Manuscript accepted on : 04 October 2017
Published online on: --
GSTM1/GSTT1 Gene Polymorphism in North Indian Population and their Association to Hypertension
Ritambhara1, Anup Kumar2, Daya Shankar Lal Srivastava3, Sivakumar Vijayaraghavalu4, and Munish Kumar1
1Department of Biochemistry, Faculty of Sciences, University of Allahabad, Allahabad, UP- 211002.
2Department of Statistics, Faculty of Sciences, University of Allahabad, Allahabad, UP-211002.
3Department of Biotechnology and Molecular Medicine, Pt B.D. Sharma Post Graduate Institute of Medical Sciences, Rohtak, Haryana-124001.
4Department of Biomedical Engineering, Lerner Research Institute, Cleveland Clinic Foundation, 9500 Euclid Avenue, Cleveland, Ohio - 44195, USA.
Corresponding Author E-mail: munishkp@gmail.com
DOI : http://dx.doi.org/10.13005/bbra/2569
ABSTRACT: Hypertension has multifactorial background and is a life style related disorder. Arterial hypertension is one of the most common types of hypertension and associated to oxidative stress known to be implicated in its pathogenesis. Glutathione S-transferases (GSTs) are phase II detoxification enzymes that play an important role in cellular defense against oxidative. GSTT1 and GSTM1 gene polymorphisms are reported to be associated with arterial hypertension in several populations and ethnic groups. GSTM1and GSTT1null genotype results in the impaired enzyme activity and consequently greater vulnerability towards various diseases such as cancer. The present study has been done to assess whether GSTT1 and GSTM1 null genotypes are associated with arterial hypertension among North Indian population. We have enrolled 99 arterial hypertension patients and 99 age and sex matched control individuals. Genotyping of GSTM1 and GSTT1 was done by multiplex PCR. We found that GSTM1 null genotype may have risk for hypertension (OR 1.75, CI 0.93-3.2), while for GSTT1 null genotypes shows protection toward hypertension (OR 0.254, CI-0.113-0.57). GSTM1 null genotypes serve as potential genetic factor and may be an independent risk factor for development of hypertension whereas GSTT1 null may have role in protection against hypertension in North Indian populations.
KEYWORDS: GSTM1; GSTT1; Hypertension; Gene Polymorphism; GST (glutathione S-transferase); SBP (systolic blood pressure); DBP (diastolic blood pressure); LDL (low density lipoprotein); HDL (high density lipoprotein)
Download this article as:| Copy the following to cite this article: Ritambhara R, Kumar A, Srivastava D. S. L, Vijayaraghavalu S, Kumar M. GSTM1/GSTT1 Gene Polymorphism in North Indian Population and their Association to Hypertension. Biosci Biotech Res Asia 2017;14(4). |
| Copy the following to cite this URL: Ritambhara R, Kumar A, Srivastava D. S. L, Vijayaraghavalu S, Kumar M. GSTM1/GSTT1 Gene Polymorphism in North Indian Population and their Association to Hypertension. Biosci Biotech Res Asia 2017;14(4). Available from: https://www.biotech-asia.org/?p=28053 |
Introduction
In developing and developed countries, more than 25% adults are prone to death from cardiovascular disease associated directly or indirectly to arterial hypertension [1, 2]. Human arterial hypertension have complex, multifactorial and polygenic niche that interact with several genetic factors and other environmental factors [3, 4]. In a meta-analysis review by Raghupathy et al it is reported that 20.6% men and 20.9% women suffer from hypertension. This ratio would be perpetually increases to 22.9% and 23.6% in men and women respectively by 2025 worldwide [5]. Several studies reported varying prevalence of hypertension in different states of India such as study done in rural Kerala reported 18% hypertension prevalence [6]. Some studies showed hypertension increased prevalence in age groups, for instance a study reported 37% prevalence among population in the age range of 30-60; while a different study reported 55% prevalence in the