In Silico Evaluation of Berberine from Tinospora Cordifolia as a Potential Inhibitor of Kras in Colorectal Cancer: Molecular Docking and Pharmacokinetic Insights
Department of Pharmacology, Vignan Institute of Pharmaceutical Technology, Andhra Pradesh, India,
Corresponding author Email id: bharathimarni@gmail.com
DOI : http://dx.doi.org/10.13005/bbra/3503
ABSTRACT:Colorectal cancer (CRC) is among the most common and lethal cancers approximately 40–50% of colorectal cancers, >90% of pancreatic ductal adenocarcinomas, and ~30% of lung adenocarcinomas harbor activating KRAS mutat approximately 40–50% of colorectal cancers, >90% of pancreatic ductal adenocarcinomas, and ~30% of lung adenocarcinomas harbor activating KRAS mutations ions worldwide. Mutations in KRAS play a central role in driving tumor progression and resistance to targeted therapies. Natural compounds, particularly berberine derived from Tinospora cordifolia, have been reported to exert anticancer effects; however, their direct interaction with KRAS has not been well established. This study explored the therapeutic potential of berberine against KRAS using molecular docking and In-silico Absorption, Distribution, Metabolism, Excretion, and Toxicity (ADMET) analyses. Docking simulations performed with CB-Dock revealed a strong binding affinity, supported by a Pro SA-web Z-score of –6.9 and favourable binding interactions. Pharmacokinetic evaluation through Swiss ADME and pkCSM indicated high intestinal absorption (97.15%), moderate blood–brain barrier penetration, good oral bioavailability, and an acceptable toxicity profile. Structural validation using the Ramachandran plot confirmed the reliability of the protein–ligand complex, with 96% of residues located in the most favored regions. Collectively, these findings highlight berberine as a promising lead candidate for KRAS-targeted therapy in colorectal cancer, warranting further preclinical validation.
KEYWORDS:ADMET; Berberine; Colorectal cancer; KRAS; Molecular docking; pkCSM; Swiss ADME; Tinospora cordifolia
Introduction
Colorectal cancer (CRC) continues to pose a serious global health burden, ranking as the third most frequently diagnosed cancer and the second leading cause of cancer-related deaths.1 In 2020 alone, approximately 1.9 million new cases and more than 930,000 deaths were reported worldwide.2 The disease arises through a multistep process driven by genetic and molecular alterations, with KRAS, APC, and TP53 being among the most critical contributors. Mutations in KRAS, occurring in nearly 35–45% of CRC patients, are particularly significant.3 These mutations, most commonly at codons 12 and 13, lead to constitutive activation of the MAPK/ERK and PI3K/AKT pathways, thereby promoting uncontrolled cell proliferation, invasion, and therapeutic resistance.4
Clinically, CRC cases harboring KRAS mutations are especially challenging to treat. Patients with these mutations are largely unresponsive to anti-EGFR monoclonal antibodies such as cetuximab and panitumumab, limiting therapeutic options and worsening prognosis.5 Despite decades of research, KRAS has been labeled an “undruggable” target due to its smooth surface topology and strong affinity for GTP/GDP. Although covalent inhibitors specific to the KRAS^G12C^ mutation have recently been approved for clinical use, these agents are mutation-specific and do not cover the broader mutation spectrum commonly observed in CRC.6
The RAS/MAPK and PI3K/AKT pathways are central regulators of cell proliferation, survival, and metabolism. KRAS, a member of the RAS family (originally identified in the Rat Sarcoma virus, hence “RAS”), functions as a molecular switch that cycles between an active GTP-bound state and an inactive GDP-bound state. Oncogenic mutations—most commonly at codons 12, 13, or 61—impair GTP hydrolysis, locking KRAS in a constitutively active conformation that drives uncontrolled downstream signaling.
Upon activation, KRAS engages two major effector cascades
The RAF–MEK–ERK (MAPK/ERK) pathway, which promotes transcription of genes involved in cell cycle progression (e.g., via cyclin-dependent kinases, CDKs).
The PI3K–AKT–mTOR axis, which enhances cell survival, protein synthesis, and glucose metabolism.
Both pathways are frequently hyperactivated in cancers with KRAS, EGFR, or other upstream alterations. EGFR (epidermal growth factor receptor), a receptor tyrosine kinase, acts as a key upstream activator of RAS. Ligand binding induces EGFR dimerization and autophosphorylation, creating docking sites for adaptor proteins (e.g., GRB2–SOS) that catalyze GDP-to-GTP exchange on RAS.
In addition to these canonical routes, mutant KRAS can activate NF-κB, a transcription factor that upregulates pro-inflammatory and anti-apoptotic genes such as PTGS2 (encoding cyclooxygenase-2, COX-2). IKBKB (IκB kinase β) is a critical regulator of NF-κB activation and is often co-opted in KRAS-driven tumors to support immune evasion and survival.
Several oncogenic kinases intersect with or amplify KRAS signaling, including:
SRC: a non-receptor tyrosine kinase that modulates integrin and growth factor signaling;
MET: the hepatocyte growth factor receptor, implicated in invasion and resistance;
KIT and ABL1: drivers in gastrointestinal stromal tumors and chronic myeloid leukemia, respectively;
JAK2: a mediator of cytokine receptor signaling that can cross-talk with RAS pathways.
