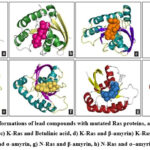
Virtual Screening for Lead Molecules in Aegle Marmelos (L.) Correa. on Mutated Ras Proteins
1Department of Botany, Assumption College (Autonomous), Changanassery, Kerala, India.
2Department of Botany, CMS College Kottayam (Autonomous), Kerala, India.
Corresponding Author Email: rogimon@cmscollege.ac.in
DOI : http://dx.doi.org/10.13005/bbra/3498
ABSTRACT:India's rich biodiversity is complemented by its vast knowledge of medicinal plants and traditional practices, with plant-based compounds having been harnessed as therapeutic agents for centuries. Cancer, a leading cause of global mortality, is fundamentally driven by DNA abnormalities that disrupt the function of key metabolic proteins. Ras proteins show a vital role in cell signalling, regulating cell proliferation and apoptosis, but mutations in these proteins may results in uncontrolled cell growth. This study investigated the anticancer potential of phytochemicals against mutated H-RAS, N-RAS, and K-RAS (RAS proteins) using molecular docking analysis. Aegle marmelos, rich in over a hundred phytochemicals with reported anticancer properties, was the source of 89 molecules screened as ligands. Seventy-eight molecules exhibited strong binding affinity (∆Gbind ≤ -5 kcal/mol) for H-Ras, 74 for K-Ras, and 77 for N-Ras, with α-amyrin and lupeol emerging as top leads for H-Ras, betulinic acid, β-amyrin, and lupeol for K-Ras, and βand α-amyrins for N-Ras. α-Amyrin emerged as a promising lead molecule against mutated Ras proteins, showing strong potential against H-Ras and N-Ras, and comparable efficacy to top leads against K-Ras. To overcome the challenges observed in drug likeness prediction connected with alpha amyrin, nano emulsion formulations strategies can improve its bioavailability and efficacy. Further validation of the lead molecules' anticancer efficacy requires comprehensive preclinical and clinical studies to confirm their biological activity.
KEYWORDS:Aegle marmelos; Anticancer; Auto Dock; Molecular docking; Phytochemicals; Ras proteins
Introduction
Cancer is a severe metabolic disorder recognized by abnormal proliferation of cells that spread all over the body and affect the normal functions of the body systems including nervous, circulatory, endocrine, and digestive systems.1 The World Health Organization data proclaimed that cancer is one of the major mortality which killed around ten million people in the year, 2020 (https://www.who.int/news-room/fact-sheets/detail/cancer) and the number of cancer patients will be increased to 21 million by 2030.2 Cancer is caused by several factors such as gene mutations, disruption of the immune system, and the effect of various carcinogens.1 The consequences of this diseased condition is driven by both genetic and epigenetic changes is the disruption of signaling pathways of cells that controls mitosis, growth, and death.3 Consequently, this interruption leads to abnormal proliferation of cells as a result of the inhibition of the normal events of the cell cycle such as cell cycle arrest and programmed cell death.4
Ras genes encode for Ras proteins or GTPases, function as molecular switches that regulate cellular signalling pathways controlling cell differentiation, proliferation, and programmed cell death. Ras proteins become oncogenes when mutated and have a crucial role in human cancer which has inspired multiple attempts to find RAS inhibitors.5Mutant RAS genes fuel some of the deadliest cancers, including pancreatic, colorectal, and lung cancers, by sending non-stop signals that drive tumor growth and cell proliferation.6 The oncogenic mutations on the H-Ras, K-Ras, or N-Ras genes observed in most human tumors alter the normal functioning of those signaling pathways there by resulting in the development of tumours.7 Ras is a protooncogene that becomes an oncogene because of a single base change that results in an alteration in a single amino acid in the encoded protein. The major Ras mutation is the change of the amino acid at position 12 in H-Ras and K-Ras and at position 61 in and N-Ras.8,9 Only these few, very specific mutations can make a Ras proto-oncogene to an oncogene. These few changes in the amino acid residues are directly involved in the binding and splitting of the GTP.7 Ras mutations are found in around 19% of cancer patients and are often linked to unfavorable prognoses. Despite extensive research, K-Ras has remained a difficult target to tackle, largely due to its molecular structure that resists small-molecule binding. 10 Despite the high similarity between the protein products of the three Ras genes, K-Ras mutations are significantly more prevalent in cancer and RASopathies, and the analysis reveals a correlation between RAS protein expression levels (KRAS > NRAS > HRAS) and mutation frequencies.11
Cancer treatment comprises surgery, chemotherapy, immunotherapy, radiotherapy, photodynamic therapy, cancer vaccinations, and stem cell transformation as individually or in combination are effective but results in severe side effects such as limited bioavailability, toxicity, fast clearance, non-specificity, and restriction in metastasis. Current anticancer drugs often affect rapidly dividing normal cells, such as those in bone marrow and hair follicles, leading to significant side effects. The extensive investigation for the alternative treatment proposes the use of phytochemicals and their derived analogs will be the most possible option for the better and nontoxic cancer treatment.12
Aegle marmelos(L.) Correa., Bael, a member of the Rutaceae family, is widely recognized for its medicinal properties and has been utilized in traditional medicine systems.13 Previous investigations on the ethnopharmacological properties of the plant showed that the phytochemicals in the plant possess radio-protective,anti-neoplastic, chemoprotective, and chemopreventive effects, effective in cancer prevention and treatment.14 Present scenario, computer-aided drug discovery approaches significantly contribute to drug discovery experiments and their interpretation to expedite the entire process. The current study was designed to validate the anticancerous activity and to identify the lead phytochemicals against mutated Ras proteins in Aegle marmelos (L.) Correa, by in silico method.
