing.
References
- World Health Organization; Special Programme for Research and Training in Tropical Diseases. Dengue case classification. In: Dengue: Guidelines for Diagnosis, Treatment, Prevention and Control. New ed. World Health Organization; 2009:10–12.
- Quirino-Teixeira AC, Rozini SV, Barbosa-Lima G, et al. Inflammatory signaling in dengue-infected platelets requires translation and secretion of nonstructural protein 1. Blood Adv. 2020;4(9):2018–2031. doi:10.1182/bloodadvances.2019001169.
CrossRef - Hottz ED, Lopes JF, Freitas C, et al. Platelets mediate increased endothelium permeability in dengue through NLRP3 inflammasome activation. 2013;122(20):3405–3414. doi:10.1182/blood-2013-05-504449
CrossRef - Hottz ED, Medeiros-de-Moraes IM, Vieira-de-Abreu A, et al. Platelet activation and apoptosis modulate monocyte inflammatory responses in dengue. J Immunol. 2014;193(4):1864–1872. doi:10.4049/jimmunol.1400091
CrossRef - Barbosa-Lima G, Hottz ED, De Assis EF, et al. Dengue virus–activated platelets modulate monocyte immunometabolic response through lipid droplet biogenesis and cytokine signaling. J Leukoc Biol. 2020;108(4):1293–1306. doi:10.1002/JLB.4MA0620-658R
CrossRef - Bozza FA, Cruz OG, Zagne SMO, et al. Multiplex cytokine profile from dengue patients: MIP-1β and IFN-γ as predictive factors for severity. BMC Infect Dis. 2008;8:86. doi:10.1186/1471-2334-8-86
CrossRef - Zhao L, Huang X, Hong W, et al. Slow resolution of inflammation in severe adult dengue patients. BMC Infect Dis. 2016;16:291. doi:10.1186/s12879-016-1596-x
CrossRef - Pan P, Zhang Q, Liu W, et al. Dengue virus M protein promotes NLRP3 inflammasome activation to induce vascular leakage in mice. J Virol. 2019;93(21):e00996-19. doi:10.1128/JVI.00996-19
CrossRef - Chen HR, Chuang YC, Lin YS, et al. Dengue virus nonstructural protein 1 induces vascular leakage through macrophage migration inhibitory factor and autophagy. PLoS Negl Trop Dis. 2016;10(7):e0004828. doi:10.1371/journal.pntd.0004828
CrossRef - Tsai JJ, Jen YH, Chang JS, et al. Frequency alterations in key innate immune cell components in peripheral blood of dengue patients detected by FACS analysis. J Innate Immun. 2011;3(5):530–540. doi:10.1159/000322904
CrossRef - Sung PS, Huang TF, Hsieh SL. Extracellular vesicles from CLEC2-activated platelets enhance dengue virus–induced lethality via CLEC5A/TLR2. Nat Commun. 2019;10:2402. doi:10.1038/s41467-019-10360-4
CrossRef - Brasier AR, Ju H, Garcia J, et al. A three-component biomarker panel for prediction of dengue hemorrhagic fever. Am J Trop Med Hyg. 2012;86(2):341–348. doi:10.4269/ajtmh.2012.11-0469