In this context, natural phytochemicals have gained attention as potential anticancer agents because of their multi-targeted actions, relatively low toxicity, and established use in traditional medicine. Tinospora cordifolia (commonly known as Guduchi), an important medicinal plant in Ayurveda, contains berberine, a bioactive isoquinoline alkaloid.7 Berberine is well-documented for its antioxidant, anti-inflammatory, pro-apoptotic, and anti-proliferative effects across a range of cancer models.8 It is also known to influence major oncogenic signaling pathways, including NF-κB, MAPK, and Wnt/β-catenin, all of which are strongly implicated in CRC pathogenesis.9-11
Despite these promising properties, the direct molecular interaction between berberine and KRAS has not been fully explored. To bridge this gap, the present study employs an integrated in silico approach involving molecular docking, ADMET prediction, and structural validation. The findings aim to provide mechanistic insights into berberine’s potential as a KRAS-targeted therapeutic candidate, laying the groundwork for further experimental and clinical investigations in CRC.12,13
Materials and Methods
Collection of berberine targets
The 2D structure of berberine (PubChem CID: 2353) was downloaded in SDF format from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/). Potential targets of berberine were predicted using: Swiss Target Prediction (http://www.swisstargetprediction.ch/ ) Pharm Mapper (http://www.lilab-ecust.cn/pharmmapper/) All predicted targets were standardized and converted to official gene names using the UniProt database (https://www.uniprot.org/).
Collection of colorectal cancer targets
CRC-associated targets were retrieved from the following disease-related databases using the keyword “colorectal cancer”: Gene Cards (https://www.genecards.org/) OMIM (https://www.omim.org/) DisGeNET (https://www.disgenet.org/).In Gene Cards, targets with a relevance score >10 were considered to ensure reliability.
Identification of the common targets:
The overlapping targets between berberine and CRC were identified using an online Venn diagram tool (http://bioinformatics.psb.ugent.be/webtools/Venn/ ). These common targets were considered potential therapeutic targets of berberine against CRC.
Ligand Preparation and Optimization
The active compound selected for the present study was berberine, a prominent isoquinoline alkaloid obtained from Tinospora cordifolia, known for its diverse pharmacological activities, including anticancer effects. The 3D structure of berberine was retrieved from the PubChem database (CID: 2353) in SDF format.
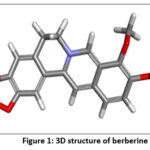 |
Figure 1: 3D structure of berberine
|
Target Protein Selection and Preparation
The protein target selected was KRAS protein complexed with a GDP analogue (PDB ID: 6OIM), a critical oncogenic driver in colorectal carcinoma. The 3D crystallographic structure was downloaded from the RCSB Protein Data Bank.14 Protein preparation involved removal of water molecules, heteroatoms, and co-crystallized ligands using Discovery Studio Visualizer, preparing the structure for molecular docking.
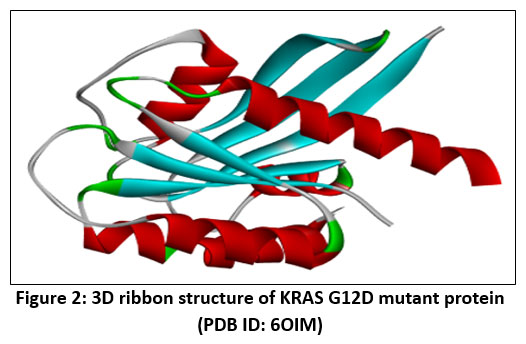 |
Figure 2: 3D ribbon structure of KRAS G12D mutant protein (PDB ID: 6OIM)
|
Structure Validation of Protein
To assess the stereochemical reliability and structural integrity of the selected protein, a Ramachandran plot was generated using the PROCHECK server.14 This analysis examined the phi (ϕ) and psi (ψ) torsional angles of amino acid residues, thereby evaluating the overall backbone geometry. The results confirmed that the majority of residues were located within the most favored regions, which is an essential prerequisite for reliable docking and simulation studies. Such validation ensures that the protein structure is of high quality, minimizing the likelihood of misleading or spurious binding interactions during molecular docking experiments.
Molecular Docking Studies
Docking studies were performed using Auto Dock Vina v1.2.2, integrated with the CB-Dock2 server to identify the optimal binding cavity.15 A grid box was automatically adjusted around predicted pockets, ensuring accurate cavity selection. Binding affinity (ΔG, kcal/mol) was used as the scoring function. To validate the docking protocol, the native co-crystallized ligand was redocked into the active site, and the root mean square deviation (RMSD) was calculated to confirm docking reliability (acceptable RMSD <2.0 Å).
ADMET Prediction
To evaluate the pharmacokinetic characteristics and potential toxicity of berberine, in silico ADMET analyses (Absorption, Distribution, Metabolism, Excretion, and Toxicity) were carried out using two widely recognized web-based tools.
SwissADME was employed to predict key physicochemical properties, compliance with Lipinski’s rule of five, gastrointestinal absorption, and blood–brain barrier permeability.16
pkCSM was applied to estimate water solubility, intestinal absorption, volume of distribution, interactions with cytochrome P450 enzymes, and multiple toxicity endpoints, including Ames mutagenicity, hERG channel inhibition, and hepatotoxicity.17
These computational predictions provide an early assessment of the compound’s drug-likeness and safety profile, offering valuable insights before advancing to in vitro or in vivo validation.