Materials and Methods
Target molecule preparation
The three Ras proteins viz., H-Ras (1P2U), K-Ras (5UFE) and N-Ras (5UHV) were used for docking simulations. The three-dimensional structures of these proteins were sourced from RCSB Protein Data Bank. The oncogenic mutations reported by Pierotti et al., (2003)8 were done in the three dimensionalmolecular structures of these Ras proteins using Swiss PDB Viewer 4.0.115 (Fig. 1). The glycine residue at the 12 position of 1P2U and 5UFE was mutated to valine and aspartic acid respectively. In 5UHV, the glutamine residue at the 61 position was mutated to lysine. The mutated 1P2U, 5UFE and 5UHV were used as the receptor molecules for docking and the mutated residues were taken as active residues for the detection of active sites.
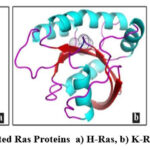 |
Figure 1: Mutated Ras Proteins a) H-Ras, b) K-Ras, c) N-Ras
|
Preparation of ligand molecules
The literature survey revealed that more than 100 phytochemicals have been so far reported from A. marmelos. Out of these, 89 molecules with molecular weights under 700 g/mol were selected for docking studies. The Canonical Simplified Molecular Input Line Entry Systems (Canonical SMILES) of the ligands were obtained from PubChem and 3D structures were created with CORINA.16The ligands chosen for docking are shown in Table 1.
Docking
Docking studies were performed using AutoDock 4.2, with the selected phytochemicals docked into the Ras protein’s binding site according to Morris et al., (2009).17 This tool leverages Monte Carlo Simulated Annealing and Lamarckian genetic algorithm for possible orientations of ligands at the Ras protein binding site. The active site residues of the proteins 1P2U, 5UFE and 5UHV were VAL12, ASP12 and LYS61 respectively. Grid dimensions of 60×60×60 points with 0.375Å spacing were specified for each protein. The active site was centered in the grid, with XYZ coordinates defined as: 1P2U – 24.721Å, 7.378Å, 38.718Å, 5UFE-10.117Å, 6.296Å, 3.87Å and 5UHV – 5.423Å, 24.44Å, 11.983Å respectively. All remaining parameters were maintained at their default values. After docking, the binding affinity and possible orientations of the ligand-protein complexes were analyzed and ranked by their binding energies using cluster analysis. Molecules with a binding energy ≤ -5 kcal/mol were considered potential hit compounds.16 The docked structures were visualized and images were created using PyMol visualization software.18
Table 1: The list of prioritized phytochemicals in Aegle marmelos
| Sl.
