Investigating the Ability of Lysinibacillus spp. to Detoxify Heavy Metals Using Bibliometric and Network Analysis
1Department of Agriculture, Jharkhand Rai University (JRU), Raja Ulatu, Jharkhand, India.
2Department of Biotechnology, Panskura Banamali College (Autonomous), Panskura, West Bengal, India.
Corresponding Author Email: bhattacharyya_nandan@rediffmail.com
DOI : http://dx.doi.org/10.13005/bbra/3490
ABSTRACT:Heavy metal contamination represents a persistent global environmental challenge due to its non-biodegradable nature, ecological toxicity, and severe risks to human health. Microbial bioremediation has emerged as a sustainable alternative to conventional physicochemical methods, with Lysinibacillus species gaining increasing attention for their exceptional metal tolerance and detoxification capabilities. The present study provides a comprehensive bibliometric and network-based evaluation of global research trends related to Lysinibacillus-mediated bioremediation of heavy metals. Bibliometric data were retrieved from the Dimensions AI database and analysed using VOSviewer to identify publication trends, leading countries, institutions, journals, and collaborative networks. To complement this analysis, a curated gene-species interaction network was constructed and analysed in Cytoscape to elucidate functional relationships among key microbial taxa and metal-resistance genes. The results reveal a sharp increase in research output since 2017, with India, China, and the United States emerging as major contributors. Network analysis identified Lysinibacillus sphaericus as a central and multifunctional node strongly associated with critical metal-resistance genes such as merA, arsB, and czcC, as well as pathways involved in organic pollutant degradation. The integrated findings highlight the ecological and biotechnological significance of Lysinibacillus, particularly L. sphaericus, as a keystone organism for designing effective synthetic microbial consortia for complex heavy metal remediation strategies.
KEYWORDS:Bibliometric; Bioremediation; Consortia; Gene-species interaction network; Heavy metal; Lysinibacillus
Introduction
The pervasive issue of heavy metal contamination has emerged as a significant concern due to its detrimental impacts on animals, plants, and humans, as well as its disruptive effects on ecosystems.1 Heavy metals are characterized by their non-biodegradable and persistent nature, which exacerbates environmental degradation and poses serious health risks.2 Although HMs and other metalloids occur naturally in the Earth’s crust, their recalcitrant properties hinder degradation processes. Bioaccumulation from diverse sources, including air, soil, and water, allows these substances to penetrate biota and move upward through food webs over time.3 Natural phenomena and, increasingly, anthropogenic activities introduce HMs into the environment. Urbanization, industrialization, and intensive agriculture frequently contaminate soils with cadmium (Cd), chromium (Cr), and related pollutants.4 Documented soil pollution cases involving these metals span Malaysia.5-7 Even at low concentrations, HMs can disrupt metabolic functions; at higher levels, they interfere with functional groups and essential metal ions in biological systems. By modifying active sites on biomolecules, they become toxic to both microbes and higher organisms. Agricultural runoff compounds the problem by delivering mercury and other toxins to aquatic ecosystems, ultimately threatening consumers of contaminated fish.8 In animals, chronic cadmium exposure damages renal proximal tubules and may induce osteoporosis by disturbing calcium metabolism.9, 10 Conventional physicochemical remediation techniques, though rapid and effective, are costly, energy-intensive, generate toxic sludge, and are impractical for large areas.11 Bioremediation offers a more economical and eco-friendly alternative; microorganisms can immobilize, transform, or remove non-biodegradable contaminants.12 Among these microbes, Lysinibacillus species exhibit exceptional tolerance, adaptability, and metal binding capacity, making them promising agents for HM clean-up.13, 14 Scientific interest in Lysinibacillus-mediated detoxification has risen sharply since the mid-2010s,15 yet few studies have systematically mapped the global research landscape. Bibliometric analysis fills this gap by revealing patterns, influential works, and research fronts.16 Accordingly, the present study combines Dimensions AI data with VOSviewer network visualizations to chart the evolution and structure of scholarship on Lysinibacillus and heavy-metal bioremediation.17, 18 Recent advances in systems biology and computational tools, such as Cytoscape, have enabled visualization and analysis of complex gene-microbe interaction networks for consortia development.19 These analyses are essential for pinpointing key functional genes, assessing modular structures within networks, and guiding the design of synthetic microbial consortia for targeted remediation.20, 21 This study aims to provide a comprehensive understanding of the role of Lysinibacillus species in heavy metal bioremediation by integrating bibliometric and network-based approaches. The objectives are to analyze global research trends, influential contributors, and collaboration patterns through bibliometric mapping; to identify major research themes and emerging hotspots associated with Lysinibacillus mediated remediation; to construct and evaluate a curated gene species interaction network in Cytoscape for identifying key functional genes, central microbial species, and important network clusters; to highlight the bioremediation potential of Lysinibacillus sphaericus and related taxa based on their genetic attributes and network positioning; and to use these insights to inform the development of synthetic microbial consortia and outline future research directions in heavy-metal bioremediation.