age range of 40-60 [7, 8]. Study comparing the elderly population from urban and rural areas reported rate of hypertension to be 64% and 55% respectively [9]. In addition to correlating the increased hypertension prevalence to age, some studies also reported the link between hypertension and socio-economic status. Mohan et al. reported 8.4% commonness of hypertension among men and women be owned to be low socio economic level [10]. A comparative study of hypertension performed in urban areas of Chennai, India in low income and high income groups found the percentage frequency rate to be 54% and 40% [11], whereas in a different study Mishra et al reported the prevalence to be 12 % in slums of Delhi, India [12]. Smoking is one of the major risk factors to cause hypertension and cardiovascular disease [13]. Cigarette smoke contains PAHs (polycyclic aromatic hydrocarbons) and hydroxilated metabolites of benzo (α) pyrene which requires bioactivation by phase I xenobiotic metabolizing enzymes (cytochrome p450s) and generates more reactive metabolites that may react with DNA and forms DNA adduct. Phase II xenobiotic enzymes glutathione S-transferase (GSTs) play an important role in detoxification process. It catalyses the conjugation of reduced glutathione with substances that contain electrophilic center [14, 15, 16]. GST’s actions are known to follow phase I xenobiotic metabolism, which is elicited by cytochrome P450 [17]. CYP enzyme catalyses addition of functional groups such as epoxides which provides electrophilic center to reduced glutathione linked to phase II detoxification [18]. Several reports suggested that some genes influence the blood pressure regulation, including IGFBP1, POR, MAWBP, XDH, GSTM1 and GSTT1 [4, 15].
GSTM1 gene locus has been found on chromosome 1p13.3. Three variants have been reported, that differ by C to G substitution at base position 534, and gene deletion [19]. The GSTT1 gene has been located on chromosome 22q11.2, one genotypes has been defined as GSTT-0 while genotype with one functional allele has been denoted as GSTT1 [20]. The deletion polymorphism study of GSTM1 and GSTT1 by gene deletion shows functional abnormality of enzyme in individuals with GSTM1 and GSTT1 null genotypes [19]. Marinho et.al reported that GSTT1 deletion was associated with protective effect on hypertension [4]. However, deletion of GSTM1 has been reported in rats having cerebral stroke associated to hypertension contributing to oxidative stress [21]. Alexey et.al reported that null genotype of GSTT1 polymorphism could be a key player in susceptibility to increased risk of cerebral stroke with association to hypertension [22]. GSTs may be related to risk of various diseases such as alzheimer [23], myocardial infarction [24], Parkinson’s, cancer [25] and diabetes [26]. In the present case control study, we have examined whether the loss of activity of GSTs enzyme due to deletion polymorphism in GSTT1 and GSTM1 may affect the risk in developing hypertension.
Materials and Methods
The institutional ethical committee approved to conduct this study. The selection of individuals in both control and cases groups was based on standard questionnaire. Blood (in EDTA vials) was collected from all subjects who have participated in the study.
Study Population
Selection of normal individual
The study groups consist of 99 normal individuals from North India and have no reported hypertension. The selection method based on those people who have normal blood pressure and normal lipid profile. Smoking habits were noted through detailed questionnaire.
Selection of hypertensive individual
The study group consists of 99 hypertensive individuals from Allahabad city of north India. All subjects examined under same criteria as normal individual and only those were included in the study that have elevated blood pressure and high lipid profile. Selection was based on same questionnaire as control.