Structural Quality Factor (Z-score)
The Z-factor of the protein-ligand complex was calculated to evaluate the quality of the structural model post-docking. A significantly negative Z-score (typically below -3) indicates a reliable protein structure with non-random, biologically plausible conformations. This was computed using ProSA-web, a robust structure assessment tool based on knowledge-based potentials.
Results
Table 1: Shows the predicted overlapping targets of berberine in colorectal cancer. Key hits included kinases (PI3K family, CDKs, MAPKs), oncogenes (SRC, MET, KIT, ABL1), and signaling regulators (JAK2, IKBKB, PTGS2). These proteins are strongly implicated in CRC progression, highlighting the multi-target nature of berberine.
| PIK3CB
PIK3CG MAPKAPK2 PARP10 CHEK2 CDK8 MAPK10 AGPAT2 ALOX5AP CHEK1 KIT SRC MKNK1 IKBKB MAPK14 LRRK2 TGM2 SCD XBP1 TYMS DHFR ROCK1 ACHE HTR2B BCHE ADRA2C ADRA2B CHRM1 PTGS2 DRD3 CDK2 |
SIRT2
ADORA2A ADORA3 GRM5 QPCT CSF1R CNR2 CBFB PIM2 MET HTR3A GRK5 CYP11B1 IMPDH2 F3 CYP19A1 BCAT2 TRPM8 ICAM1 SELE MAOB PIK3CD CDK4 ROCK2 ATR AOC3 GRK3 PLK1 GRK2 PGR DRD4 |
HSD17B1
PNMT PARP2 SIGMAR1 CYP2D6 RAC1 CDC42 RPS6KB1 AURKA AURKB CYP11B2 PRF1 GRIA1 TBXAS1 HPGD SLC1A3 PIM1 PRKACA CHRM4 JAK2 LCK CDK9 PTPN1 PTGES RPS27 NPY5R MAP4K4 NTRK1 ABL1 NR3C2 |
Molecular Docking
Berberine showed strong binding affinity for the KRAS protein, with a Vina score of –8.5 kcal/mol, indicating a stable and thermodynamically favorable interaction. Docking placed berberine in Cavity C1, a large and accessible pocket (~2114 ų) suitable for ligand binding. Interaction analysis revealed hydrogen bonds with Glu63, Asp57, and Arg68, supported by hydrophobic contacts involving Gly10, Ala11, and Cys12. Aromatic residues such as Tyr96, Pro34, and Phe59 contributed through π–π stacking, while polar residues (Gln61, Gly60, Glu62) helped stabilize the complex. Overall, berberine is well-positioned in the nucleotide-binding region of KRAS, suggesting a potential role in disrupting its GTPase activity, a critical driver of CRC progression.
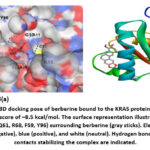 |
Figure 4: (a and b). 3D docking pose of berberine bound to the KRAS protein (Cavity C1, PDB ID: 6OIM) with a Vina score of –8.5 kcal/mol.
|
Table 2: Presents the docking results of berberine with the KRAS protein (PDB ID: 6OIM). Among the predicted binding pockets, Cavity C1 showed the most favorable binding with a Vina score of –8.5 kcal/mol and the largest cavity volume (2114 ų). Other cavities, such as C3 and C4, yielded lower binding scores (–6.7 kcal/mol) with much smaller volumes, while C5 displayed the weakest binding (–5.3 kcal/mol). These findings suggest that Cavity C1 is the most suitable site for berberine accommodation and interaction within KRAS.
| CurPocket ID | Vina Score (kcal/mol) | Cavity Volume (ų) | Center (x, y, z) | Docking Size (x, y, z) |
| C1 | -8.5 | 2114 | 2, -4, 3 | 22, 28, 22 |
| C3 | -6.7 | 143 | 9, 1, -9 | 22, 22, 22 |
| C4 | -6.7 | 140 | 16, 11, -1 | 22, 22, 22 |
| C2 | -6.2 | 267 | 18, -6, 12 | 22, 22, 22 |
| C5 | -5.3 | 81 | 20, -2, 4 | 22, 22, 22 |
Structure validation of protein
Ramachandran Plot Analysis
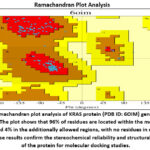 |
Figure 5: Ramachandran plot analysis of KRAS protein (PDB ID: 6OIM) generated using PROCHECK.
|
The stereochemical quality of the KRAS protein (PDB ID: 6OIM) was assessed using PROCHECK. Ramachandran plot analysis showed that 96% of non-glycine and non-proline residues were positioned within the most favored regions, while the remaining 4% occupied additionally allowed regions. Importantly, no residues were found in generously allowed or disallowed regions, confirming the high quality of the model. The protein contained 11 glycine residues, 4 proline residues, and 3 terminal residues (excluding glycine and proline), giving a total of 167 residues.
Further evaluation with G-factor analysis revealed favorable stereochemical scores: phi–psi distribution (0.05), chi1–chi2 (0.10), chi1 only (0.17), bond length (0.61), and bond angle (0.52), all exceeding the acceptable threshold of –0.5. The average G-factor (0.21) confirmed the structural reliability of the model. These results demonstrate that the selected KRAS structure is robust and suitable for docking, ADMET prediction, and simulation studies.