Materials and Methods
Data Collection
A thorough literature review was performed utilizing a prominent scientific database such as Dimension AI. This review encompassed a variety of publications such as articles, reviews, conference proceedings, and other scholarly works that concentrate on Lysinibacillus species as a strong candidate for heavy metal bioremediation. The search utilized keywords such as “Lysinibacillus and heavy metal,” “Lysinibacillus species,” “Heavy metal Bioremediation,” “environmental upgradation,” “ecosystem restoration,” and other related terms. The scope of the search was restricted to English-language publications and spanned from the establishment of these databases to the most current data available.22
Inclusion and Exclusion Criteria
The criteria for inclusion involved studies that specifically referenced Lysinibacillus species as a potential candidate for heavy metal bioremediation and explored their roles in the restoration of the ecosystem, improvement in soil nature and human health, as well as articles that examined the mechanisms of action and advantages of Lysinibacillus in removal heavy metal from soil. Conversely, the exclusion criteria eliminated studies unrelated to bioremediation, those that focused on different bacterial genera, and non-scholarly sources such as opinion articles and news reports.23
Bibliometric Analysis
Data Extraction and Preparation: The data extracted comprised publication titles, authors, publication years, journal names, keywords, abstracts, citations, and institutional affiliations. This information was subsequently organised in a systematic format for further analysis.24
Tools Used
VOSviewer was utilized to visualize and analyze co-authorship relationships, keyword co-occurrences, and citation networks, thereby facilitating the identification of thematic clusters and mapping the intellectual landscape of the field.25 Dimension AI was employed to pinpoint key authors, institutions, and countries that significantly contribute to the research on Lysinibacillus species as potential candidates for heavy metal bioremediation and monitor citation metrics and emerging research trends.26
Analysis Framework
The analytical approach included co-authorship analysis to assess collaborative networks among researchers and institutions, and citation analysis to evaluate the impact of the research (Zupic et al. 2015). The Dimension AI source library files were exported as CSV and RIS files and are listed below:
https://export.digital-science.com/2025-04-01/272248fb7d69547ac73191ecd0463df5/Dimensions-Publication-2025-04-01_09-42-53.csv.zip.
Data Compilation
Gene and species information was compiled through an extensive literature survey and validated using NCBI GenBank entries. The curated dataset included genes associated with heavy-metal resistance (such as merA, arsB, copA, czcC), organic pollutant degradation (xylE, katG, pnpA), and extracellular electron transfer (omcA, mtrA, pilA). All entries were organized into a structured Excel file to ensure consistency and accuracy.27
Network Construction in Cytoscape
The curated interaction dataset was imported into Cytoscape v3.10 in the form of a node-edge table. Nodes represented either genes or microbial species, while edges denoted validated gene-species associations. Nodes were color-coded according to cluster memberships, and the overall network layout was optimized to enhance modular visibility and structural interpretation.28
Cluster and Topology Analysis
ClusterMaker2 was employed to identify densely interconnected subgraphs within the network. Network Analyzer was then used to compute topological parameters, including centrality, node degree, and clustering coefficients. Key hub genes and species were identified based on degree values, and a bar graph depicting cluster size distribution was generated to enable comparative visualization of cluster densities.29
Rank Integration
The final ranking dataset was integrated to evaluate the relative importance of microbial species within the network. Lysinibacillus sphaericus appeared consistently within the top five species based on combined criteria such as gene abundance, network centrality, and functional relevance to heavy-metal detoxification.30
Results
Starting from 2017, this research has focused on investigating a bibliometric study on Lysinibacillus used in heavy metal bioremediation.31 The years 2024, 2023, and 2022 have shown a notable rise in publications on this topic, with 859, 650, and 653, respectively.32 In terms of publication categories, the analysis has identified 2512 articles, 1006 book chapters, 583 edited books, 72 preprints, 28 monographs, and 5 proceedings represented in Fig.1.33 Environmental science continues to dominate the field of research, with an impressive publication count of 1692, while biological science and microbiology follow just behind with 1679 and 1009 publications, respectively.34 There are 11 research domains presented in Table 1 related to the Sustainable Development Goals based on publication metrics.
Table 1: Top 16 Research fields, Sustainable Development Goal based on publication, and Co-authorship analysis
| Field of Research | Type of analysis: Co-authorship
Unit of analysis: Authors |
|||
| Research category | Publications | Author | Documents | Citations |
| Environmental Sciences | 1687 | Das, Alok Prasad | 23 | 903 |
| Yaqoob , Asim Ali | 12 | 744 | ||
| Biological Sciences | 1678 | Ibrahim, Mohamad Nasir Mohamad | 12 | 741 |
| Microbiology | 1008 | Gaur, Vivek Kumar | 11 | 660 |
| Pollution and contamination | 878 | Varjani, Sunita | 10 | 1571 |
| Engineering
|
843 | Rahman, Aminur | 10 | 194 |
| Mandal ,Abul | 9 | 163 | ||
| Agriculture, Veterinary and Food Science | 570 | Ngo,Huu Hao | 7 | 1029 |
| Chemical Engineering | 495 | Sharma ,Poonam | 7 | 527 |
| Industrial Biotechnology | 386 | Ghosh ,Sibdas | 7 | 182 |
| Environmental Engineering | 353 | Nahar,Noor | 7 | 149 |
| Crop and pest Production | 341 | Kour,Divjot | 7 | 89 |
| Environmental Biotechnology | 333 | Jass ,Jana | 6 | 156 |
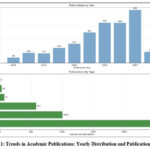 |
Figure 1: Trends in Academic Publications: Yearly Distribution and Publication Types.
