Study of Beta-Casein Gene Polymorphism in Gir Cattle from Gujarat State
1Department of Genetics, Ashok and Rita Patel Institute of Integrated Study and Research in Biotechnology and Allied Science (ARIBAS), The Charutar Vidhya Mandal (CVM) University, Vallabh Vidyanagar, Anand, Gujarat.
2Department of Biotechnology, Genetics and Bioinformatics, Natubhai V. Patel College of Pure and Applied Sciences (NVPAS), The Charutar Vidhya Mandal (CVM) University Vallabh Vidhyanagar, Anand, Gujarat.
Corresponding Author E-mail:anjali.kulkarni@cvmu.edu.in
DOI : http://dx.doi.org/10.13005/bbra/3461
ABSTRACT:Bovine milk and dairy products are vital components of traditional diets globally, with milk proteins comprising approximately 80% casein and 20% whey. Among the caseins, β-casein is a major hydrophobic protein with two common genetic variants, A1 and A2. The difference between these variants lies in a single amino acid substitution at position 67, where histidine (CCT) in A1 is replaced by proline (CAT) in A2. This study aimed to investigate the β-casein gene polymorphism in 200 Gir cattle from Shri Krishna Farm, Savarkundla using the PCR-RFLP technique. Genomic DNA was extracted using the Phenol-Chloroform-Isoamyl Alcohol method, and its quality and quantity were assessed through agarose gel electrophoresis and Nanodrop spectrophotometry. PCR amplification yielded a 121 bp fragment, which was then subjected to restriction digestion using the DdeI enzyme. The analysis revealed two genotypes—A1A2 and A2A2—with respective genotypic frequencies of 0.125 and 0.875, while the A1A1 genotype was not observed. The allele frequencies were 0.06 for A1 and 0.94 for A2, confirming the predominance of the A2 allele in Gir cattle and suggesting their potential for producing A2 milk, which is considered healthier.
KEYWORDS:Allele Frequency; Beta-Casein; Genotype; Milk; PCR-RFLP
Introduction
Bovine milk and its derivatives are integral to diets worldwide, with the casein protein family (α, β, κ) constituting approximately 80% of milk proteins. Among these, β-casein accounts for about 30–35% of total casein and is notably more water-soluble, particularly at lower temperatures, compared to other caseins. A single nucleotide polymorphism (SNP) in exon 7 of the CSN2 gene CCT → CAT results in a histidine-to-proline substitution, rather than proline-to-histidine, giving rise to the two primary β-casein alleles: A2 (Pro⁶⁷) and A1 (His⁶⁷). The A1 variant has been associated with enzymatic cleavage during digestion, yielding β-casomorphin-7 (BCM-7), an opioid peptide that may influence gut inflammation, glucose metabolism, and cardiovascular risk, whereas the A2 variant is considered safer and has been linked to better digestibility, lower inflammatory response, and beneficial lipid profiles (Mukesh et al., 2022; Cartuche-Macas et al., 2025; Sawicka-Zugaj et al., 2025).
In India, indigenous zebu (Bos indicus) breeds predominantly carry the A2 allelemaking them promising candidates for health-oriented dairy breeding programs. For example, Gir, Sahiwal, Tharparkar, and Rathi cattle often exhibit near fixation for the A2 allele or very low A1 frequencies (Yadav et al., 2023; Paradkar et al., 2023). However, crossbreeding with taurine or exotic lines such as Holstein Friesian and Jersey tends to introduce higher frequencies of A1 alleles, as observed in Frieswal and HF crosses (Kumar et al., 2019; Adesh et al., 2019). Therefore, molecular genotyping of β-casein is essential for marker-assisted selection (MAS) aimed at expanding A2 allele presence while controlling A1 introgression.