Structural quality factor (Z-score)
The structural reliability of the KRAS protein model was further evaluated using ProSA-web, which yielded a Z-score of –6.9. In the ProSA Z-score plot, the model is represented by a black dot positioned firmly within the distribution of high-quality structures—X-ray resolved proteins denoted in light blue and NMR-derived structures in dark blue. This favorable placement indicates that the overall fold and energetic profile of the KRAS structure are consistent with those of experimentally validated proteins, further supporting its suitability for molecular docking and in silico interaction studies with berberine.
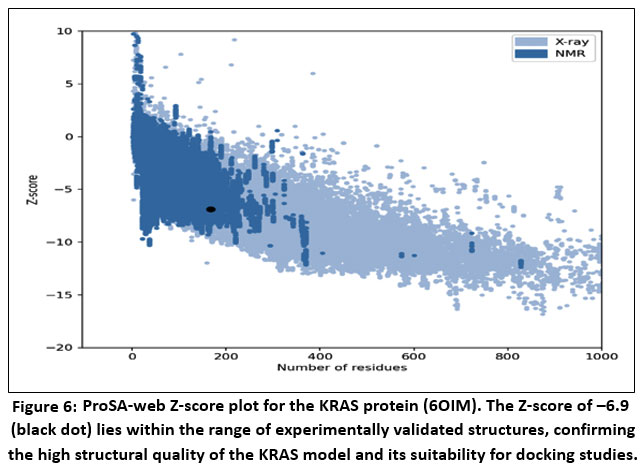 |
Figure 6: ProSA-web Z-score plot for the KRAS protein (6OIM).
|
The Z-score of –6.9 (black dot) lies within the range of experimentally validated structures, confirming the high structural quality of the KRAS model and its suitability for docking studies.
SwissADME Report
Berberine was assessed for its pharmacokinetic properties and drug-likeness using the SwissADME web tool. The results highlight several important features relevant to its therapeutic potential.
Physicochemical Characteristics
Berberine has a molecular weight of 336.36 g/mol and a relatively low topological polar surface area (TPSA) of 40.80 Ų. It contains no hydrogen bond donors, four hydrogen bond acceptors, and only one rotatable bond. Its moderate lipophilicity is indicated by a consensus LogP between 1.5 and 2.1, suggesting a balanced solubility and membrane permeability profile.
Pharmacokinetics
Despite meeting Lipinski’s rule of five, berberine shows poor gastrointestinal (GI) absorption, which may limit its oral bioavailability. It is predicted not to cross the blood-brain barrier (BBB) and functions as a substrate for P-glycoprotein (P-gp), potentially reducing its intracellular accumulation. Additionally, berberine inhibits key metabolic enzymes, including CYP1A2, CYP2C19, and CYP2D6, which could lead to drug-drug interactions.
Drug-Likeness Assessment
Berberine meets the criteria of Lipinski, Veber, and Egan filters but does not satisfy Ghose and Muegge rules, primarily due to low TPSA and borderline molecular characteristics. Its moderate bioavailability score (0.55) indicates that systemic exposure may be limited after oral administration.
Medicinal Chemistry Insights
Analysis revealed one PAINS alert and two Brenk alerts, suggesting potential off-target effects. Nonetheless, berberine has a synthetic accessibility score of 3.61, indicating it is relatively straightforward to synthesize and chemically modify.
While berberine possesses several desirable drug-like attributes, its pharmacokinetic limitations—such as low GI absorption, susceptibility to efflux, and CYP inhibition—could restrict its clinical effectiveness. These findings support the exploration of strategies like structural modification or nano-formulation to improve its bioavailability and enhance its therapeutic potential, especially in cancer treatment.
Table 3: Physicochemical properties and drug likeliness characteristics
| PHYSICOCHEMICAL PROPERTIES | |
| PROPERTY | VALUE |
| Molecular weight | ~336.36 g/mol |
| H-bond Donors | 0 |
| H-bond Acceptors | 4 |
| Topological polar surface area (TPSA) | 40.80 Ų |
| rotatable bonds | 1 |
| LogP (consensus) | ~1.5–2.1 |
| PHARMACOKINETICS | |
| PARAMETER | PREDICTION |
| GI Absorption | Low |
| BBB Permeation | No |
| P-gp Substrate | Yes |
| CYP450 Inhibiton | CYP1A2, CYP2C19, CYP2D6 inhibitor |
| Skin Permeation (logKp) | 6.75 cm/s (poor) |
| DRUG LIKELINESS RULE OF 5 AND OTHERS | |
| Rule/System | Result |
| Lipinski’s Rule | Pass (No violations) |
| Ghose Filter | Fail (Too small TPSA) |
| Veber Rule | Pass |
| Egan Rule | Pass |
| Muegge Rule | Fail |
| Bioavailability Score | 0.55 |
| MEDICINAL CHEMISTRY | |
| Filter/Flag | Result |
| PAINS alert | Yes (1 alert) |
| Brenk alert | Yes (2 alerts) |
| Leadlikeness | Not ideal |
| Synthetic Accessibility (1 = easy, 10 = hard) | 3.61 |
Table 4: ADMET Profile of Berberine Swiss ADME indicated moderate oral bioavailability (GI absorption low, bioavailability score 0.55) and P-glycoprotein substrate status, consistent with known poor solubility of berberine [Liu et al., 2010]. pkCSM, however, predicted high intestinal absorption (97.1%) and acceptable safety (non-hERG inhibitor, moderate hepatotoxicity). These findings reinforce berberine’s potential as a lead compound, although formulation strategies (e.g., nanoparticle or liposomal delivery) may be needed to overcome absorption limitations.