CrossRef - Chaloemwong J, Tantiworawit A, Rattanathammethee T, et al. Useful clinical features and hematological parameters for the diagnosis of dengue infection in patients with acute febrile illness. BMC Hematol. 2018;18:20. doi:10.1186/s12878-018-0116-1
CrossRef - Teixeira-Ferreira A, Perales J, et al. Platelet proteome reveals novel pathways of platelet activation and platelet-mediated immunoregulation in dengue. PLoS Pathog. 2017;13(5):e1006385. doi:10.1371/journal.ppat.1006385
CrossRef - Ojha A, Nandi D, Batra H, et al. Platelet activation determines the severity of thrombocytopenia in dengue infection. Sci Rep. 2017;7:41697. doi:10.1038/srep41697
CrossRef - Wilder-Smith A, Ooi EE, Horstick O, Wills B. Dengue. 2019;393(10169):350–363. doi:10.1016/S0140-6736(18)32560-1
CrossRef - Suwanmanee S, Luplertlop N. Immunopathogenesis of dengue virus-induced redundant cell death: apoptosis and pyroptosis. Viral Immunol. 2017;30(1):13–19. doi:10.1089/vim.2016.0092
CrossRef - Hidari KIPJ, Suzuki T. Dengue virus receptor. Trop Med Health. 2011;39(suppl 4):S37–S43. doi:10.2149/tmh.2011-S03
CrossRef - Kar M, Singla M, Chandele A, et al. Dengue virus entry and replication does not lead to productive infection in platelets. Open Forum Infect Dis. 2017;4(2):ofx051. doi:10.1093/ofid/ofx051
CrossRef - Simon AY, Sutherland MR, Pryzdial ELG. Dengue virus binding and replication by platelets. 2015;126(3):378–385. doi:10.1182/blood-2014-09-598029
CrossRef - Hottz ED, Oliveira MF, Nunes PCG, et al. Dengue induces platelet activation, mitochondrial dysfunction, and cell death through mechanisms involving DC-SIGN and caspases. J Thromb Haemost. 2013;11(5):951–962. doi:10.1111/jth.12178
CrossRef - Urcuqui-Inchima S, Patiño C, Torres S, et al. Recent developments in understanding dengue virus replication. Adv Virus Res. 2010;77:1–39. doi:10.1016/S0065-3527(10)77001-1
CrossRef - Bartenschlager R, Miller S. Molecular aspects of dengue virus replication. Future Microbiol. 2008;3(2):155–165. doi:10.2217/17460913.3.2.155
CrossRef - Wills B, Tran VN, Van NTH, et al. Hemostatic changes in Vietnamese children with mild dengue correlate with the severity of vascular leakage rather than bleeding. Am J Trop Med Hyg. 2009;81(4):638–644. doi:10.4269/ajtmh.2009.08-0066
CrossRef - Srichaikul T, Nimmannitya S. Haematology in dengue and dengue haemorrhagic fever. Best Pract Res Clin Haematol. 2000;13(2):261–276. doi:10.1053/beha.2000.0073
CrossRef - La Russa VF, Innis BL. Mechanisms of dengue virus-induced bone marrow suppression. Baillieres Clin Haematol. 1995;8(1):249–270. doi:10.1016/S0950-3536(05)80294-4
CrossRef - Bierman HR, Nelson ER. Hematodepressive virus diseases of Thailand. Ann Intern Med. 1965;62(5):867–884.