|
Environmental Science leads with 1692 publications, followed by Biological Science with 1679 publications. Microbiology has 1009 publications, Pollution and Contamination has 880 publications, and Engineering has 846 publications. Other fields are Agricultural, Veterinary and food science, Chemical Engineering, Environmental Biotechnology and Industrial Biotechnology.35 The coauthorship analysis represented in the Fig.2. The analysis of co-authorship by country represented in Table 2 and Fig.3 demonstrates India’s leading position with 821 documents and 26661 citations, showcasing a strong international collaboration and impact. Following India, China has 581 documents and 18875 citations, emphasizing its significant research contribution. The United States ranks third with 132 documents and 5901 citations, highlighting its active involvement in global research networks.36 Noteworthy countries like Saudi Arabia, Pakistan, and South Korea also show substantial participation in research activities, as indicated by their document numbers and citations.37
 |
Figure 2: Co-authorship analysis
|
 |
Figure 3: Major Contributing Countries
|
Table 2: Major Contributing Countries and Journals
| Type of analysis: Co-authorship, Unit of analysis: Country | Type of analysis: Citations, Unit of analysis: Source (Journal) | ||||
| Country | Documents | Citations | Source | Documents | Citations |
| India | 821 | 26661 | |||
| China | 581 | 18875 | Environmental Research and pollution research | 96 | 2255 |
| United States | 132 | 5901 | Chemosphere | 86 | 3447 |
| Pakistan | 107 | 3262 | Science of the Total Environment | 76 | 3308 |
| Malaysia | 100 | 3769 | Journal of environmental management | 69 | 4031 |
| Journal of Hazardous material | 67 | 4122 | |||
| South Korea | 93 | 4821 | |||
| Saudi Arabia | 90 | 2480 | Frontiers in Microbiology | 61 | 3070 |
| Egypt | 69 | 5327 | Ecotoxicology and Environmental Safety | 44 | 3536 |
| United Kingdom | 68 | 6073 | Microorganisms | 40 | 1058 |
| Nigeria | 60 | 1499 | Environmental pollution | 31 | 642 |
| Australia | 55 | 3322 | Environmental Research | 29 | 615 |
| Canada | 51 | 2767 | Biosource Technology | 27 | 2886 |
| Italy | 50 | 2302 | Geomicrobiology Journal | 27 | 597 |
| Spain | 48 | 1808 | World Journal of Microbiology and Biotechnology | 25 | 515 |
| Iran | 48 | 1142 | Water, Air & Soil Pollution | 24 | 282 |
| Germany | 40 | 1560 | Scientific Reports | 22 | 602 |
| Bangladesh | 32 | 688 | Applied Microbiology and Biotechnology | 19 | 840 |
| France | 29 | 1215 | International biodeterioration and biodegradation | 15 | 434 |
| Taiwan | 26 | 1185 | Journal of environmental chemical engineering | 15 | 375 |
| Sweden | 23 | 1276 | sustainability | 15 | 268 |
The analysis focuses on collaborative research through co-authorship, presented in Table3 and Fig. 4 with a focus on the organization as the unit of analysis.
 |
Figure 4: Major Contributing Organizations
|
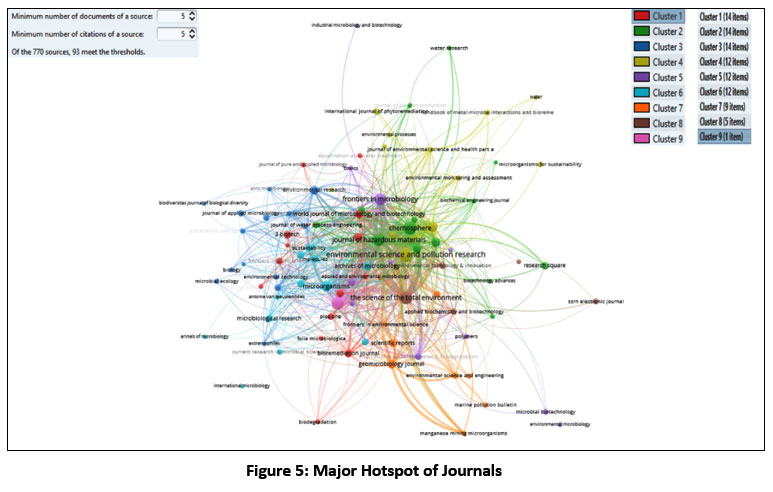 |
Figure 5: Major Hotspot of Journals
|
Table 3: List of major contributing Organizations and Co-citation analysis
| Type of analysis: Co-authorship, Unit of analysis: Organization | ||
| Name of Organization | Documents | Citations |
| Amity University | 49 | 1691 |
| Benaras Hindu University | 39 | 2047 |
| King Saud University | 39 | 966 |
| University of Punjab | 27 | 583 |
| University of Chinese academy of Science | 26 | 867 |
| Jiangsu University | 24 | 2819 |
| Lovely Professional university | 24 | 959 |
| Academy of Scientific and Innovative Research | 23 | 558 |
| Saveetha institute of Medical and Technical Sciences | 17 | 511 |
| Indian Institute of Toxicology Research | 16 | 1337 |
| Chandigarh University | 16 | 168 |
| University of Petroleum and Energy Studies | 12 | 269 |
| Sun Yat-Sen University | 11 | 241 |
| University of Technology Sydney | 10 | 1237 |
| Integral University | 9 | 587 |
| Centre for energy and environmental Sustainability | 8 | 442 |
| University of Boras | 6 | 837 |
| National Cheng Kung University | 6 | 334 |
| Ulsan National Institute Of Science and Technology | 6 | 223 |
| Gujrat Pollution control Board Gandhinagar | 5 | 770 |
Amity University leads with 49 documents and 1691 citations, demonstrating high productivity and impact. Banaras Hindu University follows with 39 documents and 2047 citations. King Saud University has 39 documents with 966 citations, showing a strong citation-to-publication ratio.38 The University of Punjab and the University of Chinese Academy have 27 and 26 documents, with citations of 583 and 867, respectively. Other significant contributors include Lovely Professional University and Academy of Scientific and Innovative Research, with 24 and 23 documents, respectively.39