| Category | Parameter | Value | Unit / Notes |
| Absorption | Caco-2 permeability | 1.734 | log Papp (10⁻⁶ cm/s) |
| Intestinal absorption (human) | 97.15 | % Absorbed | |
| Skin permeability | -2.576 | log Kp | |
| P-glycoprotein substrate | Yes | — | |
| P-glycoprotein I inhibitor | No | — | |
| P-glycoprotein II inhibitor | Yes | — | |
| Distribution | VDss (human) | 0.58 | log L/kg |
| Fraction unbound (human) | 0.262 | Fu | |
| BBB permeability | 0.198 | log BB | |
| CNS permeability | -1.543 | log PS | |
| Metabolism | CYP2D6 substrate | No | — |
| CYP3A4 substrate | Yes | — | |
| CYP1A2 inhibitor | Yes | — | |
| CYP2C19 inhibitor | No | — | |
| CYP2C9 inhibitor | No | — | |
| CYP2D6 inhibitor | Yes | — | |
| CYP3A4 inhibitor | Yes | — | |
| Excretion | Total Clearance | 1.27 | log ml/min/kg |
| Renal OCT2 substrate | No | — | |
| Toxicity | AMES toxicity | Yes | — |
| Maximum tolerated dose (human) | 0.144 | log mg/kg/day | |
| hERG I inhibitor | No | — | |
| hERG II inhibitor | No | — | |
| Oral Rat Acute Toxicity (LD50) | 2.571 | mol/kg | |
| Oral Rat Chronic Toxicity (LOAEL) | 1.89 | log mg/kg_bw/day | |
| Hepatotoxicity | Yes | — | |
| Skin Sensitisation | No | — | |
| T. pyriformis toxicity | 0.354 | log μg/L | |
| Minnow toxicity | -0.277 | log mM |
Discussion
Colorectal cancer (CRC) is frequently driven by KRAS mutations, which occur in approximately 35–45% of cases and contribute to resistance against anti-EGFR therapies [Prior et al., 2020; Moore et al., 2020]. Historically considered “undruggable,” KRAS has recently become a tractable target with the approval of covalent KRAS G12C inhibitors, spurring interest in developing broader-spectrum inhibitors.
Although berberine does not bind directly to KRAS, several lines of experimental evidence indicate that it interferes with KRAS signalling through multiple indirect mechanisms. In colorectal cancer cells harbouring KRAS mutations, berberine was shown to reduce both KRAS mRNA and protein expression, thereby attenuating activation of the downstream RAF–MEK–ERK cascade (Li et al., 2020)21. Additionally, berberine disrupts cholesterol-rich lipid rafts in the plasma membrane, which are essential for proper KRAS localization and function (Zhou et al., 2018)24. This membrane-level interference may compromise KRAS nanoclustering and signal transduction. Further, berberine suppresses upstream activators such as EGFR (Peng et al., 2015)22 and activates AMPK, leading to inhibition of mTOR and c-MYC—key effectors in KRAS-driven metabolic reprogramming Wang et al.18 Collectively, these actions position berberine as a modulator of the KRAS signaling network rather than a direct inhibitor.
Despite the identification of numerous mutation sites and varying frequencies across different cancer types, KRAS mutations predominantly (about 80%) show a preference for codon 12, with G12D, G12V, and G12C being the most prevalent variants in solid tumors. In colorectal cancer, KRAS G12C accounts for approximately 3–4% of cases and has historically been associated with resistance to anti-EGFR therapies. However, the covalent KRASG12C inhibitor adagrasib has demonstrated promising clinical activity in this subset. In the KRYSTAL-1 trial, adagrasib monotherapy achieved an objective response rate of 34% and a disease control rate of 84% in heavily pretreated KRASG12C-mutant CRC patients, with even greater efficacy observed in combination with cetuximab (response rate: 46%) [last two]. These results underscore the therapeutic potential of allele-specific inhibition, though responses remain limited to the G12C variant—highlighting the continued need for strategies targeting non-G12C KRAS mutants, which constitute the majority of KRAS-driven CRC.
In this study, berberine, an alkaloid derived from Tinospora cordifolia, demonstrated strong binding affinity to KRAS (–8.5 kcal/mol), forming hydrogen bonds with E63, D57, and R68, alongside π–π interactions with Y96 and F59. These residues are located within the nucleotide-binding region, indicating that berberine may disrupt GTP/GDP binding and impair KRAS-mediated signaling, consistent with prior observations of allosteric KRAS inhibition.
From a pharmacokinetic and safety standpoint, drug interactions and off-target effects are critical considerations. Cytochrome P450 enzymes—particularly CYP1A2, CYP2C19, and CYP2D6—metabolize a wide range of therapeutics. Inhibition or induction of these isoforms can alter drug exposure and lead to toxicity or reduced efficacy.19 Additionally, unintended blockade of the hERG potassium channel (encoded by KCNH2) is a common cause of drug-induced QT prolongation and cardiac arrhythmia, necessitating early screening in compound development.
Tools such as pkCSM (a computational platform for predicting pharmacokinetic and toxicity endpoints) are increasingly used to assess these properties in silico, including hERG inhibition, CYP interactions, and Ames mutagenicity, thereby guiding lead optimization before costly in vivo studies.