CrossRef - Sridharan A, Chen Q, Tang KF, et al. Inhibition of megakaryocyte development in bone marrow underlies dengue virus-induced thrombocytopenia in humanized mice. J Virol. 2013;87(21):11648–11658. doi:10.1128/JVI.01718-13
CrossRef - Clark KB, Noisakran S, Onlamoon N, et al. Multiploid CD61+ cells are the predominant lineage infected during acute dengue virus infection in bone marrow. PLoS One. 2012;7(12):e52902. doi:10.1371/journal.pone.0052902
CrossRef - Vogt MB, Lahon A, Arya RP, et al. Dengue viruses infect human megakaryocytes with probable clinical consequences. PLoS Negl Trop Dis. 2019;13(11):e0007837. doi:10.1371/journal.pntd.0007837
CrossRef - Basu A, Jain P, Gangodkar SV, et al. Dengue-2 virus inhibits in vitro megakaryocytic colony formation and induces apoptosis. FEMS Immunol Med Microbiol. 2008;53(1):46–51. doi:10.1111/j.1574-695X.2008.00402.x
CrossRef - Alonzo MTG, Lacuesta TLV, Dimaano EM, et al. Platelet apoptosis and apoptotic platelet clearance by macrophages in secondary dengue virus infections. J Infect Dis. 2012;205(8):1321–1329. doi:10.1093/infdis/jis180
CrossRef - Onlamoon N, Noisakran S, Hsiao HM, et al. Dengue virus-induced hemorrhage in a nonhuman primate model. 2010;115(9):1823–1834. doi:10.1182/blood-2009-09-243196
CrossRef - Krishnamurti C, Cutting MA, Peat RA, Rothwell SW. Platelet adhesion to dengue-2 virus-infected endothelial cells. Am J Trop Med Hyg. 2002;66(4):435–441. doi:10.4269/ajtmh.2002.66.435
CrossRef - Póvoa TF, Alves AMB, Oliveira CAB, et al. Pathology of severe dengue in multiple organs of human fatal cases. PLoS One. 2014;9(4):e83386. doi:10.1371/journal.pone.0083386
CrossRef - Riswari SF, Tunjungputri RN, Kullaya V, et al. Desialylation of platelets induced by von Willebrand factor is a novel mechanism of platelet clearance in dengue. PLoS Pathog. 2019;15(3):e1007500. doi:10.1371/journal.ppat.1007500
CrossRef - Sørensen AL, Rumjantseva V, Nayeb-Hashemi S, et al. Role of sialic acid for platelet lifespan. 2009;114(8):1645–1654. doi:10.1182/blood-2009-01-199414
CrossRef - Saito M, Oishi K, Inoue S, et al. Increased platelet-associated immunoglobulins correlate with thrombocytopenia and disease severity in dengue. Clin Exp Immunol. 2004;138(2):299–303. doi:10.1111/j.1365-2249.2004.02608.x
CrossRef - Honda S, Saito M, Dimaano EM, et al. Increased phagocytosis of platelets in secondary dengue virus infection. Am J Trop Med Hyg. 2009;80(5):841–845. doi:10.4269/ajtmh.2009.80.841
CrossRef - Sun DS, King CC, Huang HS, et al. Antiplatelet autoantibodies elicited by dengue virus NS1 cause thrombocytopenia and mortality in mice. J Thromb Haemost. 2007;5(11):2291–2299. doi:10.1111/j.1538-7836.2007.02700.x
CrossRef - Falconar AK. Dengue virus NS1 generates antibodies to human clotting and endothelial proteins. Arch Virol. 1997;142(5):897–916.
CrossRef - Avirutnan P, Punyadee N, Noisakran S, et al. Vascular leakage in severe dengue: role of NS1 and complement. J Infect Dis. 2019;220(5):1078–1088. doi:10.1093/infdis/jiz351
CrossRef - Nascimento EJM, Silva AM, Cordeiro MT, et al. Alternative complement pathway deregulation correlates with dengue severity. PLoS One. 2009;4(8):e6782. doi:10.1371/journal.pone.0006782
CrossRef - Nascimento EJM, Hottz ED, Garcia-Bates TM, et al. Emerging concepts in dengue pathogenesis. Crit Rev Immunol. 2014;34(3):227–240. doi:10.1615/CritRevImmunol.2014010083
CrossRef - Acosta EG, Kumar A, Bartenschlager R. Revisiting dengue virus–host cell interaction. Adv Virus Res. 2014;88:1–109. doi:10.1016/B978-0-12-800098-4.00001-5
CrossRef - Roy SK, Bhattacharjee S. Dengue virus: epidemiology, biology, and disease aetiology. Can J Microbiol. 2021;67:687–702. doi:10.1139/cjm-2020-0572
CrossRef - Ehrenkranz NJ, Ventura AK, Cuadrado RR, et al. Pandemic dengue in Caribbean countries and the southern United States. N Engl J Med. 1971;285:1460–1469.