Gir cattle, indigenous to the Gir forest region in Gujarat, are renowned for their adaptability, disease resistance, and relatively high milk fat content, making them strategically important for A2 milk production. In this study, we analyzed the A1/A2 allele and genotype frequencies in a Gir cattle herd (n = 200) using PCR–RFLP, validated Hardy–Weinberg equilibrium, and compared the frequencies with crossbred Holstein Friesian populations. Our findings confirmed a high prevalence of the A2 allele in Gir cattle, consistent with previous reports, and emphasize the need for conservation and selective breeding strategies to maintain and enhance A2 allele-rich herds in India (Mukesh et al., 2022; Yadav et al., 2023; Paradkar et al., 2023; Cartuche-Macas et al., 2025).
Materials and Methods
Experimental materials
A total of 200 blood samples were collected from Gir cattle (Bos indicus) maintained at Shree Krishna Dairy Farm, Savarkundla, Amreli district, Gujarat, India. The Gir breed, native to the Saurashtra region, is renowned for its high milk yield and disease resistance, making it one of the most important indigenous breeds for A2 milk production (Singh et al., 2020). Blood samples (3 mL each) were collected aseptically from the jugular vein using sterile 5 mL vacutainer tubes containing anticoagulant (heparin). All animals were apparently healthy, unrelated, and selected randomly to avoid sampling bias. Samples were transported on ice and stored at –20°C until further processing (Kumar et al., 2018; Yadav et al., 2021).
Genomic DNA Isolation
Genomic DNA was extracted using the phenol–chloroform–isoamyl alcohol (PCI) method, following the protocol of Patel et al. (2017) with slight modifications. Briefly, 3 mL of heparinised blood was mixed with an equal volume of Solution I (10 mM Tris-HCl, 10 mM KCl, 10 mM MgCl₂, and 20 µL Nonidet P40), vortexed, and incubated at –20°C for 15 minutes to lyse red blood cells. The lysate was centrifuged at 10,000 rpm for 10 min at 4°C to pellet nuclei. The pellet was resuspended in 800 µL Solution II (10 mM Tris-HCl, 10 mM KCl, 10 mM MgCl₂, 0.5 M NaCl, 2 mM EDTA, and 0.5% SDS) along with 5 µL Proteinase K (20 mg/mL) and incubated at 60°C overnight for complete protein digestion.
The lysate was treated with 500 µL equilibrated phenol followed by 200 µL PCI mixture (25:24:1) to remove proteins. DNA was precipitated with 2 volumes of chilled ethanol and washed with 70% ethanol to remove impurities. The DNA pellet was air-dried and resuspended in nuclease-free water.
The quality of the extracted DNA was verified by 1% agarose gel electrophoresis, while purity and concentration were determined using a NanoDrop 2000 spectrophotometer (Thermo Fisher Scientific, USA) by measuring the A260/A280 ratio. Ratios between 1.7–2.0 were considered acceptable (Khan et al., 2019).
Polymerase Chain Reaction (PCR)
The β-casein gene (CSN2) was targeted for genotyping of A1/A2 alleles. A pair of bovine-specific oligonucleotide primers reported by McLachlan et al. (2006) was used to amplify a 121 bp fragment of exon 7 of the CSN2 gene.
Table 1: Primer detail:
| Primer | Sequence (5′ → 3′) | Product Size |
| Forward | CCTTCTTTCCAGGATGAACTCCAG | 121 bp |
| Reverse | GAGTAAGAGGAGGGATGTTTTGTGGGAGGCTCT |
PCR amplification was carried out in a 25 µL reaction volume containing 100 ng genomic DNA, 10 pmol of each primer, 12.5 µL of 2× PCR Master Mix (Thermo Scientific), and nuclease-free water. The amplification was performed in a Veriti™ Thermal Cycler (Applied Biosystems, USA) using the following conditions:
Table 2: Thermal cycler conditons:
| Step | Temperature | Time | Cycles |
| Initial Denaturation | 94°C | 3 min | 1 |
| Denaturation | 94°C | 30 sec | 30Cycles |
| Annealing | 60°C | 30 sec | |
| Extension | 72°C | 30 sec | |
| Final Extension | 72°C | 5 min | 1 |
| Hold | 4°C | ∞ | — |
Amplified PCR products were confirmed by electrophoresis on 2% agarose gel stained with ethidium bromide and visualized under a UV transilluminator (Gel Doc™, Bio-Rad).