Pharmacokinetic analysis indicated moderate oral bioavailability and high intestinal absorption (97.1%) and acceptable safety (non-hERG inhibitor, moderate hepatotoxicity). In clinical pharmacology, a label of “moderate hepatotoxicity” generally corresponds to CTCAE Grade 2 liver injury, typically defined as ALT/AST elevations of >3–5 × ULN or bilirubin 1.5–3 × ULN, which usually requires dose interruption or close monitoring rather than hospitalization. Such moderate enzyme elevations are clinically relevant, especially when considering co-administration with anticancer agents that have known hepatic liabilities. Notably, adagrasib, a KRAS^G12C^ inhibitor, has been associated with ALT/AST elevations and occasional clinically significant hepatotoxicity in the KRYSTAL-1 trial, necessitating treatment modification in some patients.28 Thus, any compound predicted to induce hepatotoxicity warrants careful evaluation when used alongside KRAS-targeted therapies.
Reduced berberine derivatives such as dihydroberberine (DHBER) and tetrahydroberberine (THBER) exhibit different metabolic and toxicological profiles compared to berberine. DHBER demonstrates higher bioavailability and favorable preclinical toxicology, including lack of mutagenicity in standard assays and high LD₅₀ values, suggesting potentially lower hepatotoxic burden at equivalent systemic exposures.29 THBER and related hydrogenated berberine analogues have shown hepatoprotective or neutral effects in certain animal models, although isolated reports indicate variable hepatic outcomes depending on dose and structural modifications.30 Because these reduced derivatives undergo different metabolic pathways and may avoid formation of reactive metabolites implicated in berberine-associated hepatotoxicity, they may theoretically exhibit lower hepatotoxic risk, but this requires direct comparative in vivo studies.
Given these considerations, the pkCSM-predicted hepatotoxicity for berberine highlights the need for liver-function monitoring in future preclinical or clinical development, and also supports exploring DHBER/THBER as alternative scaffolds that may retain anticancer activity with potentially improved hepatic safety. However, limitations such as P-glycoprotein-mediated efflux and CYP450 inhibition could reduce systemic exposure, reflecting berberine’s known bioavailability challenges. These issues may be addressed through nanoparticle encapsulation, liposomal delivery, or prodrug strategies, which have previously enhanced berberine’s bioavailability. Notably, berberine exhibits polypharmacology, modulating multiple oncogenic pathways including NF-κB, Wnt/β-catenin, and MAPK. This suggests its potential both as a direct KRAS inhibitor and as a synergistic agent in CRC therapy. It is instructive to compare berberine’s profile with that of recently approved KRASG12C inhibitors such as sotorasib and adagrasib. Unlike these agents, which covalently and selectively target the KRASG12C mutant protein, berberine lacks mutation specificity and operates through pleiotropic, network-level modulation. Consequently, its antitumor potency is lower, and it is not suitable as a monotherapy for advanced KRAS-mutant cancers.26,27 However, its low cost, oral bioavailability, and minimal severe toxicity may support its use in preventive settings or as an adjunct to enhance efficacy and delay resistance in combination regimens. For instance, by dampening EGFR feedback activation or metabolic adaptation—common resistance mechanisms to direct KRAS inhibitors—berberine could theoretically complement targeted therapies Canon et al.25
Limitations: This study is limited to computational predictions. Experimental validation through in vitro KRAS-GTPase inhibition assays, CRC cell viability studies, and in vivo evaluations is required.
Future Perspectives
Conduct molecular dynamics simulations to assess the stability of the KRAS–berberine complex.
Perform experimental KRAS inhibition assays.
Investigate formulation strategies to improve pharmacokinetics.
Design berberine analogues with enhanced potency and specificity.
Collectively, these findings position berberine as a promising lead scaffold for KRAS-targeted therapies in colorectal cancer.
Conclusion
This study presents the first comprehensive in silico evidence supporting berberine, derived from Tinospora cordifolia, as a potential KRAS inhibitor in colorectal cancer. Berberine exhibited a favorable docking pose within the KRAS active site, forming key interactions with residues essential for nucleotide binding. Structural validation using Ramachandran analysis and ProSA Z-scores confirmed the reliability of the docking model, while SwissADME and pkCSM profiling indicated acceptable drug-likeness and safety. Although limitations related to absorption and metabolism were noted, these may be overcome through advanced formulation approaches, such as nanoparticles or liposomes, and structure–activity optimization. Overall, berberine emerges as a promising lead candidate for KRAS-targeted therapy, with potential for development into a novel treatment strategy for KRAS-mutant colorectal cancer. Beyond protein-coding mutations, recent studies have uncovered the presence of G-quadruplex (G4) DNA structures in the promoter region of the KRAS gene—referred to as RAS G4. These non-canonical secondary structures form in guanine-rich sequences and can modulate transcriptional activity. Stabilization of the KRAS G4 by small molecules (e.g., pyridostatin, CX-5461) has been shown to suppress KRAS expression at the mRNA level in preclinical models, offering a mutation-agnostic strategy to target KRAS-driven cancers. Although still in early development, G4-targeting approaches represent a promising complementary avenue to direct KRAS inhibitors, particularly for non-G12C variants that remain undruggable by current covalent agents. Future work should explore whether natural compounds like berberine—known to interact with nucleic acids—might also influence RAS G4 stability, potentially contributing to their observed downregulation of KRAS expression.