No. |
Phytochemicals | MolecularWeight
(g/mol) |
Molecular
Formula |
| 1 | 3,5-octadiene-2-one | 124.18 | C8H12O |
| 2 | 3-Phenylacrylamide | 147.17 | C9H9NO |
| 3 | Acetoin | 88.11 | C4H8O2 |
| 4 | Alloimperatorin | 270.28 | C16H14O4 |
| 5 | Ascorbicacid | 176.12 | C6H8O6 |
| 6 | Aurapten | 298.4 | C19H22O3 |
| 7 | Betulinicacid | 456.7 | C30H48O3 |
| 8 | Carvone | 150.22 | C10H14O |
| 9 | Carvylacetate | 194.27 | C12H18O2 |
| 10 | Caryophylleneoxide | 220.35 | C15H24O |
| 11 | Cineole | 154.25 | C10H18O |
| 12 | Cis-carveol | 152.23 | C10H16O |
| 13 | Citral | 152.23 | C10H16O |
| 14 | Citronellal | 154.25 | C10H18O |
| 15 | Cuminaldehyde | 148.2 | C10H12O |
| 16 | Decursinol | 246.26 | C14H14O |
| 17 | Dictamine | 199.2 | C12H9NO2 |
| 18 | Elemol | 222.37 | C15H26O |
| 19 | Emodin | 270.24 | C15H10O5 |
| 20 | Eugenol | 164.2 | C10H12O2 |
| 21 | Fagarine | 229.23 | C13H11NO3 |
| 22 | Farnesol | 222.37 | C15H26O |
| 23 | Flavone | 222.24 | C15H10O2 |
| 24 | Furoquinoline | 169.18 | C11H7NO |
| 25 | Geraniol | 154.25 | C10H18O |
| 26 | Haplopine | 245.23 | C13H11NO4 |
| 27 | Limonene | 136.23 | C10H16 |
| 28 | Linalool | 154.25 | C10H18O |
| 29 | Lupeol | 426.7 | C30H50O |
| 30 | Luvangetin | 258.27 | C15H14O4 |
| 31 | Marmeline | 335.44 | C22H25NO2 |
| 32 | Marmelosin | 270.28 | C16H14O4 |
| 33 | Marmesin | 246.26 | C14H14O4 |
| 34 | Marmesinin | 408.4 | C20H24O9 |
| 35 | Marmin | 332.4 | C19H24O5 |
| 36 | Methylcinnamate | 162.18 | C10H10O2 |
| 37 | Methylether | 46.07 | C2H6O |
| 38 | Methylpalmitate | 270.5 | C17H34O2 |
| 39 | Methylperillate | 180.24 | C11H16O2 |
| 40 | Myristicacid | 228.37 | C14H28O2 |
| 41 | Myrtenol | 152.23 | C10H16O |
| 42 | o-isopentenyl-halfordinol | 306.4 | C19H18N2O2 |
| 43 | p-cymene | 134.22 | C10H14 |
| 44 | Phytol | 296.5 | C20H40O |
| 45 | Piperitone | 152.23 | C10H16O |
| 46 | Plumbagin | 188.18 | C11H8O3 |
| 47 | Psoralen | 186.16 | C11H6O3 |
| 48 | Pulegone | 152.23 | C10H16O |
| 49 | Quercetin | 302.23 | C15H26O |
| 50 | Rutaretin | 262.26 | C14H14O5 |
| 51 | Rutin | 610.5 | C27H30O16 |
| 52 | Sabinene | 136.23 | C10H16 |
| 53 | Sabinol | 152.23 | C10H16O |
| 54 | Scoparone | 206.19 | C11H10O4 |
| 55 | Scopoletin | 192.17 | C10H8O4 |
| 56 | Shahidine | 279.3 | C18H17NO2 |
| 57 | Skimmianine | 259.26 | C14H13NO4 |
| 58 | Skimmin | 324.28 | C15H16O8 |
| 59 | Tembamide | 271.31 | C16H17NO3 |
| 60 | Trans-carveol | 152.23 | C10H16O |
| 61 | Umbelliferone | 162.14 | C9H6O3 |
| 62 | Valencicacid | 206.24 | C12H14O3 |
| 63 | Vanillin | 152.15 | C8H8O3 |
| 64 | Verbenone | 150.22 | C10H14O |
| 65 | Xanthotoxin | 216.19 | C12H8O4 |
| 66 | α-amyrin | 426.7 | C30H50O |
| 67 | α-cedrene | 204.35 | C15H24 |
| 68 | α-copaene | 204.35 | C15H24 |
| 69 | α-cubebene | 204.35 | C15H24 |
| 70 | α-elemene | 204.35 | C15H24 |
| 71 | α-humulene | 220.35 | C15H24O |
| 72 | α-phellandrene | 136.23 | C10H16 |
| 73 | α-pinene | 136.23 | C10H16 |
| 74 | α-zingiberine | 204.35 | C15H24 |
| 75 | β-amyrin | 426.72 | C30H50O |
| 76 | β-elemene | 204.35 | C15H24 |
| 78 | β-eudesmol | 222.37 | C15H26O |
| 79 | β-funebrene | 204.35 | C15H24 |
| 80 | β-ionone | 192.3 | C13H20O |
| 81 | β-myrcene | 142.27 | C10H16 |
| 82 | β-ocimene | 136.23 | C10H16 |
| 83 | β-phellandrene | 136.23 | C10H16 |
| 84 | β-pinene | 136.23 | C10H16 |
| 85 | β-selinene | 204.35 | C15H24 |
| 86 | β-sitosterol | 414.7 | C29H50O |
| 87 | β-sitosterolβ-D-glucoside | 576.85 | C35H60O6 |
| 88 | γ-sitosterol | 432.7 | C29H52O2 |
| 89 | γ-terpinene | 136.23 | C10H16 |