Genotyping Analysis
DNA for genotyping was isolated from human peripheral blood (in EDTA vials) by Qaigen DNA blood mini extraction kit. Multiplex PCR was done for analysis of GSTM1 and GSTT1 gene polymorphism (Kumar et.al, 2008). 50-100 ng of isolated DNA was amplified in total volume of 25µl reaction mixture containing 10 pmol of primers. CYPA1 of Exon 7 gene were co-amplified and used as internal control to differentiate between abortive PCR and successful one. The following primer sequence was used for GSTT1, GSTM1and CYPA1:
GSTT1,Forward,5׳TTCCTTACTGGTCCTCACATCTC3׳,Reverse,5׳TCACCGGATCATGGCCAGCA3׳,GSTM1,Forward,5׳GAACTCCCTGAAAAGCTAAAGC3׳,
Reverse,5׳GTTGGGCTCAAATATACGGTGG3׳,andCYPA1,Forward,5׳CTGTCTCCCTCTGGTTACAGGAAGC3׳,Reverse,5׳TTCCACCCGTTGCAGCAGGATAGCC3׳. PCR reaction was carried out in thermal cycler 8800 (Agilent Technologies, Australia). Each set of PCR reaction included both positive and negative controls. Presence and absence of GSTT1 and GSTM1 genes were detected by multiplex PCR method using following PCR cycling condition: initial denaturation at 94●C for 5 min, followed by denaturation 94●C for 1 min, annealing 59●C for 1 min and extension 72●C for 10 min. 2% agrose gel prepared for electrophoresis of multiplex PCR products and visualized by ethidium bromide staining at Geldoc XR+ system. Detection of samples positive for GSTM1 and GSTT1 genotypes matched for 215 and 480bp respectively while CYPA1 yield band of 315bp. The 315bp of CYP1A1 serve as internal control. Presence of 480bp and 215bp fragments indicates GSTT1 and GSTM1non- null genotypes respectively.
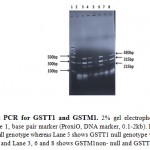 |
Figure 1: Multiplex PCR for GSTT1 and GSTM1. 2% gel electrophoresis of GSTM1 and GSTT1genotype. Lane 1, base pair marker (ProxiO, DNA marker, 0.1-2kb). Lane 2 represents both GSTT1 and GSTM1null genotype whereas Lane 5 shows GSTT1 null genotype whereas Lane 7 shows GSTM1null genotype and Lane 3, 6 and 8 shows GSTM1non- null and GSTT1non-null genotype.
|
Statistical Analysis
Statistical analysis was interpreted with SPSS software version 20.0 (SPSS, Inc.; Chicago). Descriptive measures such as mean and standard deviation were applied for normally distributed variables. Odd ratio and 95% Confidence interval (CI) were used to evaluate impact of GSTM1 and GSTT1 genotype on the risk of developing hypertension. Correlation analysis was performed for all variables which was significant at 0.01 levels (2-tailed) and 0.05 levels (2-tailed). A p-value of <0.05 considered to be statistically significant.
Results
In present study, we have 99 healthy individuals who served as controls and 99 as hypertensive patients. The majority hypertensive cases were old male and have longer persistent of hypertension. In addition, they had higher systolic blood pressure, diastolic blood pressure and LDL (low density lipoprotein) compared to controls. In present study, GSTM1 null genotypes (OR 1.75, CI 0.93-3.2, Table 1) showed non-significant increase predisposition in hypertensive individuals. This association was no longer observed after adjusting age, sex, smoking, blood pressure and lipid profile. However, GSTT1 null genotypes (OR 0.25, 0.11-0.57, Table 1) showed a significant protection toward the hypertension. In combined analysis of GSTM1 and GSTT1 genotypes, both GSTM1/GSTT1 non null variants resulted as protective factor while both GSTT1null/GSTM1 null (OR 2.85, CI 1.24-6.57, Table 1) showed 2.9 times higher risk for hypertension. We found males were 1.5 times more susceptible towards hypertension than females (Table 2). Risk estimate between smokers and non-smokers deduce that smokers were 15 times more vulnerable to have hypertension (Table 3). Our results suggest that age and systolic blood pressure (SBP) were weakly correlated, whereas HDL, total lipid and triglyceride were negatively correlated. Cholesterol levels with SBP and diastolic blood pressure (DBP) were moderately correlated whereas, HDL negatively correlated with cholesterol (Table 4). On the other side, HDL level showed strong negative correlation with age, SBP, DBP, cholesterol, LDL, triglyceride, total lipid and pulse rate. Triglyceride and total lipid were negatively correlated with age and HDL whereas pulse rate showed only negative correlation with HDL.