Acknowledgement
I sincerely thank Dr.Y.Srinivas Rao, Vignan Institute of Pharmaceutical technology, for her valuable guidance and constant support throughout this study. I also appreciate the resources and facilities provided by Vignan College, which enabled the successful completion of this work. Special thanks to the developers of the bioinformatics tools and databases used in this study, including Auto Dock Vina, Swiss ADME, pkCSM, and the Protein Data Bank, for facilitating accurate and efficient in silico analyses.
Funding Sources
The author(s) received no financial support for the research, authorship, and/or publication of this article
Conflict of Interest
We declare that this research have no commercial or financial relationships that could be perceived as a potential conflict of interest in this study.
Data Availability Statement
The manuscript incorporates all datasets produced or examined throughout this research study.
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Permission to Reproduce Material from Other Sources
Not Applicable
Author Contributions
Naga Bharathi Marni:
Jashnavi Naga Sravya Singuluri: Conceptualization, Methodology, Writing – Original Draft.
Jhansi Rama Lakshmi Saragam: Data Collection, Analysis, Writing – Review & Editing.
Usha Rani Sanapala: Visualization.
References
- Mármol I, Sánchez-de-Diego C, Pradilla Dieste A, Cerrada E, Rodriguez Yoldi MJ. Colorectal carcinoma: a general overview and future perspectives. Int J Mol Sci. 2017;18(1):197. doi:10.3390/ijms18010197
CrossRef - Sung H, Ferlay J, Siegel RL, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71(3):209-249. doi:10.3322/caac.21660
CrossRef - Prior IA, Hood FE, Hartley JL. The frequency of Ras mutations in cancer. Cancer Res. 2020;80(14):2969-2974. doi:10.1158/0008-5472.CAN-19-3682
CrossRef - Moore AR, Rosenberg SC, McCormick F, Malek S. RAS-targeted therapies: is the undruggable drugged? Nat Rev Drug Discov. 2020;19(8):533-552. doi:10.1038/s41573-020-0068-6
CrossRef - Dienstmann R, Salazar R, Tabernero J. The evolving role of molecular markers in the treatment of colorectal cancer. Nat Rev Clin Oncol. 2017;14(6):337-350. doi:10.1038/nrclinonc.2016.167
CrossRef - Canon J, Rex K, Saiki AY, et al. The clinical KRAS(G12C) inhibitor AMG 510 drives anti-tumour immunity. 2019;575(7781):217-223. doi:10.1038/s41586-019-1694-1
CrossRef - Tillhon M, Ortiz LM, Lombardi P, Scovassi AI. Berberine: new perspectives for old remedies. Biochem Pharmacol. 2012;84(10):1260-1267. doi:10.1016/j.bcp.2012.07.018
CrossRef - Chen H, Ye C, Cai B, Zhang F, Wang X, Zhang J, Zhang Z, Guo Y, Yao Q. Berberine inhibits intestinal carcinogenesis by suppressing intestinal pro-inflammatory genes and oncogenic factors through modulating gut microbiota. BMC Cancer. 2022;22(1):566. doi:10.1186/s12885-022-09635-9
CrossRef - Wu K, Yang Q, Mu Y, Zhou L, Liu Y, Zhou Q, He B. Berberine inhibits the proliferation of colon cancer cells by inactivating Wnt/β-catenin signaling. Int J Oncol. 2012;41(1):292-298. doi:10.3892/ijo.2012.1423
CrossRef - Patil JB, Kim J, Jayaprakasha GK. Berberine induces apoptosis in breast cancer cells (MCF-7) through mitochondrial-dependent pathway. Eur J Pharmacol. 2010;645(1-3):70-78. doi:10.1016/j.ejphar.2010.07.037
CrossRef - Sharma S, Pathak S. Berberine: chemopreventive potential and mechanisms of action against colorectal cancer. 2022;196:64-76. doi:10.1016/j.biochi.2022.02.015
- Ostrem JM, Peters U, Sos ML, Wells JA, Shokat KM. K-Ras(G12C) inhibitors allosterically control GTP affinity and effector interactions. 2013;503(7477):548-551. doi:10.1038/nature12796
CrossRef - Gupta SC, Kim JH, Prasad S, Aggarwal BB. Regulation of survival, proliferation, invasion, angiogenesis, and metastasis of tumor cells through modulation of inflammatory pathways by nutraceuticals. Cancer Metastasis Rev. 2010;29(3):405-434. doi:10.1007/s10555-010-9235-2
CrossRef - Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE. The Protein Data Bank. Nucleic Acids Res. 2000;28:235-242. doi:10.1093/nar/28.1.235
CrossRef - Trott O, Olson AJ. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem. 2010;31(2):455-461. doi:10.1002/jcc.21334
CrossRef - Daina A, Michielin O, Zoete V. SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness, and medicinal chemistry friendliness of small molecules. Sci Rep. 2017;7:42717. doi:10.1038/srep42717