Drug likeliness prediction
The drug-likeness properties of the lead molecules were assessed by submitting each molecule on Molinspiration property prediction (https://www.molinspiration.com) and SWISSADME (http://www.swissadme.ch/index.php) tools. The tool analyzes molecular properties in line with Lipinski’s rule of five, highlighting potential issues highlighted.20 Octanol/water partition coefficient (MiLogP) was computed to predict membrane permeability. It was determined by summing fragment contributions and applying correction factors, as per Molinspiration’s fragment-based method. The miLogP was derived by correlating calculated logP with experimental logP values for a set of drug-like molecules, which predicts oral bioavailability.21
Besides, the tool predicts other molecular properties (TPSA, MW, ON, OHNH, ROTB, and volume) were calculated based on the molecule’s three-dimensional conformation. Topological polar surface area (TPSA) is used to predict various aspects of drug absorption, including intestinal absorption, bioavailability, and penetration of the blood-brain barrier. TPSA is computed by summing the fragment-based contributions of polar fragments centered on oxygen and nitrogen atoms, as well as the surface areas of hydrogen-bonded oxygen and nitrogen atoms.The number of rotatable bonds (nrotb) is a topological descriptor that quantifies molecular flexibility and is a strong predictor of oral bioavailability. The volume of a molecule influences its transport characteristics, such as passage through the blood-brain barrier and intestinal absorption.22
Bioactivity Prediction
Bioactivity scores for each lead molecule were predicted based on properties such as GPCR ligand activity, ion channel modulation, kinase inhibition, and nuclear receptor binding using Molinspiration.22
Results
Docking studies revealed that 89 phytochemicals from A. marmelos exhibit inhibitory activity against three mutated Ras proteins. The docking scores calculated using AutoDock 4.2 indicated that there are several hit molecules on each Ras protein. Hit molecules with the lowest free energy of bindingon the three mutated Ras proteins were selected as potential lead molecules. The selected leads are listed in Table 2.
Table 2: Potential Ras inhibitors identified from Aegle marmelos
| Lead molecules | Ras proteins | ∆Gbind(kcal/mol) | Inhibition constant (KI) |
| α-amyrin | H-Ras | -9.94 | 51.69 nM |
| Lupeol | H-Ras | -9.62 | 89.46 nM |
| Betulinic acid | K-Ras | -8.51 | 577.15 nM |
| β-amyrin | K-Ras | -8.41 | 684.60 nM |
| Lupeol | K-Ras | -8.40 | 695.44 nM |
| α-amyrin | K-Ras | -8.28 | 850.02 nM |
| β-amyrin | N-Ras | -9.49 | 110.73 nM |
| α-amyrin | N-Ras | -8.78 | 367.60 nM |
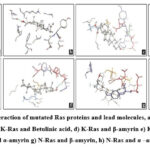
As mentioned above, the docked results of 88 molecules on mutated H-Ras (1P2U) revealed that about 78 molecules possessed free energy of binding ≤-5 kcal/mol. Similarly, 74 molecules on K-Ras (5UFE) and 77 molecules on N-Ras (5UHV) were selected as hit molecules respectively. The selected lead molecules on each Ras protein such asα-amyrin and Lupeol on H-Ras, betulinic acid, β-amyrin and lupeol on K-Ras, β-amyrin and α-amyrin on N-Ras were depicted in Fig. 2. The molecule α-amyrin was selected as lead molecule on mutated H-Ras and N-R as according to the lead selection criteria. For mutated K-Ras, it showed only a negligible difference with the lead molecules selected. Therefore, α-amyrin can be considered as the lead molecule for mutated Ras proteins.