Table 1: GSTT1, GSTM1 and GSTM1/GSTT1conjugate frequency in controls and hypertensive patients
| Genotype | Control(n=99)% | Hypertensive patients(n=99)% | Odd ratio (95% confidence interval) | P-Value | ||
| GSTT1 | ||||||
| Absent(null) | 9(9.09) | 28(28.29) | ||||
| Present(non-null) | 90(90.00) | 71(71.72) | 0.25(0.11-0.57) | 0.00054 or <0.001 | ||
| GSTM1 | ||||||
| Absent(null) | 33(33.34) | 22(22.23) | ||||
| Present(non-null) | 66(66.67) | 77(77.78) | 1.75(0.93-3.2) | 0.112 | ||
| GSTM1/GSTT1 | ||||||
| Present in both(non -null) | 90(90.91) | 77(77.78) | ||||
| Absent in both (null) | 9(9.09) | 22(22.23) | 2.85(1.24-6.57) | 0.019 | ||
Table 2: Gender susceptibility to hypertension.
| Control(N=89)% | Hypertensive patients(N=93)% | Odd ratio(95% confidence interval) | p-value | |
| Female | 23(25.84) | 32(34.40) | 1.5(0.58-0.48) |
0.258 |
| Male | 66(74.15) | 61(65.59) |
Table 3: Smoking habits between normal and hypertensive subjects.
| Smoking habits | Control(N=89)% | Hypertensive patients(N=93)% | Odd ratio(95% confidence interval) | p-value |
| Non smokers | 79(88.76) | 32(34.40) | 15.05(0.28-0.85) |
<0.001 |
| smokers | 10(11.23) | 61(65.59) |
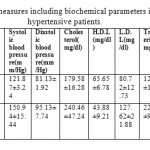 |
Table 4: Statistical measures including biochemical parameters in controls and hypertensive patients.
|
Discussion
Glutathione S-Transferases GSTM1 and GSTT1 are phase II xenobiotic metabolizing enzymes serves to catalyse glutathione conjugation and metabolize number of reactive species. GST enzymes play crucial role to prevent damage to cell against chemical carcinogens and environmental toxicants [31, 32]. These detoxifying enzymes have key role in protecting DNA from various endogenous oxidants; which otherwise could damage DNA and could result in development of cancer [33]. The prevalence range of GSTM1 null genotypes in many population based studies reported it to be 16% to 60% [35]. In our study, we found 33% individuals are GSTM1 null genotypes in controls which are similar to other reports. In case of GSTT1 polymorphism among 99 normal individuals, we found 10% frequency rate which lies in the range of reported values of European and Mediterranean population from 10.4% to 42.5% [35], whereas there is 13-28% deletion for GSTT1 in Caucasians [8]. In Indian region, the frequency of null genotype for GSTM1 and GSTT1 was found to be 22.4% and 17.6% respectively in southern region whereas it was 54% and 13% respectively in eastern region [25, 26]. The present study stated a prevalence of hypertension were 22% GSTM1 null genotype and 28% GSTT1 null genotype polymorphism respectively as determined in North Indian population. We found GSTM1 null genotypes represent independent risk factor for development of hypertension. Our findings were in accordance to Daniel and co-workers as they reported the association of null genotypes of GSTT1 and GSTM1 to hypertension in patients with type 2 diabetes in Caucasians. Ettore and co-workers reported that GSTM1 null variant was independent factor with association to hypertension in Italian region which was accordance to our result. Our result suggests that GSTM1 gene may be considered as major gene associated with susceptibility of hypertension in North Indian population. LDL (low density lipoprotein) shows positive correlation with systolic blood pressure, so it may serves as marker to primary detection of hypertension whereas HDL (high density lipoprotein) shows negative correlation. In contrast to present study, Alexy et.al concluded that GSTT1 null genotype was higher in patients suffering from essential hypertension and cerebral stroke. Abbas and co-workers demonstrated the significant association of null genotypes of GSTM1 and GSTT1 for the hypertension risk. Similarly a recent meta-analysis published by Eslami and coworkers shows that GSTM1 and GSTT1 null genotypes serve as markers for hypertension. However, Kosar et.al reported that null genotype of GSTM1 and GSTT1 were not significantly associated with hypertension. Polimati et.al stated that GSTT1 null genotypes were significantly associated with high risk of hypertension (OR-2.24). In present study males were 1.5 times more susceptible to hypertension than females in contrast a study by Polimati et.al showed the risk was higher in female hypertensive patients (OR-3.25). Recent study by Bessa and coworkers also reported that null genotypes of GSTM1/GSTT1 were main risk factors for the development of hypertension and can be served as a marker in primary detection.