The citation analysis based on journal sources presented in Fig.5 reveals that “Environmental research and Pollution Research” is the most cited journal with 96 documents contributing to 2255 citations, indicating its prominence in the field. “Chemosphere” follows closely with 86 documents and 3447 citations, reflecting a high citation rate per document. “Science of the Total Environment” ranks third with 76 documents and 3308 citations.40 Other significant journals include “Journal of Environmental Management,” “Journal of Hazardous Materials,” and “Frontiers in Microbiology,” each contributing to the research landscape with their respective document numbers and citations.41 India leads in document production with 821 documents and 26661 citations, followed by China with 581 documents and 18875 citations, and the US with 132 documents and 5901 citations.42 Pakistan stands out with 102 documents and 3262 citations. Other countries like Malaysia, Saudi Arabia, and South Korea also contribute significantly.43 India, China and the US, dominate research output and citations. China’s strong infrastructure and international collaboration are evident, while the US maintains high-quality research. India strikes a balance between output and impact. Some countries with high output have lower citation counts. The varying citation counts for countries like Egypt, and Australia, relative to their document output, indicate that their research may be highly regarded or focused on influential topics.44
In the network analysis using Cytoscape followed by NCBI represented in Fig.6, the constructed gene-species interaction network revealed a clear modular structure comprising four major functional clusters linked to heavy metal detoxification, electron transfer, aromatic compound degradation, and ancillary metabolic pathways. The heavy metal detoxification module was the largest and most interconnected, centred on the hub gene copA and linked to key metal-resistance genes such as arsB, arsC, czcC, merA, and chrA. This cluster also included several metal-tolerant microbes, Bacillus subtilis, Achromobacter xylosoxidans, Cupriavidus metallidurans, Enterobacter cloacae, and particularly Lysinibacillus sphaericus highlighting its functional dominance in metal-stressed environments.45
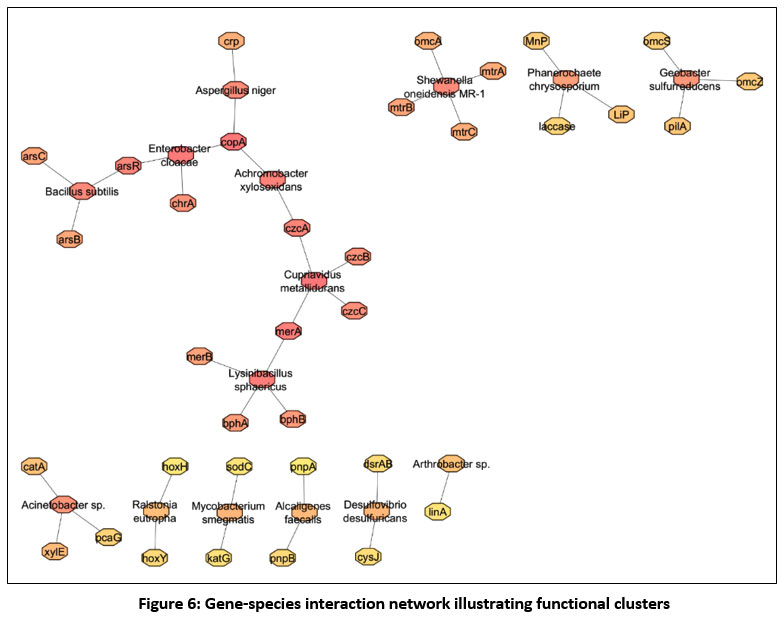 |
Figure 6: Gene-species interaction network illustrating functional clusters
|
Lysinibacillus sphaericus emerged as a significant node within this module, showing strong associations with both mercury-reduction genes (merA, merB) and biphenyl-degradation genes (bphA, bphB). This dual connectivity positions it as a bridge between metal-resistance and organic-pollutant degradation pathways. Rank integration confirmed its importance, placing it among the top five species based on gene abundance, centrality, and detoxification relevance analysis.46
A second prominent cluster consisted of extracellular-electron-transfer and ligninolytic genes such as omcA, omcS, mtrA, mtrB, mtrC, and pilA, associated with Shewanella oneidensis, Geobacter sulfurreducens, and Phanerochaete chrysosporium. This module represents organisms crucial for redox balancing and electron shuttle activity, key processes in metal reduction and wastewater treatment.47
The third cluster comprised genes involved in aromatic and xenobiotic degradation, including xylE, katG, pnpA, pnpB, and foxY, linked to versatile degraders such as Acinetobacter, Ralstonia eutropha, Mycobacterium smegmatis, and Alcaligenes faecalis. This module complements the metal-resistance pathways, indicating potential for co-removal of mixed organic-metal pollutants.48, 49
The final, smaller cluster included nodes with lower connectivity but still contributed additional metabolic functions that may support overall community resilience. Taken together, the network demonstrates that bioremediation efficiency relies on synergistic interactions across microbial groups rather than single-species performance. Lysinibacillus sphaericus, due to its central placement and multifunctional gene associations, stands out as a promising keystone organism for designing synthetic microbial consortia. The integration of network topology, functional clustering, and ranking scores provides a rational basis for selecting species combinations suited for complex contamination scenarios such as heavy-metal-rich soils, fly-ash environments, and industrial wastewater.