The binding interactions between the docked ligand and protein are as follows (Fig.3, Table3). H-Ras and α-amyrin formed a single hydrogen bond (VAL29:H64) with a bond length of 2.13 Å and O..H-O bond type. H-Ras and lupeol interacted through a single hydrogen bond (ASP119:H75) with a distance of 2.01Å and O..H-O bond type. Likewise, single H-bonds were observed between K-Ras and β-amyrin (GLN61:H63, 2.09Å, O..H-O) and between K-Ras and lupeol (GLU31:HN1, 2.07Å, N-H..O). K-Ras and α-amyrin interacted through a single hydrogen bond (ASP30:H64) with a distance of 2.194 Å and O..H-O bond type. Betulinic acid formed two H-bonds with K-Ras, specifically at GLY13:HN (1.8 Å) and LYS16:HZ1 (1.77 Å), both of N-H..O type.N-Ras interacted with β-amyrin through one H-bond (THR35:H63, 2.03 Å, O..H-O), but no interactions were seen with α-amyrin. The analysis revealed that the lead molecules generally formed robust H-bonds with Ras proteins, with bond lengths ranging from 1.7Å to 2.2Å and N-H..O and O-H..O bond types.
Table 3 : Molecular interaction between lead molecules and mutated Ras proteins
| Lead
molecules |
RAS
proteins |
H-bond |
Bond
Type |
Bond
length |
Energy
(kcal/mol) |
Phi(Å) |
Theta(Å) |
| α-amyrin | H-Ras | VAL29:H64 | O..H-O | 2.13 | -2.27 | 129.66 | 177.48 |
| Lupeol | H-Ras | ASP119:H75 | O..H-O | 2.01 | -1.31 | 121.43 | 143.66 |
| Betulinicacid | K-Ras | GLY13:HN
LYS16:HZ1 |
N-H..O
N-H..O |
1.8
1.77 |
-7.379
0.674 |
174.3
138.82 |
168.56
122.76 |
| ß-amyrin | K-Ras | GLN61:H63 | O..H-O | 2.09 | -0.28 | 138.79 | 120.13 |
| Lupeol | K-Ras | GLU31:HN1 | N-H..O | 2.07 | -4.47 | 93.21 | 157.53 |
| ß-amyrene | N-Ras | THR35:H631 | O..H-O | 2.03 | 1.4 | 130.17 | 136.78 |
| α-amyrin | N-Ras | No H-bonds |
Molinspiration property prediction tool and SWISSADME was used to evaluate the drug-likeness of the lead molecules (Table 4).
Table 4: Drug likeness properties predicted by Molinspiration and SWISSADME tools
| Drug likeness properties | α-amyrin | Betulinic acid | lupeol | β-amyrin |
| MiLogP | 8.08 | 7.04 | 8.29 | 8.02 |
| TPSA | 20.23 | 57.53 | 20.23 | 20.23 |
| Number of atoms | 31 | 33 | 31 | 31 |
| Molecular weight | 426.7 | 456.7 | 426.7 | 421.73 |
| H-bond acceptor | 1 | 3 | 1 | 1 |
| H-bond donor | 1 | 2 | 1 | 1 |
| Number of violations | 1 | 1 | 1 | 1 |
| Number of rotational bonds | 0 | 2 | 1 | 1 |
| Volume | 461.05 | 472.04 | 461.6 | 460.7 |
| Bioavailability | 0.55 | 0.85 | 0.55 | 0.55 |
| Water solubility | Low solubility | Low solubility | Low solubility | Low solubility |
| Synthetic accessibility | 6.17 | 6.04 | 5.49 | 6.17 |
The bioactivity profiles of the lead molecules were predicted using Molinspiration, covering GPCR ligands, ion channel modulators, kinase inhibitors, nuclear receptor ligands, protease inhibitors, and enzymes shown in Table 5. The bioavailability radar of each lead molecule is created using SWISSADME and in which the pink zone describes appropriate physic-chemical space for oral bioavailability. In bioavailability radar (Fig. 4), LIPO represents lipophilicity (XLOGP3), SIZE indicates molecular weight, POLAR denotes polarity (topological polar surface area), INSOLU signifies water insolubility (log S scale), INSATU reflects unsaturation (sp3 hybridization), and FLEX represents flexibility (rotatable bonds).