Conclusion
Present study suggest that GSTM1null genotypes may depict as crucial genetic factor to predict development of hypertension, whereas GSTT1 null may have role in protection against hypertension. There is significant association between systolic blood pressure and LDL which may serves as marker to primary detection of hypertension and indicates the pathogenic role in its development among different populations.
Acknowledgment
We acknowledge the Department of Science and Technology, Government of India and University Grant Commission for research support to Dr. Munish Kumar.
Conflicts of interest
No conflict of interest by authors
Funding Source
We acknowledge the Department of Science and Technology, Government of India and University Grant Commission for research support to Dr. Munish Kumar.
References
- Volpe M., Alderman M.H., Furberg C.D., Jackson R., Kostis J.B., Laragh J.H. Beyond hypertension toward guidelines for cardiovascular risk reduction. Am. J. Hypertens., 2004; 17(11):1068-74.
Cross Ref - Petrovic D., Peterlin B. GSTM1-null and GSTT1-null genotypes are associated with essential Arterial hypertension in patients with type 2 diabetes. Clinical. Biochemistry.,2014; 47:574-77.
Cross Ref - Dominiczak A.F., Negrin D.C., Clark J.S., Brosnan M.J., McBride M.W., Alexander M.Y. Genes and hypertension: from gene mapping in experimental models to vascular gene transfer strategies. Hypertension .,2000; 35(1):164-172.
Cross Ref - Marinho C., Alho I., Arduıno D., Falcao L.M., Nogueira J.B., Bicho M. GST M1/T1 and MTHFR polymorphisms as risk factors for hypertension. Biochem.Biophys. Res. Communications., 2007; 353(2): 344-50.
Cross Ref - Kearney P.M., Whelton M., Reynolds K., Muntner P., Whelton P.K., He J. Global burden of hypertension: analysis of worldwide data. Lancet .,2005; 365(9455):217-23.
Cross Ref - Kutty V.R., Balakrishnan K.G., Jayasree A.K., Thomas J. Prevalence of coronary heart disease in the rural population of Thiruvananthapuram district, Kerala, India. Int .J. Cardiol., 1993; 39:59-70.
Cross Ref - Kutty V.R., Soman C.R., Joseph A., Kumar K.V., Pisharody R. Random capillary blood sugar and coronary risk factors in a south Kerala population. J. Cardiovasc .Risk ., 2002; 9:361-67.
Cross Ref - Zachariah M.G., Thankappan K.R., Alex S.C., Sarma P.S., Vasan R.S. Prevalence, correlates, awareness, treatment, and control of hypertension in a middle-aged urban population in Kerala. Indian. Heart. J .,2003; 55:245-51.
- Hypertension study Group. Prevalence, Awareness, treatment and control of hypertension among elderly in Bangladesh and India: a multicentric study. Bulletin of the World Health Organization 2001; 79:490-500.