Discussion
The present study provides a comprehensive synthesis of global research trends and functional insights into Lysinibacillus-mediated heavy metal bioremediation by integrating bibliometric mapping with gene-species interaction network analysis. The sharp increase in publications after 2017 reflects a growing scientific recognition of microbial-based solutions as sustainable alternatives to conventional physicochemical remediation methods. This surge aligns with the global emphasis on eco-friendly technologies, circular bioeconomy concepts, and Sustainable Development Goals related to environmental protection and ecosystem restoration.
Bibliometric analysis revealed that environmental science, microbiology, and biological sciences dominate the research landscape, highlighting the inherently interdisciplinary nature of heavy metal bioremediation research. The strong representation of pollution, contamination, and engineering domains further indicates a shift from purely descriptive studies toward applied and solution-oriented research. India, China, and the United States emerged as leading contributors, reflecting differences in environmental pressures, industrialization patterns, and national research priorities. India’s prominent position can be attributed to extensive heavy metal contamination issues in soils and water bodies, coupled with strong academic interest in low-cost microbial remediation strategies. In contrast, China’s high citation impact suggests a focus on mechanistic and technologically advanced studies, while the United States maintains influence through high-quality, internationally collaborative research outputs
The dominance of journals such as Environmental Science and Pollution Research, Chemosphere, and Science of the Total Environment underscores the maturity of the field and confirms that Lysinibacillus-based remediation has transitioned from niche microbiological studies to mainstream environmental science discourse. These publication trends also indicate a gradual movement toward integrative studies combining genomics, systems biology, and environmental engineering.
The Cytoscape-based gene-species interaction network provides critical functional context to the bibliometric findings. The identification of a highly modular network architecture supports the concept that effective bioremediation is governed by cooperative microbial interactions rather than single-organism activity. Among all taxa analyzed, Lysinibacillus sphaericus consistently emerged as a central and multifunctional node, strongly associated with key heavy metal resistance genes such as merA, merB, arsB, and czcC. These genes are known to mediate mercury reduction, arsenic efflux, and multi-metal resistance, confirming the genetic basis of the organism’s high tolerance to metal-stressed environments
Notably, the dual association of L. sphaericus with both metal-detoxification genes and aromatic compound degradation genes (bphA, bphB) positions it as a critical functional bridge between inorganic and organic pollutant remediation pathways. This multifunctionality is particularly relevant for complex contamination scenarios such as fly ash-impacted soils, industrial effluents, and mixed waste streams, where metals coexist with persistent organic pollutants. The rank integration analysis further validates L. sphaericus as a keystone species based on its high network centrality, gene abundance, and detoxification relevance.
Beyond Lysinibacillus, the presence of distinct but interconnected clusters highlights the importance of functional complementarity in microbial remediation systems. The extracellular electron transfer cluster, dominated by Shewanella oneidensis and Geobacter sulfurreducens, emphasizes the role of redox-active microbes in metal reduction and electron shuttling processes. These organisms can enhance metal immobilization and transformation, particularly under anaerobic or low-oxygen conditions, thereby supporting the detoxification functions of Lysinibacillus species.
Similarly, the aromatic and xenobiotic degradation cluster, comprising genera such as Acinetobacter, Ralstonia, and Mycobacterium, indicates the system’s capacity for co-removal of organic contaminants. The coexistence of these clusters within the same network suggests that synthetic microbial consortia, rather than monocultures, are likely to achieve higher remediation efficiency, resilience, and adaptability under fluctuating environmental conditions
The integration of bibliometric trends with network topology provides a rational framework for evidence-based consortium design. While bibliometric analysis identifies influential species, genes, and research hotspots, network analysis elucidates functional dependencies and synergistic interactions. The central placement of L. sphaericus suggests that it can serve as a core organism around which auxiliary metal-reducing, electron-transferring, and organic-degrading microbes can be strategically assembled. Such consortia could be tailored for site-specific applications, including contaminated agricultural soils, industrial wastewater treatment, and fly ash-affected ecosystems.