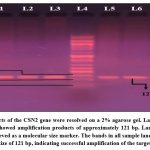 |
Figure 1: PCR products of the CSN2 gene were resolved on a 2% agarose gel. Lanes 1–3 (S37–S39) and lanes 5–8 (S40–S43) showed amplification products of approximately 121 bp. |
Restriction Fragment Length Polymorphism (PCR-RFLP) and Electrophoresis
For genotyping of A1/A2 alleles, 10 µL PCR products were digested with 4 units of DdeI restriction enzyme (New England Biolabs, USA) in a final volume of 20 µL. The digestion reaction was carried out overnight at 37°C in a water bath.
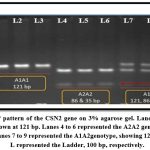 |
Figure 2: PCR-RFLP pattern of the CSN2 gene on 3% agarose gel. Lanes 1-3 represented the A1A1 genotype, as shown at 121 bp. |
The digested products were separated on a 3% agarose gel containing 1× TBE buffer at 120 V for 60–70 minutes. A 50 bp DNA ladder (Thermo Scientific) was used as a molecular size marker. The gel was visualized under a UV transilluminator and photographed for further analysis.
Gene and Genotype Frequency Calculation :
Allele frequencies were calculated using the direct counting method, following the principles of Hardy-Weinberg equilibrium (HWE) (Hardy, 1908). The Chi-square test was applied to evaluate deviations between observed and expected genotype frequencies. The allele frequencies were determined using the formulas Genotype frequency for A1A2, A1A1 and A2A2 genotype for beta casein gene was estimated as per the equation 1. Allelic frequency of A1 and A2 alleles for beta casein (CSN2) gene gene was estimated as per the equation
Equation 1: Genotype Frequency = 𝑇𝑜𝑡𝑎𝑙 𝑛𝑢𝑚𝑏𝑒𝑟 𝑜𝑓 𝑜𝑏𝑠𝑒𝑟𝑣𝑒𝑑 𝑔𝑒𝑛𝑜𝑡𝑦𝑝𝑒𝑠/ 𝑇𝑜𝑡𝑎𝑙 𝑛𝑢𝑚𝑏𝑒𝑟 𝑜𝑓 𝑔𝑒𝑛𝑜𝑡𝑦𝑝𝑒𝑠 (Table 3).
Equation 2: Allelic Frequency Equation Allelic frequency of A1 allele = 2𝐹(𝐴1𝐴1) + 𝐹(𝐴1𝐴2))/ 2(𝑇𝑜𝑡𝑎𝑙 𝑛𝑜 𝑜𝑓 𝑎𝑙𝑙𝑒𝑙𝑒𝑠)(Table 4).
Allelic frequency of A2 allele = 2𝐹(𝐴2𝐴2) + 𝐹(𝐴1𝐴2))/ 2(𝑇𝑜𝑡𝑎𝑙 𝑛𝑜 𝑜𝑓 𝑎𝑙𝑙𝑒𝑙𝑒𝑠)
where A1A1, A1A2, and A2A2 represent genotype counts, and N is the total sample size. This analysis helps assess the genetic stability of the population and the influence of evolutionary forces on genetic variation (Oliveira et al., 2021).
Table 3: Genotypic frequency of A1 and A2 allele of CSN2 gene
| Total no. of animals | Genotype frequency | Total | ||
| A1A1 | A2A2 | A1A2 | ||
| 200 | 00 | 0.125 | 0.875 | 1.0 |
Table 4: Allelic frequency of A1 and A2 allele of CSN2 gene
| Allelic frequency | p-value | |
| A1 | A2 | |
| 0.06 | 0.94 | <0.0001(P<0.05) |
(p<0.05 significant)
Results and Discussion
Results
Genomic DNA extracted from 200 Gir cattle was evaluated for quality and quantity. Electrophoretic assessment on a 1% agarose gel revealed high molecular weight, sharp DNA bands for all samples, indicating excellent integrity suitable for downstream molecular analyses such as PCR and RFLP. DNA purity, measured using a Nanodrop spectrophotometer (Nanodrop 1000, Thermo Scientific, USA), showed A260/A280 ratios ranging from 1.8 to 2.0, confirming acceptable purity for molecular applications. DNA concentrations varied between 30–250 ng/μL, providing sufficient template for PCR amplification.