- Mohan V., Shanthirani S., Deepa R., Premalatha G., Sastry N.G., Saroja R. Chennai Urban Population Study (CUPS No. 4). Intra-urban differences in the prevalence of the metabolic syndrome in southern India — the Chennai Urban Population Study. Diabet .Med., 2001; 18:280-87.
Cross Ref - Ramachandran A., Snehalatha C., Vijay V., King H. Impact of poverty on the prevalence of diabetes and its complications in urban southern India. Diabet. Med., 2002; 19(2):130-35.
Cross Ref - Misra A., Pandey R.M., Devi J.R., Sharma R., Vikram N.K., Khanna N. High prevalence of diabetes, obesity and dyslipidaemia in urban slum population in northern India. Int. J .Obes .Relat. Metab .Disord., 2001; 25(11):1722-29.
Cross Ref - Palma S., Cornetta T., Padua L. Influence of glutathione S-transferase polymorphisms on genotoxic effects induced by tobacco smoke. Mutat .Res., 2007; 633(1):1-12.
Cross Ref - Zheng Z., Park J.Y., Guillemette C., Schantz S.P., Lazarus P. Tobacco carcinogen-detoxifying enzyme UGT1A7 and its association with orolaryngeal cancer risk. J .Natl .Cancer. Inst ., 2001; 19:1411-18.
Cross Ref - Westhoff T.H., Scheid S., Tölle M. A physiogenomic approach to study the regulation of blood pressure. Physiol. Genomics., 2005; 23(1):46-53.
Cross Ref - Kumar M., Agarwal S.K., Goel S.K. Lung cancer risk in north Indian population: role of genetic polymorphisms and smoking. Mol .Cell. Biochem., 2009; 322(1-2):73-79.
Cross Ref - Hayashi S., Watanabe J., Nakachi K., Kawajiri K. Genetic linkage of lung cancer-associated MspI polymorphisms with amino acid replacement in the heme binding region of the human cytochrome P450IA1 gene. J. Biochem., 1991; 110:407-11.
Cross Ref - Sobti R.C., Sharma S., Joshi A., Jindal S.K., Janmeja A. Genetic polymorphism of the CYP1A1, CYP2E1, GSTM1 and GSTT1 genes and lung cancer susceptibility in a north Indian population. Mol.Cell .Biochem., 2004; 266(1-2):1-9.
Cross Ref - Pemble S., Schroeder K.R., Spencer S.R. Human glutathione S-transferase theta (GSTT1) cDNA cloning and the characterization of a genetic polymorphism. Biochem .J .,1994; 300(1):271-276.
Cross Ref - Geisler S.A., Olshan A.F. GSTM1, GSTT1, and the risk of squamous cell carcinoma of the head and neck: a mini HuGE review. Am. J .Epidemiol., 2001; 154(2): 95-105.
Cross Ref - Tamer L., Ercan B., Camsari A. Glutathione S-transferase gene polymorphism as a susceptibility factor in smoking-related coronary artery disease. Basic. Res.Cardiol., 2004; 99(3):223-29.
Cross Ref - Polonikov A., Vialykh E., Vasileva O., Bulgakova I., Bushueva O., Illig T. Genetic Variation in Glutathione S-Transferase Genes and Risk of Nonfatal Cerebral Stroke in Patients Suffering from Essential Hypertension. J. Mol. Neurosci., 2012; 47:511-13.
Cross Ref - Pinhel M.A., Nakazone M.A., Cacao J.C. Glutathione S-transferase variants increase susceptibility for late-onset Alzheimer’s disease: association study and relationship with apolipoprotein E varepsilon4 allele. Clin . Chem . Lab. Med., 2008; 46:439-45.
Cross Ref - Cornelis M.C., El Sohemy A., Campos H. GSTT1 genotype modifies the association between cruciferous vegetable intake and the risk of myocardial infarction. Am. J. Clin .Nutr .,2007; 86: 752-58.