CrossRef - Pires DE, Blundell TL, Ascher DB. pkCSM: predicting small-molecule pharmacokinetic and toxicity properties using graph-based signatures. J Med Chem. 2015;58(9):4066-4072. doi:10.1021/jm50178a008
CrossRef - Chen, Y., Yu, Z., Zhao, L., Liu, J., & Wang, L. (2022). Berberine in cancer therapy: Updates on its molecular mechanisms and therapeutic potential. Pharmacological Research, 175, 105999. https://doi.org/10.1016/j.phrs.2021.105999
- Han, Y., Yang, S., He, S., Zhou, H., & Liu, Z. (2011). Berberine inhibits human cytochrome P450 2D6, 2C9, and 3A4 activity in vitro. Drug Metabolism and Disposition, 39(10), 1808–1811. https://doi.org/10.1124/dmd.111.040212
CrossRef - Kong, W. J., Zhu, H. B., Yang, J. L., Liu, Q. F., Zhang, L. J., Liu, J. B., Li, Y. H., & Xiao, X. H. (2013). Berberine is a novel cholesterol-lowering drug working through a unique mechanism distinct from statins. Reproductive Toxicology, 42, 128–135. https://doi.org/10.1016/j.reprotox.2013.08.004
CrossRef - Li, Y., Zhang, Y., Wang, L., Liu, J., & Liu, G. (2020). Berberine suppresses colorectal cancer through downregulation of KRAS and inhibition of the RAS/RAF/MEK/ERK signaling pathway. European Journal of Pharmacology, 886, 173502. https://doi.org/10.1016/j.ejphar.2020.173502
CrossRef - Peng, C., Zhao, S., He, W., Chen, Y., & Wang, Y. (2015). Berberine inhibits the proliferation of human colorectal cancer cells by suppressing EGFR-mediated signaling pathways. Oncology Reports, 34(5), 2527–2534. https://doi.org/10.3892/or.2015.4247
CrossRef - Wang, Y., Zhao, Q., Han, Y., Wang, L., & Wang, T. (2021). Berberine induces apoptosis and inhibits glycolysis in KRAS-mutant colorectal cancer via AMPK/mTOR/c-MYC signaling. Cell Death & Disease, 12(3), 298. https://doi.org/10.1038/s41419-021-03584-7
- Zhou, Y., Wang, F., Liu, X., Wu, B., & Liu, J. (2018). Berberine disrupts lipid rafts and inhibits RAS membrane localization in colon cancer cells. Biochemical Pharmacology, 155, 225–234. https://doi.org/10.1016/j.bcp.2018.07.008
CrossRef - Canon, J., Rex, K., Saiki, A. Y., Mohr, C., Cooke, K., Bagal, D.,& Lipford, J. R. (2019). The clinical KRAS(G12C) inhibitor AMG 510 drives anti-tumour immunity. Nature, 575(7781), 217–223.
CrossRef - Strickler, J. H., et al. (2022). Adagrasib in KRASG12C-mutated colorectal cancer. New England Journal of Medicine, 387(1), 44–54.
- Yaeger, R., et al. (2023). KRYSTAL-1: Activity and safety of adagrasib with or without cetuximab in KRASG12C-mutant colorectal cancer. Journal of Clinical Oncology, 41(4), 755–765.
- Fakih MG, Desai J, Kuboki Y, et al. Sotorasib or adagrasib in KRAS G12C–mutated colorectal cancer: results from the KRYSTAL-1 study. N Engl J Med. 2023;389.
- Li Y, Zhao J, Wang C, et al. Pharmacological advances of dihydroberberine: a metabolite of berberine. Pharmacol Res. 2020; 158:104867.
- Amin F. Majdalawieh, Sarah M. Yousef, Imad A. Abu-Yousef, Gheyath K. Nasrallah. Immunomodulatory and Anti-Inflammatory Effects of Berberine in Lung Tissue and its Potential Application in Prophylaxis and Treatment of COVID-19. Biosci. (Landmark Ed) 2022, 27(5), 166.
CrossRef
Abbreviations List
ADMET: Absorption, Distribution, Metabolism, Excretion, and Toxicity
CRC: Colorectal cancer
MAPK: Mitogen-activated protein kinase
ERK: Extracellular signal-regulated kinase
PI3K: Phosphatidylinositol 3-kinase
AKT: Ak strain transforming (also known as Protein Kinase B, or PKB)
GTP: Guanosine triphosphate
GDP: Guanosine diphosphate
NF-κB: Nuclear factor kappa-light-chain-enhancer of activated B cells
CDKs: Cyclin-dependent kinases
KRAS: Kirsten rat sarcoma viral oncogene homolog
EGFR: Epidermal growth factor receptor
hERG: Human Ether-à-go-go-Related Gene (often refers to the gene KCNH2)
SRC: Rous sarcoma virus oncogene, Src family tyrosine kinase
MET: Mesenchymal-epithelial transition factor (or c-Met)
KIT: Tyrosine-protein kinase KIT (or c-Kit)
ABL1: Abelson murine leukemia viral oncogene homolog 1
JAK2: Janus kinase 2
IKBKB: Inhibitor of nuclear factor kappa-B kinase subunit beta
PTGS2: Prostaglandin-endoperoxide synthase 2 (also known as COX-2)
CYP1A2: Cytochrome P450 1A2 (a liver enzyme)
CYP2C19: Cytochrome P450 2C19 (a liver enzyme)
CYP2D6: Cytochrome P450 2D6 (a liver enzyme)
Other Terms
pkCSM: A web-based tool for predicting small molecule pharmacokinetic properties (no standard biological abbreviation)
Accepted on: 11-12-2025
Second Review by: Dr. Sunil Chaudhry
Final Approval by: Dr. Wagih Ghannam