PCR amplification of the β-casein gene produced the expected 121 bp fragment in all 200 samples, as confirmed on a 2% agarose gel. These PCR products were then digested with the DdeI restriction enzyme to identify A1/A2 polymorphisms.
Discussion
The predominance of the A2 allele observed in this study (A2 = 0.94; A1 = 0.06) aligns with findings from Mukesh et al. (2022), who reported an A2 allele frequency of 0.95 in indigenous breeds such as Sahiwal, Tharparkar, and Gir. Similarly, Yadav et al. (2023) documented complete fixation of the A2 allele (frequency = 1.0) in Gir cattle. The slight variation observed in the current study may be attributed to regional or herd-level differences, smaller sample sizes, or introgression from crossbreeding practices.
Pandey et al. (2019) observed an A2 allele frequency of 0.85 in Sahiwal cattle, while significantly higher A1 allele frequency (0.32) was reported in Holstein Friesian crossbreds. This aligns with the present study’s findings, where HF crossbreds showed A1 = 0.29 and A2 = 0.71, confirming that exotic germplasm contributes to increased A1 allele incidence.
At the global level, Karthickeyan et al. (2020) reported high A2 allele frequencies in Ongole and Deoni breeds, whereas Kaminski et al. (2007) reported elevated A1 allele prevalence in European dairy breeds, especially Holstein Friesians. These results highlight a general trend where indigenous breeds conserve the A2 allele, while European breeds selected for higher milk yield tend to carry more A1 alleles.
From a nutritional perspective, the predominance of the A2 allele is significant. Previous studies (Jianqin et al., 2016) have suggested that A2 milk is easier to digest and may reduce gastrointestinal discomfort, in addition to potentially lowering risks of type 1 diabetes, ischemic heart disease, and autism. Therefore, maintaining high A2 allele frequencies in indigenous breeds not only supports genetic diversity but also has public health benefits.
The absence of the A1A1 genotype in Gir cattle further underscores the breed’s suitability for selective breeding programs aimed at expanding A2 milk production. Similar observations were made by Joshi et al. (2021) in Red Sindhi and Rathi cattle, reinforcing the idea that most Indian zebu breeds are naturally predisposed toward A2 milk production. However, the presence of A1A2 heterozygotes (12.5%) indicates a risk of A1 allele propagation if crossbreeding with exotic breeds is not carefully managed.
Overall, this study supports the hypothesis that zebu cattle act as reservoirs of the A2 allele. Conservation of indigenous breeds such as Gir, Sahiwal, and Tharparkar, along with well-structured breeding policies that limit introgression of A1 alleles from exotic germplasm, will be crucial for ensuring sustainable A2 milk production in India and beyond.
Conclusion
This study establishes the predominance of the A2 allele in Gir cattle (A2 = 0.94; A1 = 0.06), confirming their genetic potential as a sustainable source of A2 milk. The findings, statistically significant (p < 0.05), highlight the role of indigenous zebu cattle in safeguarding A2 milk production, which is associated with better digestibility and reduced risk of certain health disorders compared to A1 milk. Conservation and promotion of A2 allele-rich breeds like Gir, Sahiwal, and Tharparkar will be crucial for enhancing human health, nutritional security, and long-term dairy sustainability.
Acknowledgment
The authors express their sincere gratitude to the Head of ARIBAS, Vallabh Vidyanagar, and CVM University for providing the necessary infrastructure and academic environment to conduct this research. We are deeply thankful to the owners of Krishna Gowshala and Shivaji Gowshala (Savarkundla, Gujarat) for their cooperation during sample collection. We also acknowledge the valuable veterinary assistance provided by Dr. Shubhum Joshi and Dr. Chirag Kanpariya during the fieldwork.
Funding Sources
This research was financially supported by the SHODH – Scheme of Developing High-Quality Research, Education Department, Government of Gujarat. Grant no.202210821
Conflict of Interest
The authors do not have any conflict of interest.