- Zheng Z., Park J.Y., Guillemette C., Schantz S.P., Lazarus P. Tobacco carcinogen-detoxifying enzyme UGT1A7 and its association with orolaryngeal cancer risk. J .Natl .Cancer. Inst., 2001 93(18):1411-18.
Cross Ref - Bridges B.A., Bowyer D.E., Hansen E.S. The possible involvement of somatic mutations in the development of atherosclerotic plaques, Reprot of ICPEMC subcommittee 7/1 conclusions and recommendations. Mutat . Res., 1990; 239:143-87.
Cross Ref - Capoluongo E., Onder G., Concolino P., Russo A., Santonocito C., Bernabei R. GSTM1-null polymorphism as possible risk marker for hypertension: Results from the aging and longevity study in the Sirente Geographic Area (ilSIRENTE study). Clinica. Chimica . Acta ., 2009; 399:92-96.
Cross Ref - Abbas S., Raza S.T., Chandra A., Rizvi S., Ahmed F., Eba A., Mahdi F. Association of ACE, FABP2 and GST genes polymorphism with essential hypertension risk among a North Indian population. Ann. Hum. Biol., 2015; 42:461-69.
Cross Ref - Eslami S., Sahebkar A. Glutathione-Stransferase M1 and T1 null genotypes are associated with Hypertension risk: a systematic review and meta-analysis of 12 studies. Curr. Hypertens. Rep., 2014; 416:32.
- Polimanti R., Piacentini S., Lazzarin N., Re M.A., Manfellotto D., Fuciarelli M. Glutathione S-transferase variants as risk factor for essential hypertension in Italian patients. Mol. Cell .Biochem., 2011; 357(1-2):227-33.
Cross Ref - Bessa S.S., Ali E.M., Hamdy S.M. The role of glutathione-Stransferase M1 and T1 gene polymorphisms and oxidative stressrelated parameters in Egyptian patients with essential hypertension. Eur. J .Intern. Med ., 2009; 20:625-30.
Cross Ref - Hussain K., Salah N., Hussain S., Hussain S. Investigate the Role of Glutathione S Transferase (GST) Polymorphism in Development of Hypertension in UAE Population. Iran. Red. Crescent .Med. J.,2012, 14(8): 479-82.
- Ghosh P., Basu A., Mahata J., Basu S., Sengupta M., Das J.K. Cytogenetic damage and genetic variants in the individuals susceptible to arsenic-induced cancer through drinking water. Int. J .Cancer., 2006; 118:2470-80.
Cross Ref - Vettriselvi V., Vijayalakshmi K., Paul S.F., Venkatachalam P. Genetic variation of GSTM1, GSTT1 and GSTP1 genes in a South Indian population. Asian. Pac. J. Cancer. Prev., 2006; 7(2):325-28.
- Garte S., Gaspari L., Alexandrie A.K., Ambrosone C., Autrup H., Autrup J.L. Metabolic gene polymorphism frequencies in control populations. Cancer. Epidemiol. Biomarkers. Prev., 2001; 10(12):1239-48.
- Zaki M.A., Moghazy T.F., El-Deeb M.M., Mohamed A.H., Mohamed N.A. Glutathione S-transferase M1, T1 and P1 gene polymorphisms and the risk of developing type 2 diabetes mellitus in Egyptian diabetic patients with and without diabetic vascular complications. Alexandria. Journal. of .Medicine., 2015; 51: 73-82.
Cross Ref - Hayes J.D., Strange R.C. Glutathione S-transferase polymorphisms and their biological consequences. Pharmacology., 2000; 61(3):154-66.
Cross Ref - Andreassi M.G., Botto N. DNA damage as a new emerging risk factor in atherosclerosis. Trends.Cardiovasc. Med., 2003; 13(7):270-75.
Cross Ref - Mahmoudi M., Mercer J., Bennett M. DNA damage and repair in atherosclerosis. Cardiovas. Res., 2006; 71(2): 259-68.
Cross Ref

This work is licensed under a Creative Commons Attribution 4.0 International License.