Overall, the findings reinforce the notion that Lysinibacillus sphaericus is not merely a metal-tolerant bacterium but a systems-level contributor to complex remediation networks. Its multifunctional genetic repertoire and strong network connectivity make it an ideal candidate for next-generation bioremediation strategies that integrate microbiology, systems biology, and environmental engineering.
Conclusion
This study provides an integrated bibliometric and network-based evaluation of Lysinibacillus species in heavy-metal bioremediation. The bibliometric analysis reveals a sharp rise in global research output, led by India, China, and the United States, with strong contributions from major journals and institutions in environmental and biological sciences. These trends highlight growing interdisciplinary interest spanning microbiology, environmental science, and biotechnology. Complementing this, the Cytoscape gene-species network analysis identifies Lysinibacillus sphaericus as a central, multifunctional organism linked to key heavy-metal resistance genes (merA, merB, czcC) and aromatic-compound degradation genes (bphA, bphB). Its bridging position between metal-detoxification and organic-pollutant pathways underscores its suitability for multi-species remediation systems. The presence of supportive clusters involving electron-transfer microbes (Shewanella, Geobacter) and xenobiotic degraders (Acinetobacter, Ralstonia) further emphasizes the value of synergistic consortia. Overall, the combined findings establish L. sphaericus as a strong candidate for next-generation microbial consortia and provide a clear foundation for future applied and interdisciplinary research in bioremediation.
Acknowledgements
We are grateful to the Panskura Banamali College (Autonomous) Research Centre (P.B.C.R.C.) and the Department of Microbiology of Panskura Banamali College (Autonomous) for providing us with the basic research infrastructure, which has enabled us to conduct this work.
Funding Sources
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflict of Interest
The authors do not have any conflict of interest.
Data Availability Statement
The raw data is available in the following data repository link: https://mega.nz/folder/eVVDRC7b#eXLralskEXUytahqb CKLew
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
All the authors involved in the manuscript must assure their consent for the publication of the manuscript. Furthermore, individuals who provide writing assistance should be identified by the authors, and they must disclose the funding source for this assistance.
Clinical Trial Registration
This research does not involve any clinical trials.
Permission to reproduce material from other sources
Not Applicable
Author Contributions
Kajari Roy: Conceptualization, Methodology, Data Collection, Writing – Original Draft.
Nandan Bhattacharyya: Review, Editing and Supervision.
References:
- Mitra S, Chakraborty AJ, Tareq AM, Emran TB, Nainu F, Khusro A, Idris AM, Khandaker MU, Osman H, Alhumaydhi FA, Simal-Gandara J. Impact of heavy metals on the environment and human health: Novel therapeutic insights to counter the toxicity. J King Saud Univ Sci. 2022;34:101865. doi:10.1016/j.jksus.2022.101865
CrossRef - Edo GI, et al. Environmental persistence, bioaccumulation, and ecotoxicology of heavy metals. Chem Ecol. 2024;40(3):322–349. doi:10.1080/02757540.2024.2306839
CrossRef - Adewumi AJ, Ogundele OD. Hidden hazards in urban soils: A meta-analysis review of global heavy metal contamination (2010–2022), sources and its ecological and health consequences. Sustain Environ. 2024;10(1):2293239. doi:10.1080/27658511.2023.2293239
CrossRef - Cui Z, et al. Effects of ageing on surface properties of biochar and bioavailability of heavy metals in soil. Agriculture. 2024;14(9):1631. doi:10.3390/agriculture14091631
CrossRef - Abdullah SRS, Al-Baldawi IA, Almansoory AF, Purwanti IF, Al-Sbani NH, Sharuddin SSN. Plant-assisted remediation of hydrocarbons in water and soil: application, mechanisms, challenges and opportunities. 2020;247:125932. DOI: 10.1016/j.chemosphere.2020.125932
CrossRef - Abedi Sarvestani R, Aghasi M, Niknejad H. Health risk assessment of trace elements (Pb, Cd, Cu, Fe) in agricultural soil in Kerman City, Southeast of Iran. Nat Hazards. 2024;120(1):339–367. doi:10.1007/s11069-023-06218-0
CrossRef - Wang, H., Li, W., Zhu, C., & Tang, X. (2021). Analysis of heavy metal pollution in cultivated land of different quality grades in Yangtze River Delta of China. International Journal of Environmental Research and Public Health, 18(18), 9876. https://doi.org/10.3390/ijerph18189876