CrossRef - Reis SRN, Sampaio ALF, Henriques MG, et al. In vitro model for dengue virus infection. Mem Inst Oswaldo Cruz. 2007;102:983–990. doi:10.1590/S0074-02762007000800013
CrossRef - Hotta S. Experimental studies on dengue. I. Isolation and modification of the virus. J Infect Dis. 1952;90:1–9.
CrossRef - Mustafa MS, Rasotgi V, Jain S, Gupta V. Discovery of fifth serotype of dengue virus (DENV-5). Med J Armed Forces India. 2015;71:67–70. doi:10.1016/j.mjafi.2014.12.011
CrossRef - Normile D. Surprising new dengue virus throws a spanner in disease control efforts. 2013;342:415. doi:10.1126/science.342.6157.415
CrossRef - World Health Organization. Dengue and severe dengue. Updated September 16, 2025.
- Srisawat N, Gubler DJ, Pangestu T, et al. Proceedings of the 6th Asia Dengue Summit, June 2023. PLoS Negl Trop Dis. 2024;18:e0012060. doi:10.1371/journal.pntd.0012060
CrossRef - Tsheten T, Gray DJ, Clements ACA, Wangdi K. Epidemiology and challenges of dengue surveillance in the WHO South-East Asia Region. Trans R Soc Trop Med Hyg. 2021;115:583–599. doi:10.1093/trstmh/traa158
CrossRef - Bhatia R, Dash A, Sunyoto T. Changing epidemiology of dengue in South-East Asia. WHO South East Asia J Public Health. 2013;2:23–27.
CrossRef - Singhal T, Kothari V. Clinical and laboratory profile of fatal dengue cases in Mumbai, India. Am J Trop Med Hyg. 2020;103:1223–1227. doi:10.4269/ajtmh.19-0425
CrossRef - Lim SP, Noble CG, Seh CC, et al. Potent inhibition of dengue virus replication by adenosine nucleoside analogues targeting the viral polymerase. Antiviral Res. 2013;97(3):204–214. doi:10.1016/j.antiviral.2012.12.010
CrossRef - Pattnaik A, Sahoo BR, Pattnaik AK. Molecular docking and simulation studies of dengue virus NS3 protease inhibitors. J Biomol Struct Dyn. 2020;38(12):3513–3525. doi:10.1080/07391102.2019.1660306
- Sharma M, Chauhan N, Kumar R, et al. In silico identification of potential inhibitors of dengue virus NS5 RNA-dependent RNA polymerase. J Mol Model. 2019;25:241. doi:10.1007/s00894-019-4136-4
- Joshi T, Joshi T, Sharma P, et al. In silico screening of natural compounds against dengue virus NS2B–NS3 protease. J Biomol Struct Dyn. 2021;39(10):3545–3557. doi:10.1080/07391102.2020.1758786
- Kumar A, Dwivedi VD, Bharadwaj S, et al. Structure-based virtual screening of phytochemicals against dengue virus targets. Virus Res. 2020;276:197839. doi:10.1016/j.virusres.2019.197839
CrossRef - Subramanian G, Suresh Kumar M, Gayathri P. Molecular docking studies of bioactive compounds from Carica papaya against dengue virus proteins. 2018;14(9):453–459. doi:10.6026/97320630014453
- Singh A, Gautam A, Singh S, et al. In silico evaluation of phytoconstituents from Tinospora cordifolia as antiviral agents targeting dengue virus. J Appl Pharm Sci. 2020;10(8):89–97. doi:10.7324/JAPS.2020.10812
CrossRef - Rathi B, Kaur J, Kumar R. Molecular docking analysis of phytochemicals from Moringa oleifera as potential dengue virus inhibitors. Nat Prod Res. 2021;35(20):3605–3610. doi:10.1080/14786419.2019.1701221
Accepted on: 14-02-2026
Second Review by: Dr. Ichrak Jaouadi
Final Approval by: Dr. Wagih Ghannam