Table 5: Predicted bioactivity scores for the lead molecules
| Lead molecules | GPCR
ligand |
Ion channel
modulator |
Kinase inhibitor | Nuclear receptor
ligand |
Protease inhibitor | Enzyme inhibitor |
| α-amyrin | 0.22 | -0.02 | -0.41 | 0.79 | 0.19 | 0.6 |
| Betulinic acid | 0.31 | 0.03 | -0.5 | 0.93 | 0.14 | 0.55 |
| Lupeol | 0.27 | 0.11 | -0.42 | 0.85 | 0.15 | 0.52 |
| β-amyrin | 0.22 | -0.05 | -0.31 | 0.67 | 0.11 | 0.56 |
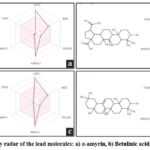 |
Figure 4: Bioavailability radar of the lead molecules: a) α-amyrin, b) Betulinic acid, c) Lupeol, d) β-amyrin
|
Discussion
Ras proteins are proto-oncogenes that are commonly mutated in many human cancers. Functional anomaly of Ras function ended up with the hyperproliferative developmental disorders and cancer and is correlated with a single amino acid substitution resulting from a mutation at specific codons 12, 13 or 61. In colon cancer (adenocarcinoma), lymphomas (Hodgekins lymphoma), leukemias (AML), thyroid and (anaplastic and follicular carcinoma) high frequencies of Ras mutations (>10%) are found in lung cancers, particularly large-cell and non-small-cell carcinomas.The K-Ras locus is predominantly affected by oncogenic mutations.12Although the protein products of the three Ras genes are identical, K-Ras is the more predominantly mutated Ras in cancer.14
The evolution of human civilization and the use of plant resources for healing have been closely linked, ultimately giving rise to modern medicine.23 A. marmelosis an important medicinal plant which has enormous ethnomedicinal applications and used in different traditional and folk medicines.24The plant is enriched with various classes of active chemical constituents such as alkaloids, steroids, coumarins, terpenoids, flavonoids and many other polyphenols that are responsible for the multifaceted pharmacological and biological activities.
Earlier investigations revealed that the hydroalcoholic extract of A. marmelos leaves possesses anticancer properties, inhibiting the growth of various cancer cells, such as K562, Raji, Jurkat, and MCF714. The antiproliferative and antioxidant activity assay on ethanolic extract of A. marmelosleaves in Dalton’s Lymphoma Ascites (DLA)-bearing mice demonstrated promising result.25
The previous investigations on the hydroalcoholic extract of A. marmelosleaves have shown anticancer effect in the animal model of Ehrlich ascites carcinoma, growth of leukemic K562, B-lymphoid Raji, T-lymphoid Jurkat, erythroleukemic HEL, melanoma Colo 38, and breast cancer cell lines MCF7 and MDA-MB-23114. The antiproliferative and antioxidant activity assay on ethanolic extract of A. marmelosleaves in mice bearing Dalton’s Lymphoma Ascites (DLA) demonstrated promising result.25
The potential anticancerous activity of the plant A. marmelosbestowed by the diverse phytoconstituents and analysis of these phytomolecules against mutated three Ras proteins were seldom investigated. Computer aided drug discovery is an indispensable technique to bring down the far-reaching experimentation mandatory for the unearthing of lead molecules. Hence, virtual screening through in silico molecular docking of 89 phytochemicals from A. marmelosagainst three mutated Ras proteins were carried out to determine effective lead molecules against various Ras proteins. The analysis of the overall results proclaimed that the molecule α-amyrin showed promising free energy of binding on all the three mutated Ras proteins even though betulinic acid and β-amyrin were identified as potential lead candidates on K-Ras and N-Ras respectively.
Theoretically, ideal drug-like molecules usually possess logP values between -0.4 and 5.6, molecular weights between 160-480 g/mol, molar refractivity between 40-130, and 20-70 atoms, following Lipinski’s rule of five, whereas natural compounds frequently deviate from these parameters.25 The drug likeness prediction of all the lead molecules showed violation in miLogP (>4.15) and by making derivatives or by changing slight alteration in the chemical structure through combinatorial techniques may enhance the property. Many researchers reported that aplenty of existing drugs violate theoretical prediction of drug likeness26 specifically naturalproducts.27 In light of this, more extensive study is required for the development of these lead molecules as drugs against mutated Ras proteins mediated tumors.