Data Availability Statement
This statement does not apply to this article.
Ethics Statement
All procedures involving animals were conducted in accordance with the guidelines set forth by the Canadian Association for Laboratory Animal Science (CALAS) and the Canadian Council on Animal Care (CCAC). Blood samples were collected by licensed veterinary doctors to ensure ethical and professional standards of animal care and use.” Information regarding cattle was obtained from records maintained by the farm owners.
Author Contributions
Shailja Jitendrabhai Vyas – Methodology; software; writing – review and editing
Anjali Kulkarni – Validation, Investigation original draff.
References
- Adesh S, Kumar R, Singh P. Genetic variability of A1/A2 β-casein in crossbred cattle populations. Indian J Anim Sci. 2019;89(4):512-518. doi:10.56093/ijas.v89i4.91234.
CrossRef - Cartuche-Macas L, Paredes M, Suárez C. Differential A1/A2 β-casein (CSN2) gene-derived allelic frequencies in Ecuadorian cattle breeds. Front Vet Sci. 2025;12:1616426. doi:10.3389/fvets.2025.1616426
CrossRef - Jaiswal KP, De S, Sarsavan A. Review on bovine beta-casein (A1, A2) gene polymorphism and their potentially hazardous effects on human health. Int J Environ Anim Conserv. 2014;3(1):1-12.
- Jianqin L, et al. Effects of milk containing only A2 beta-casein versus milk containing both A1 and A2 beta-casein proteins on gastrointestinal physiology, symptoms of discomfort, and cognitive behavior of people with self-reported milk intolerance. Nutr J. 2016;15(1):1-10. doi:10.1186/s12937-016-0147-z
CrossRef - Joshi CG, et al. Polymorphism of beta-casein gene and its association with milk production traits in Sahiwal and Holstein Friesian crossbred cattle. Indian J Anim Sci. 2021;91(6):456-460.
- Kaminski S, Cieslinska A, Kostyra E. Polymorphism of bovine β-casein and its potential effect on human health. J Appl Genet. 2007;48(3):189-198.
CrossRef - Kaminski S, et al. Polymorphism of bovine beta-casein and its potential effect on human health. J Dairy Sci. 2007;90(6):2884-2895. doi:10.3168/jds.2006-853.
- Karthickeyan SMK, Kathiravan P, Saravanan R, et al. Genetic characterization of β-casein gene variants in Indian native cattle breeds. Anim Biotechnol. 2020;31(6):498-505.
- Khan R, Sharma N, Khan F, et al. Optimization of genomic DNA extraction from cattle blood using modified phenol-chloroform method. J Anim Res. 2019;9(2):285-291.
- Kumar P, Mukesh M, Sodhi M, et al. Genotyping of β-casein gene in Indian cattle and buffalo for detection of A1/A2 alleles. Indian J Anim Sci. 2018;88(1):37-41.
- Kumar S, Patel R, Verma D. Prevalence of A1 and A2 β-casein alleles in Frieswal crossbred cows. J Dairy Res Technol. 2019;7(2):45-50. doi:10.1016/j.jdrt.2019.02.005.
- McLachlan CN. Breeding and milking cows for milk free of beta-casein A1. US Patent 7094949. Available from: http://www.wikipatents.com/USPatent-7094949/breeding-and-milking-cows-for-milk-free-of-beta-casein.
- McLachlan CNS, Elliott R, Harris DP, et al. β-casein A1, milk consumption and human health. N Z Vet J. 2006;54(4):178-180.
- Mukesh M, Kumar P, Sodhi M, et al. β-casein gene polymorphism in Indian indigenous cattle and its implications for A2 milk production. Anim Biotechnol. 2022;33(4):623-631.
- Mukesh M, Swami S, Bhakhri G, et al. Demographic pattern of A1/A2 beta casein variants indicates conservation of A2 type haplotype across native cattle breeds (Bos indicus) of India. 3 Biotech. 2022;12(8):167.