CrossRef - Charkiewicz AE, et al. Mercury exposure and health effects: What do we really know? Int J Mol Sci. 2025;26(5):2326. doi:10.3390/ijms26052326
CrossRef - Vijiyakumar N, Evan Prince S. A comprehensive review of cadmium-induced toxicity, signalling pathways, and potential mitigation strategies. Toxicol Environ Health Sci. 2025;17(1):79–94. doi:10.1007/s13530-024-00243-7
CrossRef - Tang C, et al. Cadmium exposure and osteoporosis: Epidemiological evidence and mechanisms. Toxicol Sci. 2025;205(1):1–10. doi:10.1093/toxsci/kfaf031
CrossRef - Dhokpande SR, et al. A review outlook on methods for removal of heavy metal ions from wastewater. Sep Purif Technol. 2024;350:127868. doi:10.1016/j.seppur.2024.127868
CrossRef - Wang S, Liu T, Xiao X, Li S. Advances in microbial remediation for heavy metal treatment: a mini review. Collagen Leather. 2021;3(1). DOI:10.1186/s42825-020-00042-z
CrossRef - Kumar MA, Swarnalatha AP, Naveen KN, Raju JNS, Kerur SS, Priyadharsini K. Biosorption of heavy metals using plant-derived sorbents: an environmental solution for industrial wastewater. Global Nest J. 2025;27(4). doi.org/10.30955/gnj.06874
- Tang X, Huang Y, Li Y, et al. Study on detoxification and removal mechanisms of hexavalent chromium by microorganisms. Ecotoxicol Environ Saf. 2021;208:111699. https://doi.org/10.1016/j.ecoenv.2020.111699
CrossRef - Li X, Gao Y, Ning X, Li Z. Research progress and hotspots on microbial remediation of heavy metal-contaminated soil: a systematic review and future perspectives. Environ Sci Pollut Res. 2023;30(56):118192–118212. doi.org/10.1007/s11356-023-30655-w
CrossRef - Yu JA, Chen Z, Gao W, et al. Global trends and prospects in research on heavy metal pollution at contaminated sites. J Environ Manage. 2025;383:125402. https://doi.org/10.1016/j.jenvman.2025.125402
CrossRef - Gandasari D, et al. Bibliometric and visualized analysis of social network analysis research on Scopus databases and VOSviewer. Cogent Bus Manag. 2024;11(1):2376899. doi:10.1080/23311975.2024.2376899
CrossRef - Hakkaraki VP. Exploring the evolution of bibliometric analysis: A comprehensive study of scientific publications from 1974 to 2024 using the Dimensions AI database. Asian J Inf Sci Technol. 2024;14(1):24–31. doi:10.70112/ajist-2024.14.1.3878
CrossRef - Sharma A, Tayal S, Bhatnagar S. Stress response in bacterial pathogens using network biology. Sci Rep. 2025;15:15342. https://doi.org/10.1038/s41598-025-91269-5
CrossRef - Doncheva NT, Morris JH, Gorodkin J, Jensen LJ. Cytoscape stringApp 2.0: analysis and visualization of heterogeneous biological networks. J Proteome Res. 2022;22(2):637–646. doi/full/10.1021/acs.jproteome.2c00651
CrossRef - Zhang F, Liu X, Zhang Y, et al. Gene regulatory networks in COVID-19 myocarditis. PLoS One. 2022;17(6):e0269386. https://doi.org/10.1371/journal.pone.0269386
CrossRef - Donthu N, Kumar S, Mukherjee D, Pandey N, Lim WM. How to conduct a bibliometric analysis: an overview and guidelines. J Bus Res. 2021;133:285–296. https://doi.org/10.1016/j.jbusres.2021.04.070
CrossRef - Moher D, Liberati A, Tetzlaff J, Altman DG; PRISMA Group. Preferred reporting items for systematic reviews and meta-analyses: the PRISMA statement. PLoS Med. 2009;6(7):e1000097. https://doi.org/10.1136/bmj.b2535
CrossRef - Aria M, Cuccurullo C. bibliometrix: an R-tool for comprehensive science mapping analysis. J Informetr. 2017;11(4):959–975. https://doi.org/10.1016/j.joi.2017.08.007
CrossRef - Vosoughian N, Mohammadi A, Hamayeli H. Bacteria as an efficient bacteriosystem for the synthesis of nanoparticles: a bibliometric analysis. Nano. 2021;16(14):2130014. https://doi.org/10.1142/S1793292021300140
CrossRef - Cabral JPS. Water microbiology: bacterial pathogens and water. Int J Environ Res Public Health. 2010;7:3657–3703. https://doi.org/10.3390/ijerph7103657
CrossRef - Pan P, Bhattacharyya N. Bioelectricity Production from Microbial Fuel Cell (MFC) Using Lysinibacillus xylanilyticus Strain nbpp1 as a Biocatalyst. Current Microbiology. 2023;80(8):252. doi:https://doi.org/10.1007/s00284-023-03338-5
CrossRef - Pan P, Bhattacharyya N. Electrogenic properties of Bacillus paramycoidesNBPP1 strain as a biocatalyst in the microbial fuel cell. Biofuels. 2024;15(8):1017-1028. doi:https://doi.org/10.1080/17597269.2024.2320986
CrossRef - Morris JH, Apeltsin L, Newman AM, et al. clusterMaker: a multi-algorithm clustering plugin for Cytoscape. BMC Bioinformatics. 2011;12:436. https://doi.org/10.1186/1471-2105-12-436