Conclusion
The overall results of the virtual screening of phytochemicals in A. marmelos on three mutated Ras proteins revealed that the plant has inhibitory effects on the three mutated Ras proteins. Among the 89 phytochemicals screened, most of the molecules showed moderate inhibitory activity (≤-5kcal/mol) on all the three mutated Ras proteins. In silico drug-likeness and bioactivity prediction of the lead molecules showed that they are within the range of values except miLogP. Hence, it can be concluded that by applying combinatorial chemical approach the lead molecules will enhance the violated property and ensure them as drug molecules. The identified lead molecules exhibit anti-inflammatory and antioxidant properties, which may help in reducing neuroinflammation and neurodegeneration. Alpha amyrin, betulinic acid, lupeol, and beta-amyrins are pentacyclic triterpene compounds that exhibit a range of pharmacological properties, particularly anticancer activity, making them promising candidates for drug development. A combinatorial approach and novel formulation strategies, such as nanoemulsionscan improve the lead molecule’s bioavailability and efficacy. Moreover, pharmacological validation of these lead molecules in a relevant biological system is mandatory for the development as drug molecules. Hence, further preclinical in vitro and in vivo evaluations are essential for the development of the lead molecules as drugs against the mutated Ras proteins.
Acknowledgment
We thank the Principal, Assumption College Autonomous, Changanassery, Kerala for providing the facilities and encouragement.
Conflict of Interest
The authors declare no conflict of interest, financial or otherwise.
Funding Sources
The authors received no financial support for the research, authorship and/or publication of this article.
Data Availability Statement
This statement does not apply to this article.
Ethics statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Clinical Trial Registration
This research does not involve any clinical trials.
Permission to reproduce material from other sources
Not Applicable
Author Contribution
Nisha Nisha Bhavan Chandran: Conceptualization, Methodology, Writing- Original Draft.
Rogimon Plammoottil Thomas: Supervision, Writing- Review& Editing.
References
- Tariq A., Majeed I, Khurshid A. Types of Cancers Prevailing in Pakistan and their Management Evaluation. Asian Pac J Cancer Prev. 2015;16:3605-16.doi: 10.7314/apjcp.2015.16.9.3605.
CrossRef - Iqbal J, Abbasi B A, Mahmood T, et al. Plant-derived anticancer agents: A green anticancer approach. Asian Pac J Trop Biomed. 2017;7:1129-50.https://doi.org/10.1016/j.apjtb.2017.10.016.
CrossRef - Sever R, Brugge J S. Signal transduction in cancer. Cold Spring Harb Perspect Med. 2015;5(4):a006098.doi: 10.1101/cshperspect.a006098.
CrossRef - Hassanpour S H, Dehghani M. Review of cancer from perspective of molecular. J Cancer Res Pract. 2017;4:127-9.https://doi.org/10.1016/j.jcrpr.2017.07.001.
CrossRef - Santos E, Nebreda A R. Structural and functional properties of Ras proteins. FASEB J. 1989;5:2151-63.doi: 10.1096/fasebj.3.10.2666231.
CrossRef - Tripathi B K., Hirsh N H, Qian X, et al. The pro-oncogenic noncanonical activity of a RAS•GTP:RanGAP1 complex facilitates nuclear protein export. Nat Cancer. 2024;5:1902-18.https://doi.org/10.1038/s43018-024-00847-5.
CrossRef - Simanshu D K, Nissley D V, McCormick F. Ras Proteins and Their Regulators in Human Disease. Cell. 2017;170:17-33.doi: 10.1016/j.cell.2017.06.009.
CrossRef - Pierotti M A, Sozzi G., Croce C M. Mechanisms of oncogene activation. In: Kufe DW, Pollock RE, Weichselbaum RR, et al., editors. Holland-Frei Cancer Medicine. 6th ed. Hamilton (ON): BC Decker; 2003.
- Medarde A F, Santos E. Ras in Cancer and Developmental Diseases. Genes Cancer. 2011;2:344-58.doi: 10.1177/1947601911411084.
CrossRef - Yang X, Wu H. RAS signaling in carcinogenesis, cancer therapy and resistance mechanisms. J Hematol Oncol. 2024;17(1):108.doi: 10.1186/s13045-024-01631-9.
CrossRef - Hood F E, Sahraoui Y M, Jenkins R E, Prior I A. Ras protein abundance correlates with Ras isoform mutation patterns in cancer. Oncogene. 2023;42:1224-32.doi: 10.1038/s41388-023-02638-1.
CrossRef - Khan T, Ali M, Khan A, Nisar P, Jan SA, Afridi S, Shinwari ZK. Anticancer Plants: A Review of the Active Phytochemicals, Applications in Animal Models, and Regulatory Aspects. Biomol. 2019; 27;10(1):47. doi: 10.3390/biom10010047.
CrossRef - Rahman S, Parvin R. Therapeutic Potential of Aegle marmelos (L.)-An overview. Asian Pac J Trop Dis. 2014;4:71-7.doi: 10.1016/S2222-1808(14)60318-2.