CrossRef - Mukesh M, et al. Demographic pattern of A1/A2 beta casein variants indicates conservation of A2 type haplotype across native cattle breeds (Bos indicus) of India. Sci Rep. 2022;12(1):1-10. doi:10.1038/s41598-022-10123-4.
CrossRef - Mukesh M, Malviya N, Yadav AK. Demographic pattern of A1/A2 beta-casein variants in Indian cattle breeds. 2022. Available from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC9276908/.
- Oliveira RORG, de Carvalho ME, Rodrigues MHD, et al. Allele and genotype frequency for milk beta-casein in dairy cattle in the northern region of Tocantins State, Brazil. Braz J Vet Res Anim Sci. 2021;58:e186603.
CrossRef - Pandey AK, Chauhan JB, Kumar P, et al. Allelic frequency distribution of β-casein gene variants in Indian cattle and buffalo breeds. J Anim Breed Genet. 2019;136(2):132-140.
- Pandey A, Thakur MS, Pandey Y. Association study of polymorphic variants of beta (B) casein gene with milk production traits (lactose, SNF and density) in Malvi, Nimari, Sahiwal and H.F. crossbreeds. Indian J Anim Res.
CrossRef
- Pandey AK, et al. Comparative genotypic analysis of beta-casein gene in Sahiwal and Holstein Friesian crossbred cattle. Int J Livest Res. 2019;9(3):1-7. doi:10.5455/ijlr.20190319010055
- Paradkar PH, Rao A, Singh R. Casein genotypes in Indian Gir and crossbred exotic cattle. Asian J Dairy Food Res. 2023;42(3):121-127. Available from: https://www.arccjournals.com/journal/asian-journal-of-dairy-and-food-research/DR-1659
- Patel JB, Chauhan JB. Polymorphism of the prolactin gene and its relationship with milk production in Gir and Kankrej cattle. J Nat Sci Biol Med. 2017;8(2):167-170.
CrossRef - Ramesha KP, Rao A, Basavaraju M, et al. Genetic variants of β-casein in cattle and buffalo breeding bulls in Karnataka state of India. Indian J Biotechnol. 2016;15:178-181.
- Sambrook J, Russell DW. Molecular Cloning: A Laboratory Manual. 3rd ed. Cold Spring Harbor Laboratory Press; 2001.
- Sawicka-Zugaj W, Jankowski T, Nowak K. Analysis of the frequency of the A1 and A2 alleles in the beta-casein gene and the A, B, and E alleles in the kappa-casein gene in local cattle breeds. Int J Mol Sci. 2025;26(5):2212. Available from: https://www.mdpi.com/1422-0067/26/5/2212
CrossRef - Singh RB, Kumar A, Chauhan A, et al. Indigenous cattle breeds of India: Current status and conservation. Indian J Anim Sci. 2020;90(9):1201-1208.
- Tokarchuk T, Kozhevnikova I, Kudrina M. Bos taurus β-casein: Protein structure, gene polymorphism, effect on the human gastrointestinal tract. Agric Biol. 2023;58(4):598-608.
CrossRef - Yadav A, Sharma R, Kumar P, et al. Evaluation of A1/A2 β-casein polymorphism in Indian cattle and buffalo. Trop Anim Health Prod. 2021;53:393.
- Yadav A, Tomar SS, Jha AK, et al. Molecular characterization of β-casein gene variants in Gir and Holstein Friesian crossbred cattle. Trop Anim Health Prod. 2023;55:177.
- Yadav AK, Pandey A, Pandey Y. Status of A1/A2 beta-casein gene and its association with lactose, SNF%, and density% of milk in Gir and HF crossbred cows. Pharma Innov J. 2023;12(6):268-271. Available from: https://www.thepharmajournal.com/special-issue?ArticleId=21068
- Yadav S, et al. Status of A1/A2 beta-casein gene in Gir and Holstein Friesian crossbred cattle. Indian J Anim Sci. 2023;93(2):123-127.
Accepted on: 14-10_2025
Second Review by: Dr. Muhammad Umar
Final Approval by: Dr. Ghulam Md Ashraf