CrossRef - Utriainen M, Morris JH. clusterMaker2: a major update to clusterMaker. BMC Bioinformatics. 2023;24:134. https://doi.org/10.1186/s12859-023-05225-z
CrossRef - Zupic I, Čater T. Bibliometric methods in management and organization. Organ Res Methods. 2015;18(3):429–472. https://doi.org/10.1177/1094428114562
CrossRef - Zhang L, Xu W, Wang J. Global trends in microbial bioremediation of heavy metals. J Environ Manage. 2023;331:117264. https://doi.org/10.1016/j.jenvman.2023.117264
CrossRef - Elgarahy AM, Elwakeel KZ, Mohammad SH, Elshoubaky GA. A critical review of biosorption of dyes, heavy metals and metalloids from wastewater. Clean Eng Technol. 2021;4:100209. https://doi.org/10.1016/j.clet.2021.100209
CrossRef - Vigil TN, Johnson GC, Jacob SG, Spangler LC, Berger BW. Microbial Mineralization with Lysinibacillus sphaericus for Selective Lithium Nanoparticle Extraction. Environmental Science & Technology. Published online September 12, 2024. doi:https://doi.org/10.1021/acs.est.4c06540
CrossRef - Yu Y, Li Y, Zhang Z, et al. A bibliometric analysis using VOSviewer of publications on COVID-19. Annals of Translational Medicine. 2020;8(13):816-816. doi:https://doi.org/10.21037/atm-20-4235
CrossRef - Donthu N, Kumar S, Mukherjee D, Pandey N, Lim WM. How to conduct a bibliometric analysis: An overview and guidelines. Journal of Business Research. 2021;133:285-296. doi:https://doi.org/10.1016/j.jbusres.2021.04.070
CrossRef - Bhaskar P, Tiwari CK. Charming or chilling? A comprehensive review of ChatGPT’s in education sector. International Journal of Information and Learning Technology. Published online March 8, 2025. doi:https://doi.org/10.1108/ijilt-05-2024-0097
CrossRef - Arunachalam S, Viswanathan B. A historiographic analysis of fuelcell research in Asia – China racing ahead. Current Science. 2008;95(1):36-49. https://doi.org/10.2307/24103275
- Nadeem SM, Ahmad M, Zahir ZA, Javaid A, Ashraf M. The role of mycorrhizae and plant growth promoting rhizobacteria in improving crop productivity under stressful enhttps://doi.org/10.1016/j.biotechadv.2013.12. 005vironments. Biotechnol Adv. 2014;32(2):429–448.
CrossRef - Kumar A, Rani M, Thakur AK. Microbial remediation of heavy metals in polluted soil. Contemp Adv Sci Technol. 2024;7(2):135–148.DOI:10.70130/CAST.2024.7110
CrossRef - Tufail MA, et al. Recent advances in bioremediation of heavy metals and persistent organic pollutants: a review. Sci Total Environ. 2022;850:157961. https://doi.org/10.1016/j.scitotenv.2022.157961
CrossRef - Ali H, Khan E, Sajad MA. Phytoremediation of heavy metals: concepts and applications. 2013;91(7):869–881. http://dx.doi.org/10.1016/j.chemosphere.2013.01.075
CrossRef - Azubuike CC, Chikere CB, Okpokwasili GC. Bioremediation techniques–classification based on site of application: principles, advantages, limitations and prospects. World Journal of Microbiology and Biotechnology. 2016;32(11). doi:https://doi.org/10.1007/s11274-016-2137-x
CrossRef - Chakraborty J, Das S. Application of spectroscopic techniques for monitoring microbial diversity and bioremediation. Appl Spectrosc Rev. 2017;52(1):1–38. https://doi.org/10.1080/05704928.2016.1199028
CrossRef - Das S, Dash HR, Chakraborty J. Genetic basis and importance of metal resistant genes in bacteria for bioremediation of contaminated environments with toxic metal pollutants. Appl Microbiol Biotechnol. 2016;100(7):2967–2984. https://doi.org/10.1007/s00253-016-7364-4
CrossRef - Tejaswini MSSR, Zia J. Mercury bioremediation using extremophiles. Water Air Soil Pollut. 2025;236:1–24. https://doi.org/10.1007/s11270-025-08476-z
CrossRef - Saffarini D, Brockman K, Beliaev A, Bouhenni R, Shirodkar S. Shewanella oneidensis and extracellular electron transfer to metal oxides. In: Ehrlich HL, Newman DK, Kappler A, eds. Bacteria–Metal Interactions. Cham, Switzerland: Springer; 2015:21–40. https://doi.org/10.1007/978-3-319-18570-5_2
CrossRef - Ray RR, Pattnaik S. Alcaligenes faecalis: a bacterium for sustainable management of environment. Environ Qual Manag. 2024;34(1):e22189. https://doi.org/10.1002/tqem.22189
CrossRef - Reineke W, Schlömann M. Microbial degradation of pollutants. In: Maier RM, Pepper IL, Gerba CP, eds. Environmental Microbiology. 3rd ed. Cham, Switzerland: Springer; 2023:161–290. https://doi.org/10.1007/978-3-662-66547-3_6
CrossRef
Accepted on: 06-01-2026
Second Review by: Dr. Mytham Jabouri Abaulhussain
Final Approval by: Dr. Wagih Ghannam