CrossRef - Baliga M S, Thilakchand K R, Rai M P, Rao S, Venkatesh P. Aegle marmelos (L.) Correa (Bael) and its Phytochemicals in the Treatment and Prevention of Cancer. Integr Cancer Ther. 2013;12:187-96.doi: 10.1177/1534735412451320.
CrossRef - Guex N, Peitsch M C. SWISS-MODEL and the Swiss-PdbViewer: an environment for protein modelling. Electrophoresis. 1997;18:2714-23.doi: 10.1002/elps.1150181505.
CrossRef - Nisha N C, Sreekumar S, Evans D A, Biju C K. In vitro and in silico validation of anti-cobra venom activity and identification of lead molecules in Aegle marmelos (L.) Correa. Curr Sci. 2018;114(6):1214-9.doi:10.18520/cs/v114/i06/1214-1221.
CrossRef - Morris G. M, Huey R, Lindstrom W, et al. AutoDock4and AutoDock Tools4: Automated docking with selective receptor flexibility. J Comput Chem. 2009;30:2785-91.doi: 10.1002/jcc.21256.
CrossRef - The PyMOL Molecular Graphics System, Version 1.2r3pre, Schrödinger, LLC.
- Lipinski C A, Lombardo F, Dominy B W, Feeney P J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev. 2001;46(1-3):3-26.doi: 10.1016/s0169-409x(00)00129-0.
CrossRef - Nisha N C, Sreekumar S, Biju C K, Krishnan P N. Identification of lead compounds with cobra venom neutralizing activity in three Indian medicinal plants. Int J Pharm Pharm Sci. 2014;6(2):536-41.
- Wardecki D, Dołowy M, Bober-Majnusz K. Assessment of Lipophilicity Parameters of Antimicrobial and Immunosuppressive Compounds. Molecules. 2023;28(6):2820.https://doi.org/10.3390/molecules28062820.
CrossRef - Nisha N C, Sreekumar S, Biju C K. Identification of lead compounds with cobra venom detoxification activity in Andrographis paniculata (Burm. F.) Nees through in silico method. Int J Pharm Pharm Sci. 2016;8(7):212-7.
- Petrovska B B. Historical review of medicinal plants’ usage. Pharmacogn Rev. 2012;6(11):1-5.doi: 10.4103/0973-7847.95849.
CrossRef - Vaza J S, Bhalerao SA. Bael (Aegle marmelos L.) as a potential medicinal tree: an overview. World J Pharm Sci. 2018;7(11):1406-14.
- Chockalingam V, Kadali S S, Gnanasambantham P. Antiproliferative and antioxidant activity of Aegle marmelos (Linn.) leaves in Dalton’s Lymphoma Ascites transplanted mice. Indian J Pharmacol. 2012;44(2):225-9.doi: 10.4103/0253-7613.93854.
CrossRef - Kadam R U, Roy N. Recent trends in drug-likeness prediction: A comprehensive review of in silico methods. Indian J Pharm Sci. 2007;69(5):609-15.
CrossRef - Ganesan A. The impact of natural products upon modern drug discovery. Curr Opin Chem Biol. 2008;12(3):306-7.doi: 10.1016/j.cbpa.2008.03.016.
CrossRef
Abbreviations list
∆Gbind– free energy of binding
DNA – Deoxy ribo nucleic acid
kcal/mol – Kilo calories per molecule
H-RAS – Harvey rat sarcoma viral oncogene
N-RAS – Neuroblastoma rat sarcoma viral oncogene
K-RAS – Kirsten rat sarcoma viral oncogene
etc– Et cetera
A. marmelos– Aegle marmelos
L. – Linnaeus
viz. – videlicet/namely
RCSB –Research Collaboratory for Structural Bioinformatics
g/mol – gram per mol
et al. – etaliae/ and others
Fig. – Figure
SMILES – Simplified Molecular Input Line Entry Systems
nrotb – number of rotatable bonds
miLogP– Molinspiration Logarithm of the Partition Coefficient
logP – logarithms of the Octanol-Water Partition Coefficient
TPSA- Total Polar Surface Area
MW – Molecular weight
ON – Hydrogen bond acceptors / Oxygen and Nitrogen atoms
OHNH – Hydrogen bond donors / Hydroxyl and amine groups
ROTB – Rotatable bonds
TPSA – Topological Polar Surface Area
Accepted on: 21-01-2026
Second Review by: Dr. Ankur Vashi
Final Approval by: Dr. Wagih Ghannam